ATP-dependent Clp protease proteolytic subunit
ATP-dependent Clp protease proteolytic subunit (ClpP) is an enzyme that in humans is encoded by the CLPP gene.[5][6] This protein is an essential component to form the protein complex of Clp protease (Endopeptidase Clp).
| CLPP | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | CLPP, PRLTS3, DFNB81, caseinolytic mitochondrial matrix peptidase proteolytic subunit | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 601119 MGI: 1858213 HomoloGene: 4385 GeneCards: CLPP | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Structure
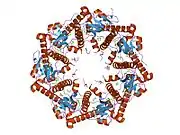
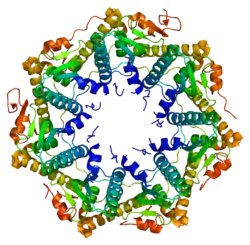
Enzyme ClpP is a highly conserved serine protease present throughout bacteria and also found in the mitochondria and chloroplasts of eukaryotic cells.[7][8] The ClpP monomer is folded into three subdomains: the "handle", the globular "head", and the N-terminal region. By itself, ClpP can assemble into a tetradecamer complex (14-members) and form a closed proteolytic chamber. A fully assembled Clp protease complex has a barrel-shaped structure in which two stacked ring of proteolytic subunits (ClpP or ClpQ) are either sandwiched between two rings or single-caped by one ring of ATPase-active chaperon subunits (ClpA, ClpC, ClpE, ClpX or ClpY). ClpXP is presented in almost all bacteria while ClpA is found in the Gram-negative bacteria, ClpC in Gram-Positive bacteria and cyanobacteria. ClpAP, ClpXP and ClpYQ coexist in E. Coli while only ClpXP complex in present in humans.[9]
Some bacteria have multiple ClpPs, like P. aeruginosa, which has two distinct ClpP isoforms ClpP1 and ClpP2. These isoforms have differences in assembly and functional characteristics. P. aeruginosa produces two forms of the ClpP peptidase, PaClpP114 and PaClpP17P27, which in complex with ClpX or ClpA form functional proteases. PaClpP2 is not able to form an active peptidase on its own but it needs PaClpP1 to be active.[10]
Function
In bacteria, it was shown that ClpP is able to cleave full-length proteins without being associated with ClpA but the degradation is at a much slower rate. Fully functional Clp protease requires the participation of AAA+ ATPase. These ClpX chaperons recognize, unfold and transfer protein substrates to proteolytic core formed by ClpP tetradecamer. The proteolytic sites of ClpP subunits contain hydrophobic grooves which recruit substrate and host the catalytic triad Asp-His-Ser.[11] In several bacteria, such as E. coli, proteins tagged with the SsrA peptide (ANDENYALAA) encoded by tmRNA are digested by Clp proteases.[12] Proteases target damaged or misfolded proteins, transcription factors and signaling proteins in bacteria to coordinate complex cell responses and thus they have robust importance for the physiology and virulence of bacteria.[13]
In P. aeruginosa, ClpP1 is expressed constitutively throughout growth whereas ClpP2 expression is induced 10-fold in stationary phase. Quorum-sensing transcription factor LasR activates expression of ClpP2 in stationary phase. ClpP1 and ClpP2 have differential cleavage specificities which contributes to total peptidase activity of PaClpP17P27. Peptidase and protease action of PaClpP17P27 produces cleavage products that enhance biofilm formation in P. aeruginosa.[10]
The protein encoded by this gene belongs to the peptidase family S14 and hydrolyzes proteins into small peptides in the presence of ATP and magnesium. The protein is transported into mitochondrial matrix and is associated with the inner mitochondrial membrane.[6]
Clinical significance
ClpP protease is a major contributor for mitochondrial protein quality control system and removing damaged or misfolded proteins in mitochondrial matrix. Defects in mitochondrial Clp proteases have been associated with the progression of neurodegenerative diseases while upregulation of ClpP proteases has been implicated in preventing premature aging.[14]
Recessive CLPP mutations were recently observed in the human Perrault variant associating with ovarian failure and sensorineural hearing loss, in parallel with growth retardation. The clinical phenotype was accompanied by the accumulation of ClpP associating partner chaperon ClpX, mtRNA, and inflammatory factors. The disease pathological cause probably involves deficient clearance of mitochondrial components and inflammatory tissue destruction.[15]
ClpP has been shown to be over-expressed in the tumour cells of a subset of cancer patients. This can be exploited by therapeutic agents, including by the hyperactivation of ClpP to cause selective cancer cell lethality.[16]
Intracellular proteases have a role in bacterial virulence. Deletion of ClpP causes growth inhibition or loss of virulence in many bacterial species which makes them a good target for developing new antimicrobial agents. Currently there are no approved antimicrobial agents that target bacterial proteases.[17] PaClpP2 is required for proper biofilm development in opportunistic pathogen P. aeruginosa. Deletion of PaClpP2 or mutation of PaClpP2 active site notably decrease biofilm thickness of P. aeruginosa. This founding has relevance in development of new antimicrobial agents against P. aeruginosa.[10]
References
- GRCh38: Ensembl release 89: ENSG00000125656 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000002660 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Bross P, Andresen BS, Knudsen I, Kruse TA, Gregersen N (December 1995). "Human ClpP protease: cDNA sequence, tissue-specific expression and chromosomal assignment of the gene". FEBS Letters. 377 (2): 249–252. doi:10.1016/0014-5793(95)01353-9. PMID 8543061. S2CID 22019074.
- "Entrez Gene: CLPP ClpP caseinolytic peptidase, ATP-dependent, proteolytic subunit homolog (E. coli)".
- Katayama-Fujimura Y, Gottesman S, Maurizi MR (April 1987). "A multiple-component, ATP-dependent protease from Escherichia coli". The Journal of Biological Chemistry. 262 (10): 4477–4485. doi:10.1016/S0021-9258(18)61217-7. PMID 3549708.
- Corydon TJ, Bross P, Holst HU, Neve S, Kristiansen K, Gregersen N, Bolund L (April 1998). "A human homologue of Escherichia coli ClpP caseinolytic protease: recombinant expression, intracellular processing and subcellular localization". The Biochemical Journal. 331 ( Pt 1) (1): 309–316. doi:10.1042/bj3310309. PMC 1219353. PMID 9512494.
- Hamon MP, Bulteau AL, Friguet B (September 2015). "Mitochondrial proteases and protein quality control in ageing and longevity". Ageing Research Reviews. 23 (Pt A): 56–66. doi:10.1016/j.arr.2014.12.010. PMID 25578288. S2CID 205667759.
- Mawla GD, Hall BM, Cárcamo-Oyarce G, Grant RA, Zhang JJ, Kardon JR, et al. (June 2021). "ClpP1P2 peptidase activity promotes biofilm formation in Pseudomonas aeruginosa". Molecular Microbiology. 115 (6): 1094–1109. doi:10.1111/mmi.14649. PMC 8141546. PMID 33231899.
- Wang J, Hartling JA, Flanagan JM (November 1997). "The structure of ClpP at 2.3 A resolution suggests a model for ATP-dependent proteolysis". Cell. 91 (4): 447–456. doi:10.1016/s0092-8674(00)80431-6. PMID 9390554. S2CID 14136820.
- Gottesman S, Roche E, Zhou Y, Sauer RT (May 1998). "The ClpXP and ClpAP proteases degrade proteins with carboxy-terminal peptide tails added by the SsrA-tagging system". Genes & Development. 12 (9): 1338–1347. doi:10.1101/gad.12.9.1338. PMC 316764. PMID 9573050.
- Hall BM, Breidenstein EB, de la Fuente-Núñez C, Reffuveille F, Mawla GD, Hancock RE, Baker TA (February 2017). O'Toole G (ed.). "Two Isoforms of Clp Peptidase in Pseudomonas aeruginosa Control Distinct Aspects of Cellular Physiology". Journal of Bacteriology. 199 (3). doi:10.1128/JB.00568-16. PMC 5237113. PMID 27849175.
- Luce, K; Weil, AC; Osiewacz, HD (2010). "Mitochondrial Protein Quality Control Systems in Aging and Disease". Protein Metabolism and Homeostasis in Aging. Advances in Experimental Medicine and Biology. Vol. 694. pp. 108–25. doi:10.1007/978-1-4419-7002-2_9. ISBN 978-1-4419-7001-5. PMID 20886760.
- Gispert S, Parganlija D, Klinkenberg M, Dröse S, Wittig I, Mittelbronn M, et al. (December 2013). "Loss of mitochondrial peptidase Clpp leads to infertility, hearing loss plus growth retardation via accumulation of CLPX, mtDNA and inflammatory factors". Human Molecular Genetics. 22 (24): 4871–4887. doi:10.1093/hmg/ddt338. PMC 7108587. PMID 23851121.
- Ishizawa J, Zarabi SF, Davis RE, Halgas O, Nii T, Jitkova Y, et al. (May 2019). "Mitochondrial ClpP-Mediated Proteolysis Induces Selective Cancer Cell Lethality". Cancer Cell. 35 (5): 721–737.e9. doi:10.1016/j.ccell.2019.03.014. PMC 6620028. PMID 31056398.
- Culp, Elizabeth; Wright, Gerard D. (April 2017). "Bacterial proteases, untapped antimicrobial drug targets". The Journal of Antibiotics. 70 (4): 366–377. doi:10.1038/ja.2016.138. ISSN 1881-1469. PMID 27899793. S2CID 31720089.
Further reading
- Kang SG, Dimitrova MN, Ortega J, Ginsburg A, Maurizi MR (October 2005). "Human mitochondrial ClpP is a stable heptamer that assembles into a tetradecamer in the presence of ClpX". The Journal of Biological Chemistry. 280 (42): 35424–35432. doi:10.1074/jbc.M507240200. PMID 16115876.
- Kang SG, Maurizi MR, Thompson M, Mueser T, Ahvazi B (December 2004). "Crystallography and mutagenesis point to an essential role for the N-terminus of human mitochondrial ClpP". Journal of Structural Biology. 148 (3): 338–352. doi:10.1016/j.jsb.2004.07.004. PMID 15522782.
- Kang SG, Ortega J, Singh SK, Wang N, Huang NN, Steven AC, Maurizi MR (June 2002). "Functional proteolytic complexes of the human mitochondrial ATP-dependent protease, hClpXP". The Journal of Biological Chemistry. 277 (23): 21095–21102. doi:10.1074/jbc.M201642200. PMID 11923310.
- de Sagarra MR, Mayo I, Marco S, Rodríguez-Vilariño S, Oliva J, Carrascosa JL, Casta ñ JG (October 1999). "Mitochondrial localization and oligomeric structure of HClpP, the human homologue of E. coli ClpP". Journal of Molecular Biology. 292 (4): 819–825. doi:10.1006/jmbi.1999.3121. PMID 10525407.
- Corydon TJ, Bross P, Holst HU, Neve S, Kristiansen K, Gregersen N, Bolund L (April 1998). "A human homologue of Escherichia coli ClpP caseinolytic protease: recombinant expression, intracellular processing and subcellular localization". The Biochemical Journal. 331 ( Pt 1) (Pt 1): 309–316. doi:10.1042/bj3310309. PMC 1219353. PMID 9512494.
External links
- MEROPS entry for ClpP
- Human CLPP genome location and CLPP gene details page in the UCSC Genome Browser.