1q21.1 deletion syndrome
1q21.1 deletion syndrome is a rare aberration of chromosome 1. A human cell has one pair of identical chromosomes on chromosome 1. With the 1q21.1 deletion syndrome, one chromosome of the pair is not complete, because a part of the sequence of the chromosome is missing. One chromosome has the normal length and the other is too short.
| 1q21.1 deletion syndrome | |
|---|---|
| Other names | Chromosome 1q21.1 microdeletion syndrome, 1q21.1 microdeletion, Monosomy 1q21.1, Del(1)(q21), 1q21.1 contiguous gene deletion, 1q21.1 deletion, Chromosome 1q21.1 deletion syndrome, Chromosome 1q21.1 deletion syndrome, 1.35-Mb. |
| Specialty | Genetics. |
| Symptoms | Delayed development, intellectual disability, physical abnormalities, neurological abnormalities and psychiatric problems.[1] |
| Usual onset | Infancy.[2] |
| Causes | Deletion of genetic material on the long arm (or q arm) of chromosome 1 at position 21.1.[3] |
| Diagnostic method | Chromosomal microarray analysis.[4] |
| Differential diagnosis | 22q11.2 microdeletion syndrome.[4] |
| Frequency | 0.015% of the population.[5] |
In 1q21.1, the '1' stands for chromosome 1, the 'q' stands for the long arm of the chromosome and '21.1' stands for the part of the long arm in which the deletion is situated.
The syndrome is a form of the 1q21.1 copy number variations, and it is a deletion in the distal area of the 1q21.1 part. The CNV leads to a very variable phenotype, and the manifestations in individuals are quite variable. Some people who have the syndrome can function in a normal way, while others have symptoms of intellectual impairment and various physical anomalies.[1]
1q21.1 microdeletion is a very rare chromosomal condition. Only 46 individuals with this deletion have been reported in medical literature as of August 2011.[3]
Signs and symptoms
Approximately 75% of all children with a 1q21.1 microdeletion exhibit delayed development, notably in motor skills such as sitting, standing, and walking.[1] Individuals may have generalized mild learning difficulties; about 30% of those diagnosed with 1q21.1 deletion syndrome are affected.[4]
Dysmorphic craniofacial traits are common, however, they are highly varied and thus difficult to identify. Microcephaly has been reported in 39% of those with the 1q21.1 deletion.[4][2]
It is not clear whether the list of symptoms is complete. Very little information is known about the syndrome. The syndrome can have completely different effects on members of the same family.[1][4]
Genitourinary abnormalities include vesicoureteral reflux, hydronephrosis, inguinal hernia, cryptorchidism, and genital malformations. There have been two reported cases of Mayer-Rokitansky-Kuster-Hauser syndrome alongside 1q21.1 deletion syndrome.[6][4][1]
The majority of affected people have normal neurologic physical examinations, however hypotonia and tremors are quite common. Seizures affect roughly 16% of children and typically begin during infancy.[1][4]
Possible psychiatric and behavioral abnormalities include autism spectrum disorder, attention deficit hyperactivity disorder, and mood disorders. Furthermore, distal 1q21.1 microdeletions have been found in 0.2%-0.6% of those with schizophrenia.[4][7]
A common deletion is between 1.0-1.9Mb. Mefford states that the standard for a deletion is 1.35Mb.[8] The largest deletion seen on a living human is over 5 Mb.
Causes
1q21.1 deletion is autosomal dominant, with 18%-50% of deletions developing de novo and 50%-82% inherited from a parent.[4][1]
The majority of affected individuals are missing a 1.35 million DNA base pair sequence, also written as as 1.35 megabases, in the q21.1 region of chromosome 1. Though the exact size of the deleted region differs. This deletion affects one of each cell's two copies of chromosome 1.[1]
The symptoms caused by a 1q21.1 microdeletion are most likely due to the loss of many genes in this region. Researchers are attempting to determine which missing genes may contribute to 1q21.1 deletions' specific characteristics. Because some people with a 1q21.1 microdeletion have no clear associated traits, it is hypothesized that additional genetic and environmental factors influence the development of symptoms.[1]
Due to this genetic misprint, the embryo may experience problems in the development during the first months of pregnancy. Approximately 20 to 40 days after fertilization, something goes wrong in the construction of the body parts and brain, which leads to a chain reaction.[9]
Genetics
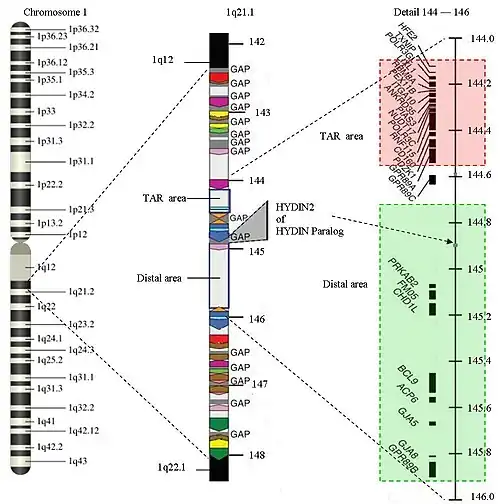
The structure of 1q21.1
The structure of 1q21.1 is complex. The area has a size of approximately 6 Megabase (Mb) (from 141.5 Mb to 147.9 Mb). Within 1q21.1 there are two areas where the CNVs can be found: the proximal area or TAR area (144.1 to 144.5) and the distal area (144.7 to 145.9). The 1q21.1 deletion syndrome will commonly be found in the distal area, but an overlap with the TAR-area is possible. 1q21.1 has multiple repetitions of the same structure (areas with the same color in the picture have equal structures) Only 25% of the structure is not duplicated. There are several gaps in the sequence. There is no further information available about the DNA-sequence in those areas up until now. The gaps represent approximately 700 Kilobase. New genes are expected in the gaps. Because the gaps are still a topic of research, it is hard to find the exact start and end markers of a deletion. The area of 1q21.1 is one of the most difficult parts of the human genome to map.
Because of the repetitions in 1q21.1, there is a larger chance of an unequal crossing-over during meiosis. CNVs occur due to non-allelic homologous recombination mediated by low copy repeats (sequentially similar regions).
Typing
A common deletion is restricted to the distal area. This is a Class I-deletion.
In some cases the deletion is so large that the proximal area is involved as well, the so-called Class II-deletion. There are some complex cases in which both the proximal area and the distal area are affected, while the area in between is normal. There are also some a-typical variants.
Diagnosis
The recurring distal 1.35-Mb heterozygous deletion within the position of 145-146.35 Mb in the reference genome confirms the diagnosis of 1q21.1 deletion. The copy number of sequences can be determined using chromosome microarrays or targeted deletion analysis by fluorescence in situ hybridization. [4]
Differential diagnosis
22q11.2 microdeltion shares several characteristics with 1q21.1 microdeletion including developmental delays, learning disabilities, intellectual disability, and behavioral deviations. Those with recurrent 1q21.1 microdeletion, on the other hand, do not have the distinctive facial traits seen in the 22q11.2 microdeletion syndrome.[4][5]
Management
Several examinations should be performed to determine the level of disability and the needs of a person with the 1q21.1 deletion. Because of the vast range of symptoms, ophthalmologic, cardiac, developmental, neurological, and psychiatric evaluations may be helpful in identifying the manifestations of 1q21.1 deletion. Different developmental abnormalities of the skull and genitourinary system can be identified with brain imaging and renal ultrasounds. Consultations with a medical geneticist or genetic counselor can aid in determining the cause of 1q21.1 deletion. [4]
Because of the variability in 1q21.1 deletion, management is dependent on specific symptoms. Occupational and physical therapy, as well as special learning programs, may be used to manage symptoms.[4]
Epidemiology
Nonallelic homologous recombination (NAHR), mediated by low copy repeats (LCRs), is a well-known mechanism of copy number alterations in an array of genomic diseases. Individually, these conditions are uncommon, yet collectively, they impact a significant portion of the population. 1q21.1 deletions are estimated to occur in about 0.015% of the population. However, these CNVs are incompletely penetrant, therefore it is likely that the actual prevalence in the general population is greater than the current predictions.[5]
Research
On several locations in the world people are studying on the subject of 1q21.1 deletion syndrome. The syndrome was identified for the first time with people with heart abnormalities. The syndrome has later been found in patients with schizophrenia. Research is done on patients with a symptom of the syndrome, to find more patients with the syndrome.
There may be a relation between autism and schizophrenia. Literature shows that nine locations have been found on the DNA where the syndromes related to autism or schizophrenia can be found, the so-called "hotspots": 1q21.1, 3q29, 15q13.3, 16p11.2, 16p13.1, 16q21, 17p12, 21q11.2 and 21q13.3. With a number of hotspots, either autism and schizophrenia were observed depending on the copy-number variation (CNV) at that location.
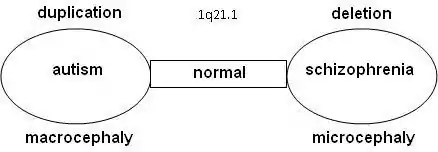
Statistical research showed that schizophrenia is more common in combination with 1q21.1 deletion syndrome. On the other side, autism is significantly more common with 1q21.1 duplication syndrome. Further research confirmed that the odds on a relation between schizophrenia and deletions at 1q21.1, 3q29, 15q13.3, 22q11.21 en Neurexin 1 (NRXN1) and duplications at 16p11.2 are at 7.5% or higher.[10][11]
Common variations in the BCL9 gene, which is in the distal area, confer risk of schizophrenia and may also be associated with bipolar disorder and major depressive disorder.[12]
Research is done on 10–12 genes on 1q21.1 that produce DUF1220-locations. DUF1220 is an unknown protein, which is active in the neurons of the brain near the neocortex. Based on research on apes and other mammals, it is assumed that DUF1220 is related to cognitive development (man: 212 locations; chimpanzee: 37 locations; monkey: 30 locations; mouse: 1 location). It appears that the DUF1220-locations on 1q21.1 are in areas that are related to the size and the development of the brain. The aspect of the size and development of the brain is related to autism (macrocephaly) and schizophrenia (microcephaly). It has been proposed that a deletion or duplication of a gene that produces DUF1220-areas might cause growth and development disorders in the brain[13]
Another relation between macrocephaly with duplications and microcephaly with deletions has been seen in research on the HYDIN Paralog or HYDIN2. This part of 1q21.1 is involved in the development of the brain. It is assumed to be a dosage-sensitive gene. When this gene is not available in the 1q21.1 area, it leads to microcephaly. HYDIN2 is a recent duplication (found only in humans) of the HYDIN gene found on 16q22.2.[14] Research on the genes CHD1L and PRKAB2 within lymphoblast cells[15] lead to the conclusion that anomalies appear with the 1q21.1-deletion syndrome:
- CHD1L is an enzyme which is involved in untangling the chromatides and the DNA repair system. With 1q21.1 deletion syndrome a disturbance occurs, which leads to increased DNA breaks. The role of CHD1L is similar to that of helicase with the Werner syndrome
- PRKAB2 is involved in maintaining the energy level of cells. With 1q21.1-deletion syndrome this function was attenuated.
GJA5 has been identified as the gene that is responsible for the phenotypes observed with congenital heart diseases on the 1q21.1 location. In case of a duplication of GJA5 tetralogy of Fallot is more common. In case of a deletion other congenital heart diseases than tetralogy of Fallot are more common.[16]
References
- "1q21.1 microdeletion: MedlinePlus Genetics". medlineplus.gov. Retrieved 2023-07-25.
- "1q21.1 microdeletion syndrome — About the Disease — Genetic and Rare Diseases Information Center". rarediseases.info.nih.gov. Retrieved 2023-07-25.
- "Overview: 1q21.1 microdeletion syndrome". Genetic and Rare Diseases Information Center (GARD). Office of Rare Diseases Research • U.S. National Institutes of Health. 8 August 2011. Archived from the original on 2 June 2013. Retrieved 9 September 2013.
- Haldeman-Englert, Chad R.; Jewett, Tamison (24 February 2011). 1q21.1 Recurrent Microdeletion. Seattle (WA): University of Washington, Seattle. PMID 21348049 – via PubMed.
- Gillentine, M.A.; Lupo, P.J.; Stankiewicz, P.; Schaaf, C.P. (July 2008). "An estimation of the prevalence of genomic disorders using chromosomal microarray data". Journal of Human Genetics. 63 (7): 795–801. doi:10.1038/s10038-018-0451-x. ISSN 1434-5161. PMC 6019170. PMID 29691480.
- Chen, Mei-Jou; Wei, Shin-Yi; Yang, Wei-Shiung; Wu, Tsai-Tzu; Li, Huei-Ying; Ho, Hong-Nerng; Yang, Yu-Shih; Chen, Pei-Lung (July 2015). "Concurrent exome-targeted next-generation sequencing and single nucleotide polymorphism array to identify the causative genetic aberrations of isolated Mayer-Rokitansky-Küster-Hauser syndrome". Human Reproduction (Oxford, England). 30 (7): 1732–1742. doi:10.1093/humrep/dev095. ISSN 1460-2350. PMID 25924657.
- Stefansson, Hreinn; Rujescu, Dan; Cichon, Sven; Pietiläinen, Olli P. H.; Ingason, Andres; Steinberg, Stacy; Fossdal, Ragnheidur; Sigurdsson, Engilbert; Sigmundsson, Thordur; Buizer-Voskamp, Jacobine E.; Hansen, Thomas; Jakobsen, Klaus D.; Muglia, Pierandrea; Francks, Clyde; Matthews, Paul M.; Gylfason, Arnaldur; Halldorsson, Bjarni V.; Gudbjartsson, Daniel; Thorgeirsson, Thorgeir E.; Sigurdsson, Asgeir; Jonasdottir, Adalbjorg; Jonasdottir, Aslaug; Bjornsson, Asgeir; Mattiasdottir, Sigurborg; Blondal, Thorarinn; Haraldsson, Magnus; Magnusdottir, Brynja B.; Giegling, Ina; Möller, Hans-Jürgen; Hartmann, Annette; Shianna, Kevin V.; Ge, Dongliang; Need, Anna C.; Crombie, Caroline; Fraser, Gillian; Walker, Nicholas; Lonnqvist, Jouko; Suvisaari, Jaana; Tuulio-Henriksson, Annamarie; Paunio, Tiina; Toulopoulou, Timi; Bramon, Elvira; Di Forti, Marta; Murray, Robin; Ruggeri, Mirella; Vassos, Evangelos; Tosato, Sarah; Walshe, Muriel; Li, Tao; Vasilescu, Catalina; Mühleisen, Thomas W.; Wang, August G.; Ullum, Henrik; Djurovic, Srdjan; Melle, Ingrid; Olesen, Jes; Kiemeney, Lambertus A.; Franke, Barbara; Sabatti, Chiara; Freimer, Nelson B.; Gulcher, Jeffrey R.; Thorsteinsdottir, Unnur; Kong, Augustine; Andreassen, Ole A.; Ophoff, Roel A.; Georgi, Alexander; Rietschel, Marcella; Werge, Thomas; Petursson, Hannes; Goldstein, David B.; Nöthen, Markus M.; Peltonen, Leena; Collier, David A.; St Clair, David; Stefansson, Kari (September 2008). "Large recurrent microdeletions associated with schizophrenia". Nature. 455 (7210): 232–236. Bibcode:2008Natur.455..232S. doi:10.1038/nature07229. hdl:11858/00-001M-0000-0012-C93F-6. ISSN 1476-4687. PMC 2687075. PMID 18668039.
- Mefford HC, Sharp AJ, Baker C, et al. (October 2008). "Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes". N. Engl. J. Med. 359 (16): 1685–99. doi:10.1056/NEJMoa0805384. hdl:2066/71235. PMC 2703742. PMID 18784092.
- A. Ploeger; 'Towards an integration of evolutionary psychology and developmental science: New insights from evolutionary developmental biology'
- Levinson DF, Duan J, Oh S, et al. (March 2011). "Copy number variants in schizophrenia: confirmation of five previous findings and new evidence for 3q29 microdeletions and VIPR2 duplications". Am J Psychiatry. 168 (3): 302–16. doi:10.1176/appi.ajp.2010.10060876. PMC 4441324. PMID 21285140.
- Ikeda M, Aleksic B, Kirov G, et al. (February 2010). "Copy number variation in schizophrenia in the Japanese population". Biol. Psychiatry. 67 (3): 283–6. doi:10.1016/j.biopsych.2009.08.034. PMID 19880096. S2CID 26047827.
- Li J, Zhou G, Ji W, et al. (March 2011). "Common variants in the BCL9 gene conferring risk of schizophrenia". Arch. Gen. Psychiatry. 68 (3): 232–40. doi:10.1001/archgenpsychiatry.2011.1. PMID 21383261.
- e.g.: Dumas L, Sikela JM (2009). "DUF1220 domains, cognitive disease, and human brain evolution". Cold Spring Harb. Symp. Quant. Biol. 74: 375–82. doi:10.1101/sqb.2009.74.025. PMC 2902282. PMID 19850849.
- Doggett NA, Xie G, Meincke LJ, et al. (Dec 2006). "A 360-kb interchromosomal duplication of the human HYDIN locus". Genomics. 88 (6): 762–71. doi:10.1016/j.ygeno.2006.07.012. PMID 16938426.
- Harvard C (2011). "Understanding the impact of 1q21.1 copy number variant". Orphanet Journal of Rare Diseases. 6: 54. doi:10.1186/1750-1172-6-54. PMC 3180300. PMID 21824431.
- Soemedi, R.; et al. (2011). "DPhenotype-Specific Effect of Chromosome 1q21.1 Rearrangements and GJA5 Duplications in 2436 Congenital Heart Disease Patients and 6760 Controls". Hum. Mol. Genet. 21 (7): 1513–1520. doi:10.1093/hmg/ddr589. PMC 3298277. PMID 22199024.
Further reading
- Genetics of Mental Retardation, Karger. Knight S. (ed) Chapter One: 'A Parent's' Perspective' contains description and photos of female with 1q21.1 microdeletion
- Brunet A, Armengol L, Heine D, et al. (2009). "BAC array CGH in patients with Velocardiofacial syndrome-like features reveals genomic aberrations on chromosome region 1q21.1". BMC Med. Genet. 10: 144. doi:10.1186/1471-2350-10-144. PMC 2805625. PMID 20030804.
- Brunetti-Pierri N, Berg JS, Scaglia F, Belmont J, Bacino CA, Sahoo T, et al. (2008). "Recurrent reciprocal 1q21.1 deletions and duplications associated with microcephaly or macrocephaly and developmental and behavioral abnormalities". Nat Genet. 40 (12): 1466–71. doi:10.1038/ng.279. PMC 2680128. PMID 19029900.
- Crespi B, Stead P, Elliot M (2010). "Evolution in health and medicine Sackler colloquium: Comparative genomics of autism and schizophrenia". Proc Natl Acad Sci U S A. 107 (Suppl 1): 1736–41. Bibcode:2010PNAS..107.1736C. doi:10.1073/pnas.0906080106. PMC 2868282. PMID 19955444.