Degradome sequencing
Degradome sequencing (Degradome-Seq), [1][2] also referred to as parallel analysis of RNA ends (PARE), is a modified version of 5'-Rapid Amplification of cDNA Ends (RACE) using high-throughput, deep sequencing methods such as Illumina's SBS technology. The degradome encompasses the entire set of proteases that are expressed at a specific time in a given biological material, including tissues, cells, organisms, and biofluids.[3] Thus, sequencing this degradome offers a method for studying and researching the process of RNA degradation. This process is used to identify and quantify RNA degradation products, or fragments, present in any given biological sample.[4] This approach allows for the systematic identification of targets of RNA decay and provides insight into the dynamics of transcriptional and post-transcriptional gene regulation.[4]
Degradome sequencing is a complex process which includes multiple steps such as isolating RNA fragments in a given sample as well as ligation and reverse transcription to form complementary DNA (cDNA) strands.[4] This cDNA can be sequenced, and the results are compared with a transcriptome, or reference genome, in order to determine and characterize the abundance of the RNA fragments identified in this process.[4]
Methods
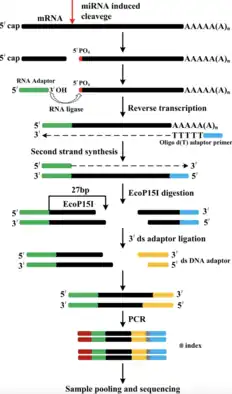
Technique
In general, the basic steps necessary for degradome sequencing include:
- RNA isolation and separation: The RNA is isolated from the sample and size-fractionated to separate and classify small RNA molecules.[5]
- Library sequencing: The RNA is then ligated and used to form cDNA strands via reverse transcription. The cDNA is then denatured and replicated via polymerase chain reaction (PCR). This yields a library generated and sequenced using high-throughput, deep sequencing methods.[5]
- Analysis of data: The sequences yielded from these procedures are processed using bioinformatics tools to remove low-quality results, sequences that are adapters, and other forms of non-viable results.[5] Following this screening step, the reads that remain are then compared with the reference genome (transcriptome) to determine the cleavage sites and their specific location.[5]
- Analysis of cleavage sites: The cleavage sites are analyzed to identify the target transcripts of the enzyme that degrades RNA.[5] Cleavage sites are identified in the 3' untranslated region (UTR) of mRNAs. In addition, sequence motifs and other targets involved in miRNA-mediated degradation can be identified via the analysis of cleavage sites.[5]
Analysis of Sequenced Raw Data
When analyzing the raw data derived from degradome sequencing, software tools like CleaveLand, PAREsnip, and miRferno are beneficial resources for researchers.[6]
CleaveLand Data Analysis Methodology
Degradome sequencing data and structural RNAs are used to remove all degradome sequences with exact matches to structural RNAs. The cDNA database is then used to map degradome sequences to cDNA sequences. The degradome sequences with many transcriptome hits are normalized. Then, query sequences of mRNA are generated for the matching degradome sequence. These query sequences are mapped to small RNAs, and a complementarity search is performed to match query sequences to small RNAs. A signal is then released to initiate noise analysis which works to distinguish and separate spurious results from real targets. Lastly, the resulting output of data analysis includes a list of all mRNA targets with the associated alignments for the small RNA-mRNA pairs.[7]
Applications
Introduction
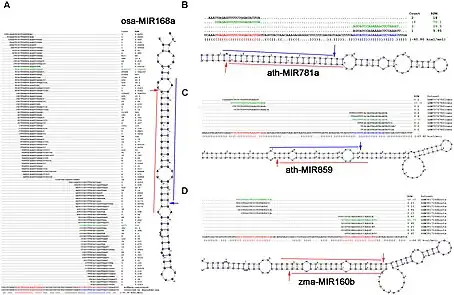
The applications of degradome sequencing include identifying microRNA (miRNA) targets, establishing mRNA methods of decay, and finding novel non-coding RNA fragments. In particular, this tool has been used to determine miRNA targets in numerous organisms, such as plants[8] and mammals.[9] Degradome sequencing has also been used to study the role of RNA decay pathways in cancer[9] and identify new types of non-coding RNAs.[8]
Ultimately, degradome sequencing is a powerful tool for the comprehensive analysis of RNA degradation with a variety of applications in biological research as well as medicine.
MicroRNA Research
MicroRNAs are a class of small noncoding RNA created by removing stem-loop precursors.[10] MiRNAs play a role in controlling gene expression post-transcriptionally in addition to during transcription via RNA silencing.[11] In order to accomplish this, the RNA-induced silencing complex (RISC) processes pre-microRNAs into mature microRNAs.[12] Mature miRNAs target specific mRNA species for regulation, often via the RISC complex disassembling specific mRNA sequences to inhibit translation.[12]
MiRNAs are highly conserved across a variety of species, so degradome sequencing is used in research to identify mRNA targets in many species.[13] Degradome sequencing has been used to identify miRNA cleavage sites,[13] because miRNAs can cause endonucleolytic cleavage of mRNA by extensive and often perfect complementarity to mRNAs.[1][2] Degradome sequencing revealed many known and novel plant miRNA and small interfering RNA (siRNA) targets.[1][2][14][15][16][17] Recently, degradome sequencing also has been applied to identify animal (human and mouse) miRNA-derived cleavages.[18][19][20]
Tracking microRNA Processing Signals by Degradome Sequencing Data Analysis
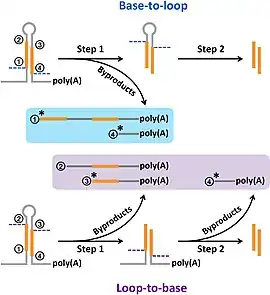
In this study, researchers tracked and reported miRNA processing intermediates. Degradome signals on miRNA precursors were extracted and processed for 15 different species. The use of degradome sequencing in this study allowed for the collection of data that supported the analysis and processing of many miRNA precursors, with a greater ratio of high-confidence miRNAs annotated in miRBase, an miRNA database, than those considered low-confidence. Additionally, this study highlighted the importance of degradome sequencing as a technique in the study of miRNA annotation. In particular, the processing signal distribution provided by degradome sequencing data allowed the researchers to propose a new model for the method by which miRNAs are diced and to determine the frequency with which the loop-to-base mode of processing occurred. Ultimately, the results of this study are indicative of the impressive capability of degradome sequencing data to track miRNA processing signals, providing novel insights into miRNA processing and function.[21]
The RNA Degradome: A Precious Resource for Deciphering RNA Processing and Regulation Codes in Plants
In this study, researchers developed a model in which biologists could use data derived from degradome sequencing to determine the effect of transcriptional and/or post-transcriptional regulation on patterns of gene expression in plants. In particular, this model applies degradome sequencing data to establish the method by which small RNAs (sRNAs) mature and guide the process of targeted gene regulation. The results of this study demonstrate the vast potential applications of degradome sequencing analysis in future research regarding RNA biology in eukaryotes. In particular, degradome sequencing data can be used to track non-coding RNA (ncRNA) processing signals which would be a valuable tool if expanded to include animal-based research.[8]
External links
- starBase database: a database for exploring microRNA cleavage sites from degradome sequencing (Degradome-Seq) data.
Cancer Research
Degradome sequencing can be used to identify cleavage sites of RNAs by sequencing the 5' end of the cleaved RNA fragments.[4] This technique has been widely used in cancer research to identify potential targets of RNA-degrading enzymes involved in cancer progression. As such, degradome sequencing has provided a new method of discovering markers for earlier diagnosis and prognosis determination in cancer patients. Given the established role of extracellular proteases in promoting tumor development and growth across different tissues, degradome sequencing also holds important implications for discovering novel therapeutic targets for cancer treatments.[22]
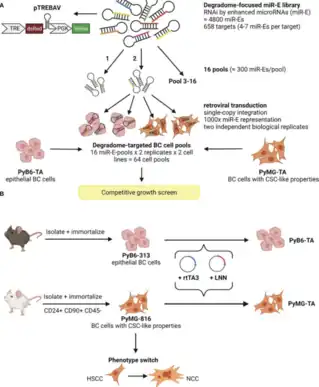
Degradome-Focused RNA Interference Screens to Identify Proteases Important for Breast Cancer Cell Growth
In this study, researchers utilized degradome sequencing to analyze all genome-encoded proteases involved in cell growth associated with breast cancer. These genetic screens were performed in two breast cancer cell lines in mice which were phenotypically distinct. One of these was a stem-cell like breast cancer cell line that altered its behavior under varied environmental conditions, such as the availability of oxygen and nutrients. Degradome sequencing, followed by a multistep selection process, revealed 100 protease genes that played a role in the growth of breast cancer cells. While the role of many of these protease genes in breast cancer growth was supported by previous research, this study found some proteases previously unknown to be involved in cancer growth. Additionally, this study revealed that environmental factors, such as nutrient and oxygen abundance, affect the extent to which breast cancer cells rely on specific proteases identified via degradome sequencing.[9]
The results of this study were validated by using individual knockdown constructs in mice which functionally diminished the proteases of interest and affected the expression of breast cancer cells. These results indicate the high degree of reliability of degradome sequencing in identifying proteases involved in the growth of breast cancer cell lines in mouse models. Ultimately, this study concluded that degradome sequencing is a beneficial research tool for discovering and analyzing the functions of proteases in the proliferation of breast cancer. This holds many important implications for the potential degradome sequencing possesses as a diagnostic tool in early breast cancer detection and treatment.[9]
References
- German MA, Pillay M, Jeong DH, Hetawal A, Luo S, Janardhanan P, Kannan V, Rymarquis LA, Nobuta K, German R, De Paoli E, Lu C, Schroth G, Meyers BC, Green PJ (2008). "Global identification of microRNA-target RNA pairs by parallel analysis of RNA ends". Nat. Biotechnol. 26 (8): 941–946. doi:10.1038/nbt1417. PMID 18542052. S2CID 13187064.
- Addo-Quaye C, Eshoo TW, Bartel DP, Axtell MJ (2008). "Endogenous siRNA and miRNA targets identified by sequencing of the Arabidopsis degradome". Curr. Biol. 18 (10): 758–762. doi:10.1016/j.cub.2008.04.042. PMC 2583427. PMID 18472421.
- Trindade, Fábio; Ferreira, Rita; Amado, Francisco; Vitorino, Rui (2015-01-01), Makowski, Gregory S. (ed.), "Chapter Five - Biofluid Proteases Profiling in Diabetes Mellitus", Advances in Clinical Chemistry, Elsevier, 69: 161–207, doi:10.1016/bs.acc.2014.12.004, PMID 25934362, retrieved 2023-04-17
- Lin, Shih-Shun; Chen, Yihua; Lu, Mei-Yeh Jade (2019). "Degradome Sequencing in Plants". Plant MicroRNAs. Methods in Molecular Biology. Vol. 1932. pp. 197–213. doi:10.1007/978-1-4939-9042-9_15. ISBN 978-1-4939-9041-2. ISSN 1940-6029. PMID 30701502. S2CID 73413403.
- Li, Yong-Fang; Zhao, Miao; Wang, Menglei; Guo, Junqiang; Wang, Li; Ji, Jie; Qiu, Zongbo; Zheng, Yun; Sunkar, Ramanjulu (2019-11-18). "An improved method of constructing degradome library suitable for sequencing using Illumina platform". Plant Methods. 15 (1): 134. doi:10.1186/s13007-019-0524-7. ISSN 1746-4811. PMC 6859640. PMID 31832076.
- Folkes, Leighton; Moxon, Simon; Woolfenden, Hugh C.; Stocks, Matthew B.; Szittya, Gyorgy; Dalmay, Tamas; Moulton, Vincent (2012-07-01). "PAREsnip: a tool for rapid genome-wide discovery of small RNA/target interactions evidenced through degradome sequencing". Nucleic Acids Research. 40 (13): e103. doi:10.1093/nar/gks277. ISSN 1362-4962. PMC 3401462. PMID 22467211.
- Addo-Quaye, Charles; Miller, Webb; Axtell, Michael J. (2009-01-01). "CleaveLand: a pipeline for using degradome data to find cleaved small RNA targets". Bioinformatics. 25 (1): 130–131. doi:10.1093/bioinformatics/btn604. ISSN 1367-4803. PMC 3202307. PMID 19017659.
- Ma, Xiaoxia; Yin, Xiaopu; Tang, Zhonghai; Ito, Hidetaka; Shao, Chaogang; Meng, Yijun; Xie, Tian (2020). "The RNA degradome: a precious resource for deciphering RNA processing and regulation codes in plants". RNA Biology. 17 (9): 1223–1227. doi:10.1080/15476286.2020.1757898. ISSN 1547-6286. PMC 7549650. PMID 32338184.
- Hölzen, Lena; Syré, Kerstin; Mitschke, Jan; Brummer, Tilman; Miething, Cornelius; Reinheckel, Thomas (2022-10-12). "Degradome-focused RNA interference screens to identify proteases important for breast cancer cell growth". Frontiers in Oncology. 12: 960109. doi:10.3389/fonc.2022.960109. ISSN 2234-943X. PMC 9598039. PMID 36313646.
- Cui, Huachun; Xie, Na; Tan, Zheng; Banerjee, Sami; Thannickal, Victor; Abraham, Edward; Liu, Gang (July 2014). "The human long noncoding RNA lnc-IL7R regulates the inflammatory response". European Journal of Immunology. 7 (44): 2084–2095. doi:10.1002/eji.201344126. PMC 4107034. PMID 24723426.
- Cao, Chunyu; Ding, Yifei; Kong, Xiangjun; Feng, Guangde; Xiang, Wei; Chen, Long; Yang, Fang; Zhang, Ke; Chu, Mingxing; Wang, Pingqing; Zhang, Baoyun (2018-04-01). "Reproductive role of miRNA in the hypothalamic-pituitary axis". Molecular and Cellular Neuroscience. 88: 130–137. doi:10.1016/j.mcn.2018.01.008. ISSN 1044-7431. PMID 29414103. S2CID 24050423.
- Redfern, Andrew D.; Colley, Shane M.; Beveridge, Dianne J.; Ikeda, Naoya; Epis, Michael R.; Li, Xia; Foulds, Charles E.; Stuart, Lisa M.; Barker, Andrew; Russell, Victoria J.; Ramsay, Kerry; Kobelke, Simon J.; Li, Xiaotao; Hatchell, Esme C.; Payne, Christine (2013-04-16). "RNA-induced silencing complex (RISC) Proteins PACT, TRBP, and Dicer are SRA binding nuclear receptor coregulators". Proceedings of the National Academy of Sciences. 110 (16): 6536–6541. Bibcode:2013PNAS..110.6536R. doi:10.1073/pnas.1301620110. ISSN 0027-8424. PMC 3631673. PMID 23550157.
- Thomson, DW; Bracken, CP; Goodall, GJ (2011-06-07). "Experimental strategies for microRNA target identification". Nucleic Acids Research. 39 (16): 6845–6853. doi:10.1093/nar/gkr330. PMC 3167600. PMID 21652644.
- Yang JH, Li JH, Shao P, Zhou H, Chen YQ, Qu LH (2011). "starBase: a database for exploring microRNA–mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data". Nucleic Acids Res. 39 (Database issue): 1–8. doi:10.1093/nar/gkq1056. PMC 3013664. PMID 21037263.
- Henderson, I. R.; Jacobsen, S. E. (2008). "Sequencing sliced ends reveals microRNA targets". Nature Biotechnology. 26 (8): 881–882. doi:10.1038/nbt0808-881. PMC 2989925. PMID 18688239.
- Wu, L.; Zhang, Q.; Zhou, H.; Ni, F.; Wu, X.; Qi, Y. (2009). "Rice MicroRNA Effector Complexes and Targets". The Plant Cell Online. 21 (11): 3421–35. doi:10.1105/tpc.109.070938. PMC 2798332. PMID 19903869.
- Pantaleo, V.; Szittya, G.; Moxon, S.; Miozzi, L.; Moulton, V.; Dalmay, T.; Burgyan, J. (2010). "Identification of grapevine microRNAs and their targets using high throughput sequencing and degradome analysis". The Plant Journal. 62 (6): 960–76. doi:10.1111/j.0960-7412.2010.04208.x. PMID 20230504.
- Shin, C.; Nam, J. W.; Farh, K. K. H.; Chiang, H. R.; Shkumatava, A.; Bartel, D. P. (2010). "Expanding the MicroRNA Targeting Code: Functional Sites with Centered Pairing". Molecular Cell. 38 (6): 789–802. doi:10.1016/j.molcel.2010.06.005. PMC 2942757. PMID 20620952.
- Karginov, F. V.; Cheloufi, S.; Chong, M. M. W.; Stark, A.; Smith, A. D.; Hannon, G. J. (2010). "Diverse Endonucleolytic Cleavage Sites in the Mammalian Transcriptome Depend upon MicroRNAs, Drosha, and Additional Nucleases". Molecular Cell. 38 (6): 781–8. doi:10.1016/j.molcel.2010.06.001. PMC 2914474. PMID 20620951.
- Bracken, CP; Szubert, JM; Mercer, TR; Dinger, ME; Thomson, DW; Mattick, JS; Michael, MZ; Goodall, GJ (2011-03-22). "Global analysis of the mammalian RNA degradome reveals widespread miRNA-dependent and miRNA-independent endonucleolytic cleavage". Nucleic Acids Research. 39 (13): 5658–5668. doi:10.1093/nar/gkr110. PMC 3141239. PMID 21427086.
- Yu, Dongliang; Xu, Min; Ito, Hidetaka; Shao, Weishan; Ma, Xiaoxia; Wang, Huizhong; Meng, Yijun (2018). "Tracking microRNA Processing Signals by Degradome Sequencing Data Analysis". Frontiers in Genetics. 9: 546. doi:10.3389/fgene.2018.00546. ISSN 1664-8021. PMC 6246748. PMID 30487815.
- Fan, Jia; Ning, Bo; Lyon, Christopher J.; Hu, Tony Y. (2017-01-01), Hu, Tony Y.; Tamanoi, Fuyuhiko (eds.), "Chapter One - Circulating Peptidome and Tumor-Resident Proteolysis", The Enzymes, Peptidomics of Cancer-Derived Enzyme Products, Academic Press, 42: 1–25, doi:10.1016/bs.enz.2017.08.001, PMID 29054266, retrieved 2023-04-17