APOBEC1
Apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1 also known as C->U-editing enzyme APOBEC-1 is a protein that in humans is encoded by the APOBEC1 gene.[5]
This gene encodes a member of the APOBEC protein family and the cytidine deaminase enzyme family. The encoded protein forms a multiple-protein RNA editing holoenzyme with APOBEC1 complementation factor (A1CF). This holoenzyme is involved in the editing of cytosine-to-uracil (C-to-U) nucleotide bases in apolipoprotein B and neurofibromin 1 mRNAs.[5]
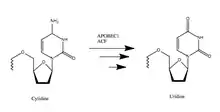
APOBEC-1 (A1) has been linked with cholesterol control, cancer development and inhibition of viral replication.[6] Its function relies on introducing a stop codon into apolipoprotein B (ApoB) mRNA, which alters lipid metabolism in the gastrointestinal tract. The editing mechanism is highly specific. A1’s deamination of the cytosine base yields uracil, which creates a stop codon in the mRNA.
A1 has been linked with both positive and negative health effects. In rodents, it has wide tissue distribution where as in humans, it is only expressed in the small intestine.[7]
Gene
APOBEC1 lies on human chromosome 12.[8]
Function
ApoB is essential in the assembly of very low density lipoproteins from lipids, in the liver and small intestine.[7] By editing ApoB, it forces only the smaller product, ApoB48, to be expressed, which greatly inhibits lipoprotein production. However, A1 is currently found only at extremely low levels in the human liver and intestine, while it is highly expressed in rodents. In humans, A1 is found exclusively in gastrointestinal epithelial cells.[6]
Mechanism
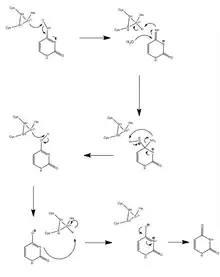
A1 modifies the cytosine base at position 6666 on the ApoB mRNA strand through a deamination.[9] An A1 dimer first binds to ACF, which forms the binding complex that is then able to eliminate the amine group from cytosine.

ACF binds to the mooring sequence, which puts A1 in position to edit the correct residue.[10] By converting cytosine to uracil, A1 changes the codon from CAA, which codes for glutamine during transcription, to UAA, a stop codon.[11] This stop codon yields the much shorter protein ApoB48 instead of ApoB100, as the mRNA is predisposed to transcript.[12] The editing amount, or expression, of A1 performs is correlated with the insulin concentration in the nucleus, the site of modification.[13][14] Tests involving A1 mutants with various deleted amino acid sequences have shown that editing activity is dependent on residues 14 to 35. Like all APOBEC proteins, A1 coordinates a zinc atom with two cysteine and one histidine residues that serve as a Lewis acid. Hydrolytic deamination of the cytosine amine group then occurs, catalyzed by the proton transfer from the nearby glutamic acid residue, and the enzymatic structure is conserved by a proline residue.[10]
Structure
The structure of A1 relies on three dimensional folds induced by a zinc complex.[15] These folds allow the enzyme to access the RNA specifically. Deletion tests with mutant strands have shown that residues 181 to 210 are integral to mRNA editing, and there is most likely a beta-turn at proline residues 190 and 191.[10] Specifically, L182, I185, and L189 are integral to the complex’s function, most likely due to their importance to dimerization.[10] Substituting these residues has no predicted impact on secondary structure, so the significant decrease in editing activity is best explained by the alteration of the side-chains, which are integral to dimer structure.[10] Amino acid replacements at these sites deactivated deamination. The C-terminal of enzyme structure is more strongly expressed in the nucleus, hence the site of modification, while the 181 to 210 residues indicate that the enzyme is in the cytoplasm. These are regulatory factors.[16]
Disease relevance
The low levels of A1 in humans are one reason why high lipid intake is damaging to health. ApoB48 is essential for the assembly and secretion of triglyceride-rich chylomicrons, which are necessary as a response to high-fat intake. ApoB100 are metabolized in the bloodstream to LDL cholesterol,[17] high levels of which are associated with atherosclerosis.[18] While A1 has a negligible impact on human lipid synthesis, at high concentrations it can be genotoxic. Its diffusion toward the nucleic membrane can lead it to mutate DNA sequences that are actively transcribed on the genome. In single growth assays, A1 has been found to impact HIV replications. Additionally, A1 has reduced Hepatitis B virus (HBV) DNA replication, although the mechanism is still not known. The antiviral properties of A1 extend to both DNA and RNA due to its deamination function, which can hinder DNA replication and consequently suppress further infection by HIV or HBV.[19] A pan-cancer study shows that A1 mRNA level is associated with adverse prognosis as well as higher rate of the human genomic insertions and deletions (indels), particularly in-frame ones, which proposes its endogenous mutator activity. [20] There has also been evidence that A1 also edits at NF1, related to tumors in nerve cells.[21]
See also
References
- GRCh38: Ensembl release 89: ENSG00000111701 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000040613 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Entrez Gene: APOBEC1 apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1".
- Rosenberg BR, Hamilton CE, Mwangi MM, Dewell S, Papavasiliou FN (2011). "Transcriptome-wide sequencing reveals numerous APOBEC1 mRNA-editing targets in transcript 3' UTRs". Nat. Struct. Mol. Biol. 18 (2): 230–6. doi:10.1038/nsmb.1975. PMC 3075553. PMID 21258325.
- Teng BB, Ochsner S, Zhang Q, Soman KV, Lau PP, Chan L (1999). "Mutational analysis of apolipoprotein B mRNA editing enzyme (APOBEC1). structure-function relationships of RNA editing and dimerization". J. Lipid Res. 40 (4): 623–35. doi:10.1016/S0022-2275(20)32141-6. PMID 10191286.
- Jarmuz A, Chester A, Bayliss J, Gisbourne J, Dunham I, Scott J, Navaratnam N (2002). "An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22". Genomics. 79 (3): 285–96. doi:10.1006/geno.2002.6718. PMID 11863358.
- Gee P, Ando Y, Kitayama H, Yamamoto SP, Kanemura Y, Ebina H, Kawaguchi Y, Koyanagi Y (2011). "APOBEC1-mediated editing and attenuation of herpes simplex virus 1 DNA indicate that neurons have an antiviral role during herpes simplex encephalitis". J. Virol. 85 (19): 9726–36. doi:10.1128/JVI.05288-11. PMC 3196441. PMID 21775448.
- Smith H (12 September 2008). "The APOBEC1 Paradigm for Mammalian Cytidine Deaminases That Edit DNA and RNA" (PDF). In Henri Grosjean (ed.). DNA and RNA Modification Enzymes: Comparative Structure, Mechanism, Functions, Cellular Interactions and Evolution. Landes Bioscience. Retrieved 24 February 2014.
- Yang Y, Ballatori N, Smith HC (2002). "Apolipoprotein B mRNA editing and the reduction in synthesis and secretion of the atherogenic risk factor, apolipoprotein B100 can be effectively targeted through TAT-mediated protein transduction". Mol. Pharmacol. 61 (2): 269–76. doi:10.1124/mol.61.2.269. PMID 11809850.
- Blanc V, Davidson NO (2011). "Mouse and other rodent models of C to U RNA editing". RNA and DNA Editing. Methods in Molecular Biology. Vol. 718. pp. 121–35. doi:10.1007/978-1-61779-018-8_7. ISBN 978-1-61779-017-1. PMC 3608419. PMID 21370045.
- von Wronski MA, Hirano KI, Cagen LM, Wilcox HG, Raghow R, Thorngate FE, Heimberg M, Davidson NO, Elam MB (1998). "Insulin increases expression of apobec-1, the catalytic subunit of the apolipoprotein B mRNA editing complex in rat hepatocytes". Metab. Clin. Exp. 47 (7): 869–73. doi:10.1016/s0026-0495(98)90128-7. PMID 9667237.
- Yang Y, Sowden MP, Yang Y, Smith HC (2001). "Intracellular trafficking determinants in APOBEC-1, the catalytic subunit for cytidine to uridine editing of apolipoprotein B mRNA". Exp. Cell Res. 267 (2): 153–64. doi:10.1006/excr.2001.5255. PMID 11426934.
- MacGinnitie AJ, Anant S, Davidson NO (1995). "Mutagenesis of apobec-1, the catalytic subunit of the mammalian apolipoprotein B mRNA editing enzyme, reveals distinct domains that mediate cytosine nucleoside deaminase, RNA binding, and RNA editing activity". J. Biol. Chem. 270 (24): 14768–75. doi:10.1074/jbc.270.24.14768. PMID 7782343.
- Lehmann DM, Galloway CA, Sowden MP, Smith HC (2006). "Metabolic regulation of apoB mRNA editing is associated with phosphorylation of APOBEC-1 complementation factor". Nucleic Acids Res. 34 (11): 3299–308. doi:10.1093/nar/gkl417. PMC 1500872. PMID 16820530.
- Nakamuta M, Chang BH, Zsigmond E, Kobayashi K, Lei H, Ishida BY, Oka K, Li E, Chan L (1996). "Complete phenotypic characterization of apobec-1 knockout mice with a wild-type genetic background and a human apolipoprotein B transgenic background, and restoration of apolipoprotein B mRNA editing by somatic gene transfer of Apobec-1". J. Biol. Chem. 271 (42): 25981–8. doi:10.1074/jbc.271.42.25981. PMID 8824235.
- Chen Z, Eggerman TL, Bocharov AV, Baranova IN, Vishnyakova TG, Csako G, Patterson AP (2010). "Hypermutation induced by APOBEC-1 overexpression can be eliminated". RNA. 16 (5): 1040–52. doi:10.1261/rna.1863010. PMC 2856876. PMID 20348446.
- Gonzalez MC, Suspène R, Henry M, Guétard D, Wain-Hobson S, Vartanian JP (2009). "Human APOBEC1 cytidine deaminase edits HBV DNA". Retrovirology. 6: 96. doi:10.1186/1742-4690-6-96. PMC 2770521. PMID 19843348.
- Niavarani, Ahmadreza; Shahrabi Farahani, Asieh; Sharafkhah, Maryam; Rassoulzadegan, Minoo (2018-01-13). "Pancancer analysis identifies prognostic high-APOBEC1 expression level implicated in cancer in-frame insertions and deletions". Carcinogenesis. 39 (3): 327–335. doi:10.1093/carcin/bgy005. ISSN 0143-3334. PMID 29346513.
- Mukhopadhyay D, Anant S, Lee RM, Kennedy S, Viskochil D, Davidson NO (2002). "C-->U editing of neurofibromatosis 1 mRNA occurs in tumors that express both the type II transcript and apobec-1, the catalytic subunit of the apolipoprotein B mRNA-editing enzyme". Am. J. Hum. Genet. 70 (1): 38–50. doi:10.1086/337952. PMC 384902. PMID 11727199.
- Blanc V, Navaratnam N, Henderson JO, Anant S, Kennedy S, Jarmuz A, Scott J, Davidson NO (March 2001). "Identification of GRY-RBP as an apolipoprotein B RNA-binding protein that interacts with both apobec-1 and apobec-1 complementation factor to modulate C to U editing". J. Biol. Chem. 276 (13): 10272–83. doi:10.1074/jbc.M006435200. PMID 11134005.
- Mehta A, Kinter MT, Sherman NE, Driscoll DM (March 2000). "Molecular cloning of apobec-1 complementation factor, a novel RNA-binding protein involved in the editing of apolipoprotein B mRNA". Mol. Cell. Biol. 20 (5): 1846–54. doi:10.1128/MCB.20.5.1846-1854.2000. PMC 85365. PMID 10669759.
- Lau PP, Chan L (Dec 2003). "Involvement of a chaperone regulator, Bcl2-associated athanogene-4, in apolipoprotein B mRNA editing". J. Biol. Chem. 278 (52): 52988–96. doi:10.1074/jbc.M310153200. PMID 14559896.
- Lau PP, Chang BH, Chan L (April 2001). "Two-hybrid cloning identifies an RNA-binding protein, GRY-RBP, as a component of apobec-1 editosome". Biochem. Biophys. Res. Commun. 282 (4): 977–83. doi:10.1006/bbrc.2001.4679. PMID 11352648.
External links
- Human APOBEC1 genome location and APOBEC1 gene details page in the UCSC Genome Browser.
Further reading
- Wedekind JE, Dance GS, Sowden MP, Smith HC (2003). "Messenger RNA editing in mammals: new members of the APOBEC family seeking roles in the family business". Trends Genet. 19 (4): 207–16. doi:10.1016/S0168-9525(03)00054-4. PMID 12683974.
- Harris RS, Liddament MT (2004). "Retroviral restriction by APOBEC proteins". Nat. Rev. Immunol. 4 (11): 868–77. doi:10.1038/nri1489. PMID 15516966. S2CID 10789405.
- Espinosa R, Funahashi T, Hadjiagapiou C, Le Beau MM, Davidson NO (1994). "Assignment of the gene encoding the human apolipoprotein B mRNA editing enzyme (APOBEC1) to chromosome 12p13.1". Genomics. 24 (2): 414–5. doi:10.1006/geno.1994.1645. PMID 7698776.
- Navaratnam N, Bhattacharya S, Fujino T, Patel D, Jarmuz AL, Scott J (1995). "Evolutionary origins of apoB mRNA editing: catalysis by a cytidine deaminase that has acquired a novel RNA-binding motif at its active site". Cell. 81 (2): 187–95. doi:10.1016/0092-8674(95)90328-3. PMID 7736571.
- Lau PP, Zhu HJ, Baldini A, Charnsangavej C, Chan L (1994). "Dimeric structure of a human apolipoprotein B mRNA editing protein and cloning and chromosomal localization of its gene". Proc. Natl. Acad. Sci. U.S.A. 91 (18): 8522–6. Bibcode:1994PNAS...91.8522L. doi:10.1073/pnas.91.18.8522. PMC 44638. PMID 8078915.
- Hadjiagapiou C, Giannoni F, Funahashi T, Skarosi SF, Davidson NO (1994). "Molecular cloning of a human small intestinal apolipoprotein B mRNA editing protein". Nucleic Acids Res. 22 (10): 1874–9. doi:10.1093/nar/22.10.1874. PMC 308087. PMID 8208612.
- Morrison JR, Pászty C, Stevens ME, Hughes SD, Forte T, Scott J, Rubin EM (1996). "Apolipoprotein B RNA editing enzyme-deficient mice are viable despite alterations in lipoprotein metabolism". Proc. Natl. Acad. Sci. U.S.A. 93 (14): 7154–9. Bibcode:1996PNAS...93.7154M. doi:10.1073/pnas.93.14.7154. PMC 38952. PMID 8692961.
- Lau PP, Zhu HJ, Nakamuta M, Chan L (1997). "Cloning of an Apobec-1-binding protein that also interacts with apolipoprotein B mRNA and evidence for its involvement in RNA editing". J. Biol. Chem. 272 (3): 1452–5. doi:10.1074/jbc.272.3.1452. PMID 8999813.
- Oka K, Kobayashi K, Sullivan M, Martinez J, Teng BB, Ishimura-Oka K, Chan L (1997). "Tissue-specific inhibition of apolipoprotein B mRNA editing in the liver by adenovirus-mediated transfer of a dominant negative mutant APOBEC-1 leads to increased low density lipoprotein in mice". J. Biol. Chem. 272 (3): 1456–60. doi:10.1074/jbc.272.3.1456. PMID 8999814.
- Hirano K, Min J, Funahashi T, Baunoch DA, Davidson NO (1997). "Characterization of the human apobec-1 gene: expression in gastrointestinal tissues determined by alternative splicing with production of a novel truncated peptide". J. Lipid Res. 38 (5): 847–59. doi:10.1016/S0022-2275(20)37210-2. PMID 9186903.
- Fujino T, Navaratnam N, Scott J (1998). "Human apolipoprotein B RNA editing deaminase gene (APOBEC1)". Genomics. 47 (2): 266–75. doi:10.1006/geno.1997.5110. PMID 9479499.
- Mehta A, Kinter MT, Sherman NE, Driscoll DM (2000). "Molecular cloning of apobec-1 complementation factor, a novel RNA-binding protein involved in the editing of apolipoprotein B mRNA". Mol. Cell. Biol. 20 (5): 1846–54. doi:10.1128/MCB.20.5.1846-1854.2000. PMC 85365. PMID 10669759.
- Lellek H, Kirsten R, Diehl I, Apostel F, Buck F, Greeve J (2000). "Purification and molecular cloning of a novel essential component of the apolipoprotein B mRNA editing enzyme-complex". J. Biol. Chem. 275 (26): 19848–56. doi:10.1074/jbc.M001786200. PMID 10781591.
- Blanc V, Navaratnam N, Henderson JO, Anant S, Kennedy S, Jarmuz A, Scott J, Davidson NO (2001). "Identification of GRY-RBP as an apolipoprotein B RNA-binding protein that interacts with both apobec-1 and apobec-1 complementation factor to modulate C to U editing". J. Biol. Chem. 276 (13): 10272–83. doi:10.1074/jbc.M006435200. PMID 11134005.
- Lau PP, Chang BH, Chan L (2001). "Two-hybrid cloning identifies an RNA-binding protein, GRY-RBP, as a component of apobec-1 editosome". Biochem. Biophys. Res. Commun. 282 (4): 977–83. doi:10.1006/bbrc.2001.4679. PMID 11352648.
- Anant S, Henderson JO, Mukhopadhyay D, Navaratnam N, Kennedy S, Min J, Davidson NO (2001). "Novel role for RNA-binding protein CUGBP2 in mammalian RNA editing. CUGBP2 modulates C to U editing of apolipoprotein B mRNA by interacting with apobec-1 and ACF, the apobec-1 complementation factor". J. Biol. Chem. 276 (50): 47338–51. doi:10.1074/jbc.M104911200. PMID 11577082.
- Lau PP, Villanueva H, Kobayashi K, Nakamuta M, Chang BH, Chan L (2001). "A DnaJ protein, apobec-1-binding protein-2, modulates apolipoprotein B mRNA editing". J. Biol. Chem. 276 (49): 46445–52. doi:10.1074/jbc.M109215200. PMID 11584023.
- Anant S, Mukhopadhyay D, Sankaranand V, Kennedy S, Henderson JO, Davidson NO (2001). "ARCD-1, an apobec-1-related cytidine deaminase, exerts a dominant negative effect on C to U RNA editing". Am. J. Physiol., Cell Physiol. 281 (6): C1904-16. doi:10.1152/ajpcell.2001.281.6.C1904. PMID 11698249. S2CID 7343760.
- Mukhopadhyay D, Anant S, Lee RM, Kennedy S, Viskochil D, Davidson NO (2002). "C-->U editing of neurofibromatosis 1 mRNA occurs in tumors that express both the type II transcript and apobec-1, the catalytic subunit of the apolipoprotein B mRNA-editing enzyme". Am. J. Hum. Genet. 70 (1): 38–50. doi:10.1086/337952. PMC 384902. PMID 11727199.
- Dance GS, Sowden MP, Cartegni L, Cooper E, Krainer AR, Smith HC (2002). "Two proteins essential for apolipoprotein B mRNA editing are expressed from a single gene through alternative splicing". J. Biol. Chem. 277 (15): 12703–9. doi:10.1074/jbc.M111337200. PMID 11815617.