Aspartate-semialdehyde dehydrogenase
In enzymology, an aspartate-semialdehyde dehydrogenase (EC 1.2.1.11) is an enzyme that is very important in the biosynthesis of amino acids in prokaryotes, fungi, and some higher plants. It forms an early branch point in the metabolic pathway forming lysine, methionine, leucine and isoleucine from aspartate. This pathway also produces diaminopimelate which plays an essential role in bacterial cell wall formation. There is particular interest in ASADH as disabling this enzyme proves fatal to the organism giving rise to the possibility of a new class of antibiotics, fungicides, and herbicides aimed at inhibiting it.
| aspartate-semialdehyde dehydrogenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
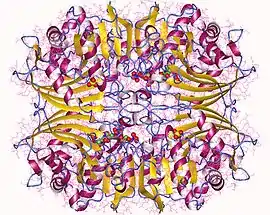 Aspartate semialdehyde dehydrogenase tetramer, Trichophyton rubrum | |||||||||
| Identifiers | |||||||||
| EC no. | 1.2.1.11 | ||||||||
| CAS no. | 9000-98-0 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
The enzyme catalyzes the reversible chemical reaction
- L-aspartate 4-semialdehyde + phosphate + NADP+ L-4-aspartyl phosphate + NADPH + H+
The 3 substrates of this enzyme are L-aspartate 4-semialdehyde, phosphate, and NADP+, whereas its 3 products are L-4-aspartyl phosphate, NADPH, and H+. However, under physiological conditions the reaction runs in the opposite direction.
This enzyme belongs to the family of oxidoreductases, specifically those acting on the aldehyde or oxo group of a donor with NAD+ or NADP+ as acceptor. The systematic name of this enzyme class is L-aspartate-4-semialdehyde:NADP+ oxidoreductase (phosphorylating). Other names in common use include aspartate semialdehyde dehydrogenase, aspartic semialdehyde dehydrogenase, L-aspartate-beta-semialdehyde:NADP+ oxidoreductase, (phosphorylating), aspartic beta-semialdehyde dehydrogenase, and ASA dehydrogenase. This enzyme participates in glycine, serine and threonine metabolism and lysine biosynthesis.
Aspartate-semialdehyde dehydrogenase may be cis-regulated by an Asd RNA motif found in the 5' UTR of some Asd genes.
Protein families
| Semialdehyde dehydrogenase, dimerisation domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Semialdhyde_dhC | ||||||||
| Pfam | PF02774 | ||||||||
| Pfam clan | CL0139 | ||||||||
| InterPro | IPR012280 | ||||||||
| |||||||||
This domain contains both N-acetyl-glutamine semialdehyde dehydrogenase (AgrC), which is involved in arginine biosynthesis, and aspartate-semialdehyde dehydrogenase,[1] an enzyme involved in the biosynthesis of various amino acids from aspartate. It also contains the yeast and fungal Arg5,6 protein, which is cleaved into the enzymes N-acetyl-gamma-glutamyl-phosphate reductase and acetylglutamate kinase. These are also involved in arginine biosynthesis. All proteins in this entry contain a dimerisation domain of semialdehyde dehydrogenase.
Structural studies
As of late 2007, 24 structures have been solved for this class of enzymes, with PDB accession codes 1BRM, 1GL3, 1MB4, 1MC4, 1NWC, 1NWH, 1NX6, 1OZA, 1PQP, 1PQU, 1PR3, 1PS8, 1PU2, 1Q2X, 1T4B, 1T4D, 1TA4, 1TB4, 1YS4, 2EP5, 2GYY, 2GZ1, 2GZ2, and 2GZ3.
References
- Hadfield A, Kryger G, Ouyang J, Petsko GA, Ringe D, Viola R (1999). "Structure of aspartate-beta-semialdehyde dehydrogenase from Escherichia coli, a key enzyme in the aspartate family of amino acid biosynthesis". J Mol Biol. 289 (4): 991–1002. doi:10.1006/jmbi.1999.2828. PMID 10369777.
- BLACK S, WRIGHT NG (1955). "Aspartic beta-semialdehyde dehydrogenase and aspartic beta-semialdehyde". J. Biol. Chem. 213 (1): 39–50. doi:10.1016/S0021-9258(18)71042-9. PMID 14353904.
- Boyer, P.D., Lardy, H. and Myrback, K. (Eds.), The Enzymes, 2nd ed., vol. 7, Academic Press, New York, 1963, p. 203-221.