Structural biology
Structural biology is a field that is many centuries old which, as defined by the Journal of Structural Biology, deals with structural analysis of living material (formed, composed of, and/or maintained and refined by living cells) at every level of organization. Early structural biologists throughout the 19th and early 20th centuries were primarily only able to study structures to the limit of the naked eye's visual acuity and through magnifying glasses and light microscopes.
| Part of a series on |
| Biochemistry |
|---|
 |
|
In the 20th century, a variety of experimental techniques were developed to examine the 3D structures of biological molecules. The most prominent techniques are X-ray crystallography, nuclear magnetic resonance, and electron microscopy. Through the discovery of X-rays and its applications to protein crystals, structural biology was revolutionized, as now scientists could obtain the three-dimensional structures of biological molecules in atomic detail.[1] Likewise, NMR spectroscopy allowed information about protein structure and dynamics to be obtained.[2] Finally, in the 21st century, electron microscopy also saw a drastic revolution with the development of more coherent electron sources, aberration correction for electron microscopes, and reconstruction software that enabled the successful implementation of high resolution cryo-electron microscopy, thereby permitting the study of individual proteins and molecular complexes in three-dimensions at angstrom resolution.
With the development of these three techniques, the field of structural biology expanded and also became a branch of molecular biology, biochemistry, and biophysics concerned with the molecular structure of biological macromolecules (especially proteins, made up of amino acids, RNA or DNA, made up of nucleotides, and membranes, made up of lipids), how they acquire the structures they have, and how alterations in their structures affect their function.[3] This subject is of great interest to biologists because macromolecules carry out most of the functions of cells, and it is only by coiling into specific three-dimensional shapes that they are able to perform these functions. This architecture, the "tertiary structure" of molecules, depends in a complicated way on each molecule's basic composition, or "primary structure." At lower resolutions, tools such as FIB-SEM tomography have allowed for greater understanding of cells and their organelles in 3-dimensions, and how each hierarchical level of various extracellular matrices contributes to function (for example in bone). In the past few years it has also become possible to predict highly accurate physical molecular models to complement the experimental study of biological structures.[4] Computational techniques such as molecular dynamics simulations can be used in conjunction with empirical structure determination strategies to extend and study protein structure, conformation and function.[5]
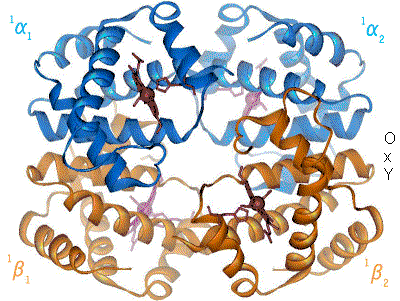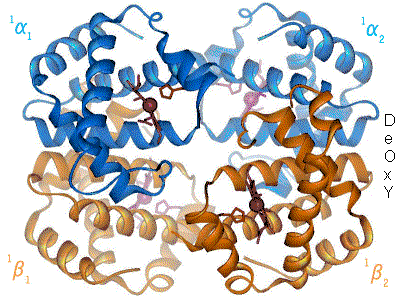
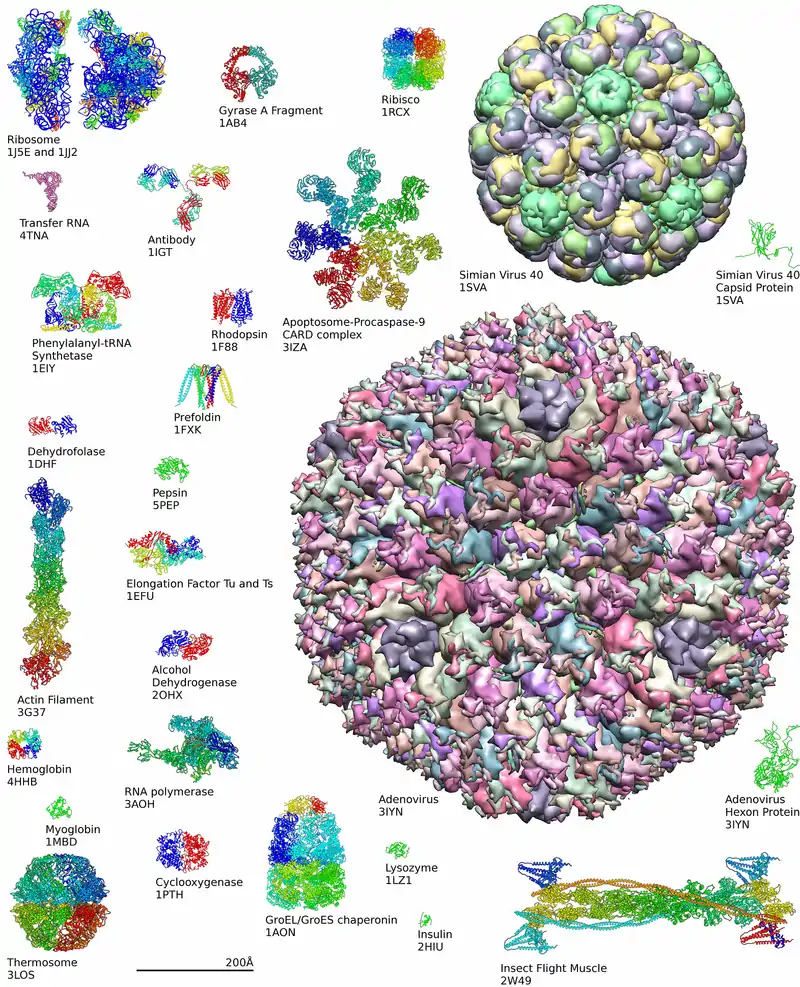
History
In 1912 Max Von Laue directed X-rays at crystallized copper sulfate generating a diffraction pattern.[6] These experiments led to the development of X-ray crystallography, and its usage in exploring biological structures.[4] In 1951, Rosalind Franklin and Maurice Wilkins used X-ray diffraction patterns to capture the first image of deoxyribonucleic acid (DNA). Francis Crick and James Watson modeled the double helical structure of DNA using this same technique in 1953 and received the Nobel Prize in Medicine along with Wilkins in 1962.[7]
Pepsin crystals were the first proteins to be crystallized for use in X-ray diffraction, by Theodore Svedberg who received the 1962 Nobel Prize in Chemistry.[8] The first tertiary protein structure, that of myoglobin, was published in 1958 by John Kendrew.[9] During this time, modeling of protein structures was done using balsa wood or wire models.[10] With the invention of modeling software such as CCP4 in the late 1970s,[11] modeling is now done with computer assistance. Recent developments in the field have included the generation of X-ray free electron lasers, allowing analysis of the dynamics and motion of biological molecules,[12] and the use of structural biology in assisting synthetic biology.[13]
In the late 1930s and early 1940s, the combination of work done by Isidor Rabi, Felix Bloch, and Edward Mills Purcell led to the development of nuclear magnetic resonance (NMR). Currently, solid-state NMR is widely used in the field of structural biology to determine the structure and dynamic nature of proteins (protein NMR).[14]
In 1990, Richard Henderson produced the first three-dimensional, high resolution image of bacteriorhodopsin using cryogenic electron microscopy (cryo-EM).[15] Since then, cryo-EM has emerged as an increasingly popular technique to determine three-dimensional, high resolution structures of biological images.[16]
More recently, computational methods have been developed to model and study biological structures. For example, molecular dynamics (MD) is commonly used to analyze the dynamic movements of biological molecules. In 1975, the first simulation of a biological folding process using MD was published in Nature.[17] Recently, protein structure prediction was significantly improved by a new machine learning method called AlphaFold.[18] Some claim that computational approaches are starting to lead the field of structural biology research.[19]
Techniques
Biomolecules are too small to see in detail even with the most advanced light microscopes. The methods that structural biologists use to determine their structures generally involve measurements on vast numbers of identical molecules at the same time. These methods include:
- Mass spectrometry
- Macromolecular crystallography
- Neutron diffraction
- Proteolysis
- Nuclear magnetic resonance spectroscopy of proteins (NMR)
- Electron paramagnetic resonance (EPR)
- Cryogenic electron microscopy (cryoEM)
- Electron crystallography and microcrystal electron diffraction
- Multiangle light scattering
- Small angle scattering
- Ultrafast laser spectroscopy
- Anisotropic terahertz microspectroscopy
- Two-dimensional infrared spectroscopy
- Dual-polarization interferometry and circular dichroism
Most often researchers use them to study the "native states" of macromolecules. But variations on these methods are also used to watch nascent or denatured molecules assume or reassume their native states. See protein folding.
A third approach that structural biologists take to understanding structure is bioinformatics to look for patterns among the diverse sequences that give rise to particular shapes. Researchers often can deduce aspects of the structure of integral membrane proteins based on the membrane topology predicted by hydrophobicity analysis. See protein structure prediction.
Applications
Structural biologists have made significant contributions towards understanding the molecular components and mechanisms underlying human diseases. For example, cryo-EM and ssNMR have been used to study the aggregation of amyloid fibrils, which are associated with Alzheimer's disease, Parkinson's disease, and type II diabetes.[20] In addition to amyloid proteins, scientists have used cryo-EM to produce high resolution models of tau filaments in the brain of Alzheimer's patients which may help develop better treatments in the future.[21] Structural biology tools can also be used to explain interactions between pathogens and hosts. For example, structural biology tools have enabled virologists to understand how the HIV envelope allows the virus to evade human immune responses.[22]
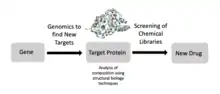
Structural biology is also an important component of drug discovery.[23] Scientists can identify targets using genomics, study those targets using structural biology, and develop drugs that are suited for those targets. Specifically, ligand-NMR, mass spectrometry, and X-ray crystallography are commonly used techniques in the drug discovery process. For example, researchers have used structural biology to better understand Met, a protein encoded by a protooncogene that is an important drug target in cancer.[24] Similar research has been conducted for HIV targets to treat people with AIDS.[23] Researchers are also developing new antimicrobials for mycobacterial infections using structure-driven drug discovery.[23]
See also
- Primary structure
- Secondary structure
- Tertiary structure
- Quaternary structure
- Structural domain
- Structural motif
- Protein subunit
- Molecular model
- Cooperativity
- Chaperonin
- Structural genomics
- Stereochemistry
- Resolution (electron density)
- Proteopedia The collaborative, 3D encyclopedia of proteins and other molecules.
- Protein structure prediction
References
- Curry, Stephen (2015-07-03). "Structural Biology: A Century-long Journey into an Unseen World". Interdisciplinary Science Reviews. 40 (3): 308–328. Bibcode:2015ISRv...40..308C. doi:10.1179/0308018815Z.000000000120. ISSN 0308-0188. PMC 4697198. PMID 26740732.
- Campbell, Iain D. (2013-01-01). "The evolution of protein NMR". Biomedical Spectroscopy and Imaging. 2 (4): 245–264. doi:10.3233/BSI-130055. ISSN 2212-8794.
- Banaszak LJ (2000). Foundations of Structural Biology. Burlington: Elsevier. ISBN 9780080521848.
- Jumper, John; Evans, Richard; Pritzel, Alexander; Green, Tim; Figurnov, Michael; Ronneberger, Olaf; Tunyasuvunakool, Kathryn; Bates, Russ; Žídek, Augustin; Potapenko, Anna; Bridgland, Alex; Meyer, Clemens; Kohl, Simon A. A.; Ballard, Andrew J.; Cowie, Andrew (2021-07-15). "Highly accurate protein structure prediction with AlphaFold". Nature. 596 (7873): 583–589. Bibcode:2021Natur.596..583J. doi:10.1038/s41586-021-03819-2. ISSN 1476-4687. PMC 8371605. PMID 34265844.
- Karplus M, McCammon JA (September 2002). "Molecular dynamics simulations of biomolecules". Nature Structural Biology. 9 (9): 646–652. doi:10.1038/nsb0902-646. PMID 12198485. S2CID 590640.
- Curry S (July 2015). "Structural Biology: A Century-long Journey into an Unseen World". Interdisciplinary Science Reviews. 40 (3): 308–328. Bibcode:2015ISRv...40..308C. doi:10.1179/0308018815z.000000000120. PMC 4697198. PMID 26740732.
- "The Nobel Prize in Physiology or Medicine 1962". NobelPrize.org. Retrieved 2022-10-01.
- Jaskolski M, Dauter Z, Wlodawer A (September 2014). "A brief history of macromolecular crystallography, illustrated by a family tree and its Nobel fruits". The FEBS Journal. 281 (18): 3985–4009. doi:10.1111/febs.12796. PMC 6309182. PMID 24698025.
- Kendrew JC, Bodo G, Dintzis HM, Parrish RG, Wyckoff H, Phillips DC (March 1958). "A three-dimensional model of the myoglobin molecule obtained by x-ray analysis". Nature. 181 (4610): 662–666. Bibcode:1958Natur.181..662K. doi:10.1038/181662a0. PMID 13517261. S2CID 4162786.
- Garman EF (March 2014). "Developments in x-ray crystallographic structure determination of biological macromolecules". Science. 343 (6175): 1102–1108. Bibcode:2014Sci...343.1102G. doi:10.1126/science.1247829. PMID 24604194. S2CID 21067016.
- "About CCP4". legacy.ccp4.ac.uk. Retrieved 2021-04-02.
- Waldrop MM (January 2014). "X-ray science: The big guns". Nature. 505 (7485): 604–606. Bibcode:2014Natur.505..604W. doi:10.1038/505604a. PMID 24476872.
- Kiel C, Serrano L (November 2012). "Structural data in synthetic biology approaches for studying general design principles of cellular signaling networks". Structure. 20 (11): 1806–1813. doi:10.1016/j.str.2012.10.002. PMID 23141693.
- Wüthrich K (November 2001). "The way to NMR structures of proteins". Nature Structural Biology. 8 (11): 923–925. doi:10.1038/nsb1101-923. PMID 11685234. S2CID 26153265.
- Henderson R, Baldwin JM, Ceska TA, Zemlin F, Beckmann E, Downing KH (June 1990). "Model for the structure of bacteriorhodopsin based on high-resolution electron cryo-microscopy". Journal of Molecular Biology. 213 (4): 899–929. doi:10.1016/S0022-2836(05)80271-2. PMID 2359127.
- Callaway, Ewen (2020-02-10). "Revolutionary cryo-EM is taking over structural biology". Nature. 578 (7794): 201. Bibcode:2020Natur.578..201C. doi:10.1038/d41586-020-00341-9. PMID 32047310. S2CID 211081167.
- Levitt M, Warshel A (February 1975). "Computer simulation of protein folding". Nature. 253 (5494): 694–698. Bibcode:1975Natur.253..694L. doi:10.1038/253694a0. PMID 1167625. S2CID 4211714.
- Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. (August 2021). "Highly accurate protein structure prediction with AlphaFold". Nature. 596 (7873): 583–589. Bibcode:2021Natur.596..583J. doi:10.1038/s41586-021-03819-2. PMC 8371605. PMID 34265844.
- Nussinov, Ruth; Tsai, Chung-Jung; Shehu, Amarda; Jang, Hyunbum (2019-02-12). "Computational Structural Biology: Successes, Future Directions, and Challenges". Molecules (Basel, Switzerland). 24 (3): E637. doi:10.3390/molecules24030637. ISSN 1420-3049. PMC 6384756. PMID 30759724.
- Iadanza MG, Jackson MP, Hewitt EW, Ranson NA, Radford SE (December 2018). "A new era for understanding amyloid structures and disease". Nature Reviews. Molecular Cell Biology. 19 (12): 755–773. doi:10.1038/s41580-018-0060-8. PMID 30237470. S2CID 52307956.
- Fitzpatrick, Anthony W. P.; Falcon, Benjamin; He, Shaoda; Murzin, Alexey G.; Murshudov, Garib; Garringer, Holly J.; Crowther, R. Anthony; Ghetti, Bernardino; Goedert, Michel; Scheres, Sjors H. W. (2017-07-13). "Cryo-EM structures of tau filaments from Alzheimer's disease". Nature. 547 (7662): 185–190. Bibcode:2017Natur.547..185F. doi:10.1038/nature23002. ISSN 1476-4687. PMC 5552202. PMID 28678775.
- Engelman, Alan; Cherepanov, Peter (2012-03-16). "The structural biology of HIV-1: mechanistic and therapeutic insights". Nature Reviews Microbiology. 10 (4): 279–290. doi:10.1038/nrmicro2747. ISSN 1740-1526. PMC 3588166. PMID 22421880. S2CID 14088805.
- Thomas SE, Mendes V, Kim SY, Malhotra S, Ochoa-Montaño B, Blaszczyk M, Blundell TL (August 2017). "Structural Biology and the Design of New Therapeutics: From HIV and Cancer to Mycobacterial Infections: A Paper Dedicated to John Kendrew". Journal of Molecular Biology. John Kendrew’s 100th Anniversary Special Edition. 429 (17): 2677–2693. doi:10.1016/j.jmb.2017.06.014. PMID 28648615.
- Wendt KU, Weiss MS, Cramer P, Heinz DW (February 2008). "Structures and diseases". Nature Structural & Molecular Biology. 15 (2): 117–120. doi:10.1038/nsmb0208-117. PMC 7097323. PMID 18250627.
