Cyclothiazomycin
The cyclothiazomycins are a group of natural products, classified as thiopeptides, which are produced by various Streptomyces species of bacteria.
 | |
| Names | |
|---|---|
| Other names
cyclothiazomycin A, 5102-II | |
| Identifiers | |
3D model (JSmol) |
|
PubChem CID |
|
| |
| |
| Properties | |
| C59H64N18O14S7 | |
| Molar mass | 1473.71 g/mol |
| Melting point | >210 °C (decomposes) |
| Solubility | soluble in water, chloroform/methanol, mixed aqueous/organic solvents (DMSO/water, methanol/water, acetone/water) |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
Infobox references | |
These compounds are ribosomally synthesized and post-translationally modified peptides (RiPPs) and can be further classified as thiopeptides. The overall structure of the cyclothiazomycins comprises a macrocyclic bicyclic peptide containing several thiazoles and thiazolines. The cyclothiazomycins are reported to have multiple inhibitory effects ranging from decreasing blood pressure to interfering with RNA transcription; they also exhibit some antibiotic activity.
History
Cylothiazomycin A was first isolated from Streptomyces sp. NR0516 in 1991.[1] The structure of cyclothiazomycin A was solved via NMR spectroscopy and chemical degradation.[2] Previously, a peptide compound 5102-II had been isolated in 1982 from Streptomyces hygroscopicus 10-22.[3] The discovery of the genes responsible for the biosynthesis of cyclothiazomycin in 2010 showed that 5102-II and cyclothiazomycin A were the same compound.[4]
Structural analogues cyclothiazomycin B1 and B2 were isolated from Streptomyces sp. A307 and solved in 2006 with the help of high-resolution mass spectrometry and NMR spectroscopy.[5]
A third analogue, cyclothiazomycin C, was discovered in 2014 with the aid of genome mining, nucleophilic 1,4-addition labeling reactions, high resolution mass spectrometry, and NMR spectroscopy.[6]
Structure
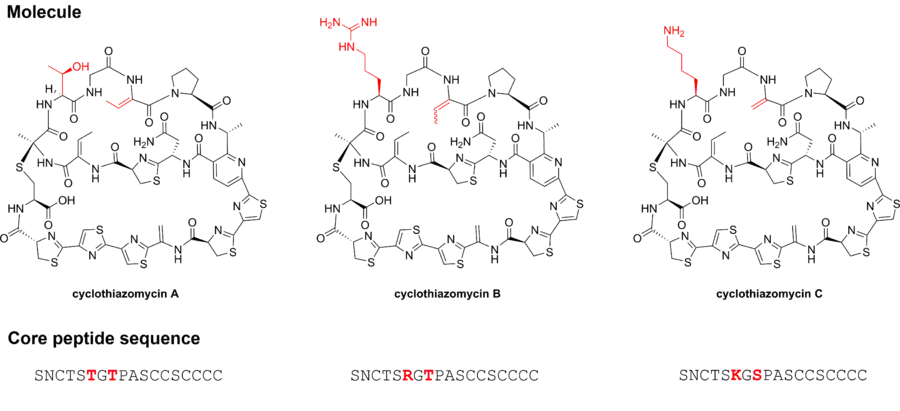
The cyclothiazomycins are thiazole-containing bicyclic and macrocyclic peptides that are structurally similar to thiostrepton and nosiheptide.[2] Formally classified as thiopeptides within the larger family of ribosomally synthesized and post-translationally modified peptides (RiPPs), the cyclothiazomycins are initially biosynthesized as peptides which subsequently undergo chemical modifications, including macrocyclization, aromatization, cyclodehydration, and dehydrogenation. Cyclothiazomycins exist as three analogues that differ at two amino acid residues in the core peptide. The image below denotes in red the differences between the analogues as well as the amino acid sequence that becomes the final compounds.
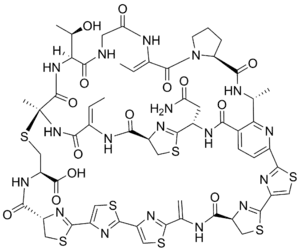
The structure of cyclothiazomycin A was established primarily by two-dimensional nuclear magnetic resonance spectroscopy. Cyclothiazomycin A contains a dehydroalanine and two dehydrohomoalanine residues within a bicyclic, macrocyclic scaffold composed of thiazolines, thiazoles, and a trisubstituted pyridine.[2]
Cyclothiazomycin B and C vary from cyclothiazomycin A at the second and third threonine residues in the core sequence of the precursor peptide. In cyclothiazomycin B, the second threonine is an arginine. Cyclothiazomycin B exists as a pair of diastereomers, cyclothiazomycin B1 and cyclothiazomycin B2, that differ at the configuration of one of its dehydrohomoalanine moieties. Cyclothiazomycin B1 and B2 interconvert by isomerization in solution but are stable in the solid state.[5]
Cyclothiazomycin C has a lysine in place of the second threonine residue found in cyclothiazomycin A and a serine residue in place of the third threonine residue, resulting in a dehydroalanine instead of the dehydrohomoalanine observed in cyclothiazomycin A.[6]
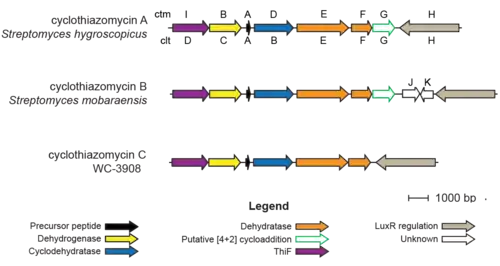
Biosynthesis
Cyclothiazomycin is part of a class of natural products that are ribosomally synthesized and post-translationally modified peptides (RiPPs). As such, cyclothiazomycin begins as a peptide that is synthesized in the bacterial ribosome. A series of chemical steps by biosynthetic enzymes transform the original peptide into the final (mature) natural product.
The gene encoding for cyclothiazomycin begins with a short open reading frame (ORF) ctmA (cltA). CtmD (cltB) encodes a "fused" TOMM cyclodehydratase believed to take part in the formation of thiazolines. The enzyme encoded by the ctmB (cltC) gene is believed to catalyze the dehydrogenation of thiazolines to thiazoles. CtmE and ctmF (ctlE and cltF) each encode a split lanthipeptide dehydratase which dehydrates serine and threonine to dehydroalanine and dehydrobutyrine. CtmG (cltG) is believed to aide in the production of the central pyridine. CtmI (cltD) encodes a ThiF-like protein. CtmH (cltH) is a LuxR-type regulatory gene. CtmJK are not present in the cyclothiazomycin A and C clusters; there is no known function. The cltN gene is believed to encode an enzyme that is involved in the formation of the tertiary thioether, however this has not been proven and cltN is not regulated by ctmH.CtmA, ctmD, ctmF, and ctmG deletion leads to the disappearance of cyclothiazomycin A.[3][4][7]
CtmH is a LuxR-type regulatory gene that was shown to regulate ctmA-ctmL.[7]
Bioactivity
Each analogue of cyclothiazomycin is believed to interact with at least one biological target. Cyclothiazomycin A has been shown to inhibit blood plasma renin leading to a decrease blood pressure; it also exhibits weak anti-fungal activity.[1][3] Cyclothiazomycin B shows inhibitory activity against RNA polymerase, and it is believed to act by reducing ribosome dependent GTPase.[5] Cyclothiazomycin B also exhibits anti-fungal activity by binding to the chitin of cell walls, thus producing a fragile cell wall.[8] Recently discovered cyclothiazomycin C has an unknown biological role, but cyclothiazomycins B and C both exhibit antibacterial activity against Gram-positive bacteria, including Bacillus anthracis.[6]
References
- Aoki, M; Ohtsuka, T; Yamada, M; Ohba, Y; Yoshizaki, H; Yasuno, H; Sano, T; Watanabe, J; Yokose, K; Seto, H (Jun 1991). "Cyclothiazomycin, a novel polythiazole-containing peptide with renin inhibitory activity. Taxonomy, fermentation, isolation and physico-chemical characterization". The Journal of Antibiotics. 44 (6): 582–8. doi:10.7164/antibiotics.44.582. PMID 2071486.
- Aoki, Masahiro; Ohtsuka, Tatsuo; Itezono, Yoshiko; Yokose, Kazuteru; Furihata, Kazuo; Seto, Haruo (1991). "Structure of cyclothiazomycin, a unique polythiazole-containing peptide with renin inhibitory activity. Part 2. Total structure". Tetrahedron Letters. 32 (2): 221–224. doi:10.1016/0040-4039(91)80860-9.
- Zhang, S.; Zhau, H; Liu, J (1982). "Studies on the agricultural antibiotics 5102-II. isolation and characterization of antibiotic 5102-II". Acta Microbiologica Sinica. 22: 145–150.
- Wang, J.; Yu, Y.; Tang, K.; Liu, W.; He, X.; Huang, X.; Deng, Z. (12 February 2010). "Identification and Analysis of the Biosynthetic Gene Cluster Encoding the Thiopeptide Antibiotic Cyclothiazomycin in Streptomyces hygroscopicus 10-22". Applied and Environmental Microbiology. 76 (7): 2335–2344. doi:10.1128/AEM.01790-09. PMC 2849233. PMID 20154110.
- Hashimoto, Masaru; Murakami, Takanori; Funahashi, Katsuyuki; Tokunaga, Takashi; Nihei, Ken-ichi; Okuno, Toshikatsu; Kimura, Takatsugu; Naoki, Hideo; Himeno, Hyouta (2006). "An RNA polymerase inhibitor, cyclothiazomycin B1, and its isomer". Bioorganic & Medicinal Chemistry. 14 (24): 8259–8270. doi:10.1016/j.bmc.2006.09.006. PMID 17010619.
- Cox, Courtney L.; Tietz, Jonathan I.; Sokolowski, Karol; Melby, Joel O.; Doroghazi, James R.; Mitchell, Douglas A. (17 June 2014). "Nucleophilic 1,4-Additions for Natural Product Discovery". ACS Chemical Biology. 9 (9): 2014–2022. doi:10.1021/cb500324n. PMC 4168802. PMID 24937678.
- Zhang, P.; Wu, H.; Chen, X.-L.; Deng, Z.; Bai, L.; Pang, X. (25 April 2014). "Regulation of the biosynthesis of thiopeptide antibiotic cyclothiazomycin by the transcriptional regulator SHJG8833 in Streptomyces hygroscopicus 5008". Microbiology. 160 (Pt_7): 1379–1392. doi:10.1099/mic.0.076901-0. PMID 24769844.
- Mizuhara, Naoko; Kuroda, Manabu; Ogita, Akira; Tanaka, Toshio; Usuki, Yoshinosuke; Fujita, Ken-ichi (2011). "Antifungal thiopeptide cyclothiazomycin B1 exhibits growth inhibition accompanying morphological changes via binding to fungal cell wall chitin". Bioorganic & Medicinal Chemistry. 19 (18): 5300–5310. doi:10.1016/j.bmc.2011.08.010. PMID 21885289.