Dokdonia
Dokdonia is a genus of bacteria in the family Flavobacteriaceae and phylum Bacteroidota.[1][3][2]
| Dokdonia | |
|---|---|
| Scientific classification | |
| Domain: | |
| Phylum: | |
| Order: | |
| Suborder: | |
| Family: | |
| Genus: | Dokdonia Yoon et al. 2005[1] |
| Type species | |
| Dokdonia donghaensis[1] | |
| Species | |
|
D. diaphoros[1] | |
| Synonyms[2] | |
| |
The genus Dokdonia was first described in 2005 by Yoon et al. near Liancourt Rocks in the Sea of Japan.[4] Dokdonia is named after Dokdo, the Korean name for the Liancourt Rocks which lies between Japan and South Korea.[5] Yoon et al. isolated the bacterium from seawater and identified the first species as Dokdonia donghaensis.[6]
There are 10 classified species (D. aurantiaca, D. diaphoros, D. donghaensis, D. eikasta, D. flava, D. genika, D. lutea, D. pacifica, D. ponticola, and D. sinensis) and many unclassified strains under the Dokdonia genus based on the NCBI taxonomy database.[7] The International Committee on Systematics of Prokaryotes (ICSP) currently recognizes nine groups of Dokdonia described to species level with D. ponticola considered not validly published.[8]
The general characteristics of Dokdonia species include gram-negative, non-motile, aerobic, catalase- and oxidase-positive, non-spore-forming rods or elongated rods. Species are usually considered relatively halophilic as they are cultivated optimally with 2% w/v sea salts (NaCl).[4][9]
Ecology and Significance
Dokdonia species have a relatively wide distribution in the water column. They have been isolated from surface seawater, marine sediment, and seaweed.[10][11][12][13][14] Dokdonia have been found across a wide range of marine environments around Korea, China, and Japan, but have also been found in Baltic and Mediterranean waters.[15][16] They tend to live a planktonic lifestyle, drifting in the water column, but they can occupy a wide range of ecological niches.[15]
Dokdonia cells are primarily heterotrophic and sustain off dissolved organic carbon in the water column.[17] It has also been shown that phototrophy can be induced in some strains under laboratory conditions, implying that bacteria in the genus Dokdonia are not obligate heterotrophs but are mixotrophic.[18] This shift in carbon source is induced by lower levels and quality of carbon source as well as lower light levels.[15] As a planktonic mixotrophic microbe, Dokdonia cells can provide a source of organic matter and carbon for higher trophic level organisms, contribute to the ocean's primary productivity, and also play an important role in transforming elements and nutrient cycling.[19]
Bacteria in the genus Dokdonia has been seen to congregate in biofilms with other bacterial species, collectively improving their resistance to bacterial predation and producing antimicrobial agents.[16]
Among the Flavobacteriaceae, Dokdonia have a high requirement for copper in order maintain regular growth and metabolism.[20] Most information known about the genus is from strains of the type species, Dokdonia donghanensis.[21] As research and cultivation Dokdonia spp. continues, insights into their diversity and adaptations contribute greatly to public knowledge of marine bacteria as a whole.
Described Species
Dokdonia donghaensis[4][22]
- Discovered by Yoon et al. in 2005 from seawater near Liancourt Rocks.
- For isolation of strain from seawater, the standard dilution plating technique was used and the isolate is cultured on marine agar at 25 °C.[4]
- Gram-negative, non-motile, non-spore-forming, slightly halophilic, rod or elongated rod-shaped with carotenoid pigments but no flexirubin-type pigments
- Cells are 1.5–25.0μm in length and 0.3–0.6μm in width. Form circular, smooth, yellow colonies with diameter of 1 – 2mm on marine agar after 3 days.
- Optimum growth occurs at 30 °C, pH 7.0–8.0 and 2% (w/v) NaCl; No anaerobic growth on marine agar and no growth on marine agar supplemented with nitrate
- Catalase-positive and oxidase-positive; DNA G+C content is 38 mol%
 Colony morphology of Dokdonia donghaensis strain MED134 on nutrient agar.
Colony morphology of Dokdonia donghaensis strain MED134 on nutrient agar.
Dokdonia aurantiaca[23]
- Discovered by Choi et al. in 2018 from seaweed sample, Zostera marina, collected from East China Sea, Republic of Korea
- For isolation of strain from seaweed sample, the seaweed sample with cleaned and immerse in saline. The saline supernatant is collected and inoculated onto marine agar for 5 days at 25 °C to culture the isolated strain.[11][23]
- Gram-negative, non-motile, aerobic, orange-coloured and rod-shaped with carotenoid pigments but no flexirubin-type pigments
- Cells are 0.68–0.76 µm in diameter and 1.76–3.04 µm in length. Form circular, convex, smooth, colonies with diameter of 1.5–2mm on marine agar after 3 days
- Optimum growth occurs with 4% (w/v) NaCl, at pH 7 and at 25 °C
- Catalase-positive and oxidase-negative; DNA G+C content is 38 mol%
Dokdonia diaphoros[24][10]
- Discovered by Khan et al. in 2006 from marine sediment at Kisarazu, Japan and was classified as Krokinobacter diaphorus. In 2012, Yoon et al. reclassified this species as Dokdonia diaphoros as the 16S rRNA gene sequence analysis has shown that the genera Dokdonia and Krokinobacter under the family Flavobacteriaceae are phylogenetically closely related[24]
- Gram-negative, aerobic rods with carotenoid pigments but no flexirubin-type pigments
- Cells are 0.5–0.7mm by 2.5–4.0mm. Form slightly convex and yellowish colonies
- Optimum growth occurs with 3% (w/v) NaCl at 20 °C
- Catalase-positive and oxidase-positive; DNA G+C content is 33 mol%
Dokdonia eikasta [10]
- Discovered by Khan et al. in 2006 from marine sediment of Kisarazu, Japan. It was initially classified as Krokinobacter eikastus. In 2012, Yoon et al. reclassified this species as Dokdonia diaphoros [24]
- Gram-negative, aerobic rods with carotenoid pigments but no flexirubin-type pigments
- Cells are 0.5–0.7mm by 2.5–4.0mm. Form slightly convex and yellowish colonies
- Optimum growth occurs with 3% (w/v) NaCl at 20 °C
- Catalase-positive and oxidase-positive; DNA G+C content is 38 mol%
Dokdonia flava [11]
- Discovered by Choi et al. from the seaweed Zostera marina from the Yellow Sea, Republic of Korea in 2018.
- Gram-negative, aerobic, non-motile, rod-shape cells with carotenoid pigments but no flexirubin-type pigments
- Cell size of 0.80–0.89 µm in diameter and 2.24–3.84 µm in length. Form circular, convex, smooth, yellowish colonies with 1–2mm in diameter on marine agar after 3 days.
- Optimum growth occurs with 4% (w/v) NaCl at pH = 7 and 25 °C
- Catalase-positive and oxidase-positive; DNA G+C content is 36 mol%.
Dokdonia genika [10]
- Discovered by Khan et al. in 2006 from the marine sediment at Odawara, Japan and described as Krokinobacter genikus. In 2012, Yoon et al. reclassified this species as Dokdonia genika.
- Gram-negative, aerobic rods with carotenoid pigments but no flexirubin-type pigments
- Cells are 0.5–0.7mm by 2.5–4.0mm. Form slightly convex and yellowish colonies
- Optimum growth occurs with 3% (w/v) NaCl at 20 °C
- Catalase-positive and oxidase-positive; DNA G+C content is 37-39 mol%
Dokdonia lutea [12]
- Discovered by Choi et al. in 2017 from the brown alga Sargassum fulvellum collected from the East China Sea, Republic of Korea.
- Gram-negative, aerobic, non-motile, rod-shaped with no flexirubin-type pigments
- Cells are 0.5 µm in diameter and 2.0–3.2 µm in length. Form circular, convex, smooth, and yellowish colonies that are 1.5–2 mm in diameter on marine agar after 3 days
- Optimum growth occurs with 5% (w/v) NaCl at pH = 8 and at 25-30 °C
- Catalase-positive and oxidase-negative; DNA G+C content is 35 mol%
Dokdonia pacifica [13]
- Discovered by Zhang et al. in 2015 from surface seawater collected from the South Pacific Gyre.
- Gram-negative, aerobic, non-flagellated, non-gliding, rod-shaped with carotenoid pigments but no flexirubin-type pigments
- Cells are 1.2–1.5mm in length, 0.3–0.4mm in width. Form circular, convex, transparent, and yellow transparent, and yellow colonies that are 0.8–1.0 mm in diameter on marine agar after 3 days
- Optimum growth occurs with 2-3% (w/v) NaCl at pH = 8 and at 28 °C
- Catalase-positive and oxidase-positive; DNA G+C content is 36 mol%
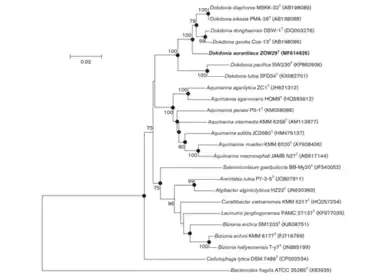 Phylogeny of Dokdonia based on 16S rRNA sequencing. Bootstrap values over 70% are shown, indicating confidence in the organization of the phylogenetic tree over 1000 replicates. Scale bar represents 0.02 nucleotide substitutions per position.
Phylogeny of Dokdonia based on 16S rRNA sequencing. Bootstrap values over 70% are shown, indicating confidence in the organization of the phylogenetic tree over 1000 replicates. Scale bar represents 0.02 nucleotide substitutions per position.
Dokdonia sinensis [14]
- Discovered by Zhou et al. in 2020 from seawater collected around Xiaoshi Island, PR China.
- Gram-negative, aerobic, non-motile, rod-shaped with no flexirubin-type pigments
- Cells are 1.0–3.0 µm in length and 0.5–0.8 µm in width. Form circular, convex, smooth, orange-pigmented colonies that are 1.0–1.5mm in diameter on marine agar after 3 days
- Optimum growth occurs with 3% (w/v) NaCl at pH = 7 and at 28 °C
- Catalase-positive and oxidase-negative; DNA G+C content is 39.5 mol%
Dokdonia ponticola[25]
- Discovered by Park et al. in 2018 from seawater collected around Oido, an island of South Koreaon in the Yellow Sea. This species is not recognized by ICSP as a species under Dokdonia genus. However, the result of 16S rRNA gene sequences showed that this strain has high similarity with the type strains of Dokdonia species, thus suggesting it belongs to Dokdonia genus.[25]
- Gram-negative, aerobic, non-motile, carrageenan-degrading, rod-shaped with no flexirubin-type pigments
- Cells are 0.2–0.5 µm in diameter and 0.5 to over 10.0 µm in length. Form circular, convex, smooth colonies with intense yellow colour. Colonies are 0.5–1.0mm in diameter on marine agar after 5 days
- Optimum growth occurs with 2% (w/v) NaCl at 20-25 °C.
- Catalase-positive and oxidase-positive; DNA G+C content is 39.7 mol%
Genome
Nine species of Dokdonia have been added to core genomic databases such as Uniprot and Genbank but not all have undergone formal review.[26]
Genome properties of D. donghaensis DSW-1T
The complete genome sequence of D. donghaensis DSW-1T can be accessed from GenBank under the accession number CP015125.[6] The complete circular genome contains 3,923,666 base pairs, 55 RNA genes, and 2,881 protein genes. The sequencing was established using the PacBio sequencing platform and funded by the National Research Foundation of Korea.[6]
Genome properties of Dokdonia sp. strain MED134
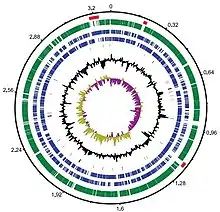
Whole-genome sequencing of Dokdonia sp. strain MED134 was done by J.Craig Venter Institute using Sanger method.[27] The genome size of Dokdonia sp. strain MED134 is 3,301,953 bp which is relatively small compared to other Bacteroidota. The number of conserved genes is similar to other Bacteroidota and it contains 170 core genes for bacteria.[27][28] Genome analysis shows that proteorhodopsins(PR)-containing marine Bacteroidota contains significantly fewer paralogous genes which form through intragenomic duplication events.[27][29] This result suggests that phototrophic Bacteroidota have similar number of gene families but less genes for each paralogous gene family which results in a reduced genome size. This finding is consistent among phototrophic bacteria, suggesting that Bacteroidota have evolved to retain only select genes from each paralogous gene family while also establishing other genetic features common among PR-containing marine Bacteroidota.[24][17]
Several key genes involved in PR-phototrophy have been identified through further genomic analysis of Dokdonia sp. strain MED134 and comparison to heterotrophic marine bacteria. Dokdonia have a relatively high abundance of peptidases and greater proportion of peptidases to total proteins compared to other bacteria. This property may indicate that the degradation of peptides may be the main carbon and nitrogen source instead of polysaccharides.[24] Phylogenetic analysis of features like conjugative transposons genes highlight horizontal gene transfer events with other marine flavobacteria.[27] Successful incorporation of PR genes into the genome has been seen to contribute to the overall fitness and survival of bacteria in the marine environment.[24]
Metabolism
Metabolic processes of Dokdonia
Dokdonia species are known to be chemoheterotrophs,[30][17] acquiring energy from organic molecules. They show maximal growth in conditions with a high concentration of dissolved organic matter.[17] Dokdonia and other members of the Bacteroidota phylum are important in the degradation of organic materials, especially during sporadic nutrient increase. They have a mechanism that allows them to attach and degrade polymeric substances efficiently.[27]
Some strains of Dokdonia are facultative double mixotrophs as they can utilize both heterotrophic and phototrophic metabolism.[18] These strains contain proteorhodopsins (PRs) which act as light-dependent proton pumps.[17][18][27] PR has a simple structure compared to other light-harvesting molecules like chlorophyll. They are a single membrane protein with retinal as its prosthetic group.[27][31] Retinal is a polyene chromophore (light-sensitive pigment) which acts as the light-absorbing molecule in Dokdonia species.[27][32] PRs allow cells to harvest energy from sunlight. When exposing to light, PRs pump protons across membrane and build proton gradient that can be used to generate ATP that powers various cellular activities.[27][18][33] The energy generated by PRs can support cell growth, degradation of complex and recalcitrant organic matter, and allow cells to uptake amino acids and peptides at lower concentrations.[27]
Dokdonia sp. strain MED134 (Dokdonia donghaensis MED134)
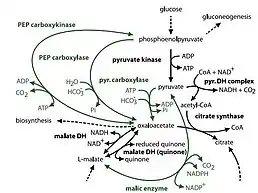
This strain uses aerobic heterotrophic metabolism; it primarily uses amino acids as carbon and nitrogen sources via expression of peptidases which break peptides down into amino acids. Dokdonia sp. strain MED134 has a Na+ ion pump on its membrane that can generate Na+ gradient to produce ATP for other cellular activities.[27]
Dokdonia sp. strain MED134 has complete Embden-Meyerhof-Farnas pathway (glycolysis), gluconeogenesis pathway, and tricarboxylic acid (TCA) cycle. The anaplerotic reaction which connects glycolysis with TCA cycle utilizes several enzymes including PEP carboxykinase and PEP carboxylase.[27]
The PRs in Dokdonia sp. strain MED134 are predicted to be heptahelical integral membrane proteins that pump H+ across membranes to build a proton gradient which generates ATP.[17][18] Depends on the organism's depth in water column, the PR's maximum absorbance wavelength changes. In near surface waters, the maximum absorption wavelength of PR is around 530 nm (green light). In deeper water, the maximum absorption wavelength is 490 nm (blue light).[17]
Dokdonia sp. PRO95
Dokdonia sp. PRO95 carries two types of rhodopsins: PR as well as a rhodopsin that is related to xanthorhodopsins (XRs). XRs are light-harvesting proton pumps that are more efficient compared to PRs.[34] The PRs of this strain are light-driven sodium-motive pumps (Na+-rhodopsins or NaRs) which pump Na+ from the cytoplasm to external medium. They can also pump H+ when Na+ is absent.[31] Unlike Dokdonia sp. strain MED134, no light-induced growth is observed in Dokdonia sp. PRO95 despite they have high genome similarity.[34]
Ecological significance of proteorhodopsin-containing Dokdonia
The presence of PRs allows Dokdonia to grow better in light compared dark conditions, especially when there is low or intermediate levels of nutrients available.[17][18] Cells with PRs are less dependent on the amount and quality of organic carbon sources.[17][27] Being able to harvest energy from light gives them an advantage during nutrient deficient periods, making the retention of PRs favoured by selection.[17]
Alpha- and gammaproteobacteria that contain PRs have enhanced cell growth due to cellular functions such as survival in environments with deficient nutrients that are promoted by light .[35][36] The correlation between PR phototrophy and increased cell growth was first observed in Dokdonia sp. strain MED134.[17][27]
Expression of proteorhodopsin encoding gene
In Dokdonia, light induces the expression of PR genes.[17][18] There is a significant increase in the expression levels of PR genes in light compared to dark conditions as light can increase the strength of the PR gene promoter.[27][18] The light-dark cycle can induce the upregulation of PR genes in Flavobacteriia that lead to population growth.[27]
Behavior
Biofilm
In the Tyrrhenian Sea off the coast of Naples, species of Dokdonia are the most abundant in biofilms on plastic debris. 4.76 ± 7.1% of genera forming biofilms belong to Dokdonia.[37]
Given that Dokdonia was found on plastics but not in the sediment or water column, it is possible that the habitat in biofilms on plastics is preferable to planktonic growth for species of Dokdonia in the Mediterranean Sea.[37] As microbial mortality is reduced in the more stable microenvironment of a biofilm, population maintenance is less dependent on environmental factors such as nutrient concentration and grazers.[38]
Response to varying nutrient availability
Due to variable expression of PR, Dokdonia sp. strain MED134 growth rate is more positively influenced by light exposure when metabolizing an energy-poor carbon source such as alanine compared to that which was observed in the presence of glucose.[39][40]
Growth rates of Dokdonia sp. Dokd-P16 are significantly affected by variations in dissolved copper concentration. The strain exhibited an 80% reduction in growth rates in 0.6 nM copper compared to those observed in 50 nM copper. This suggests that the presence of copper is crucial to Dokdonia metabolism.[41]
Iron is a common limiting nutrient in the ocean. Dokdonia sp. strain MED134 has various mechanisms make use of different forms of iron. It contains ATP-binding cassette-type transport system that can transport iron into the cell. Also, it can store iron and recycle iron from heme.[17]
References
- Parte AC. "Dokdonia". LPSN.
- "Dokdonia". www.uniprot.org.
- Parker CT, Garrity GM (2012). Parker CT, Garrity GM (eds.). "Nomenclature Abstract for Dokdonia Yoon et al. 2005 emend. Yoon et al. 2012". The NamesforLife Abstracts. doi:10.1601/nm.9754.
- Yoon JH, Kang SJ, Lee CH, Oh TK (November 2005). "Dokdonia donghaensis gen. nov., sp. nov., isolated from sea water". International Journal of Systematic and Evolutionary Microbiology. 55 (Pt 6): 2323–2328. doi:10.1099/ijs.0.63817-0. PMID 16280490.
- Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB, eds. (2011). Bergey's Manual of Systematic Bacteriology: Volume 4: The Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes. Springer Science & Business Media. ISBN 9780387685724.
- Kim K, Kwon SK, Yoon JH, Kim JF (August 2016). "Complete Genome Sequence of the Proteorhodopsin-Containing Marine Flavobacterium Dokdonia donghaensis DSW-1T, Isolated from Seawater off Dokdo in the East Sea (Sea of Korea)". Genome Announcements. 4 (4): e00804–16. doi:10.1128/genomeA.00804-16. PMC 4974333. PMID 27491981.
- taxonomy. "Taxonomy browser (Dokdonia)". www.ncbi.nlm.nih.gov. Retrieved 2022-04-03.
- "Genus: Dokdonia". lpsn.dsmz.de. Retrieved 2022-02-20.
- Yoon JH, Kang SJ, Park S, Oh TK (August 2012). "Reclassification of the three species of the genus Krokinobacter into the genus Dokdonia as Dokdonia genika comb. nov., Dokdonia diaphoros comb. nov. and Dokdonia eikasta comb. nov., and emended description of the genus Dokdonia Yoon et al. 2005". International Journal of Systematic and Evolutionary Microbiology. 62 (Pt 8): 1896–1901. doi:10.1099/ijs.0.035253-0. PMID 21984677.
- Khan ST, Nakagawa Y, Harayama S (February 2006). "Krokinobacter gen. nov., with three novel species, in the family Flavobacteriaceae". International Journal of Systematic and Evolutionary Microbiology. 56 (Pt 2): 323–328. doi:10.1099/ijs.0.63841-0. PMID 16449433.
- Choi S, Kang JW, Yoon JH, Seong CN (March 2018). "Dokdonia flava sp. nov., isolated from the seaweed Zostera marina". International Journal of Systematic and Evolutionary Microbiology. 68 (3): 899–904. doi:10.1099/ijsem.0.002607. PMID 29458481.
- Choi S, Kang JW, Lee JH, Seong CN (November 2017). "Dokdonia lutea sp. nov., isolated from Sargassum fulvellum seaweed". International Journal of Systematic and Evolutionary Microbiology. 67 (11): 4482–4486. doi:10.1099/ijsem.0.002317. PMID 28933321.
- Zhang Z, Gao X, Wang L, Zhang XH (July 2015). "Dokdonia pacifica sp. nov., isolated from seawater". International Journal of Systematic and Evolutionary Microbiology. 65 (7): 2222–2226. doi:10.1099/ijs.0.000252. PMID 25862384.
- Zhou LY, Meng X, Zhong YL, Li GY, Du ZJ, Mu DS (March 2020). "Dokdonia sinensis sp. nov., a flavobacterium isolated from surface seawater". International Journal of Systematic and Evolutionary Microbiology. 70 (3): 1617–1622. doi:10.1099/ijsem.0.003949. PMID 32228747. S2CID 213159033.
- Israelsson, Stina (2020-06-05). "Seasonal dynamics of Baltic Sea plankton activities: heterotrophic bacterial function under different biological and environmental conditions".
- Burmølle, Mette; Webb, Jeremy S.; Rao, Dhana; Hansen, Lars H.; Sørensen, Søren J.; Kjelleberg, Staffan (June 2006). "Enhanced Biofilm Formation and Increased Resistance to Antimicrobial Agents and Bacterial Invasion Are Caused by Synergistic Interactions in Multispecies Biofilms". Applied and Environmental Microbiology. 72 (6): 3916–3923. Bibcode:2006ApEnM..72.3916B. doi:10.1128/aem.03022-05. ISSN 0099-2240. PMC 1489630. PMID 16751497.
- Gómez-Consarnau L, González JM, Coll-Lladó M, Gourdon P, Pascher T, Neutze R, et al. (January 2007). "Light stimulates growth of proteorhodopsin-containing marine Flavobacteria". Nature. 445 (7124): 210–213. Bibcode:2007Natur.445..210G. doi:10.1038/nature05381. PMID 17215843. S2CID 2338098.
- Palovaara J, Akram N, Baltar F, Bunse C, Forsberg J, Pedrós-Alió C, et al. (September 2014). "Stimulation of growth by proteorhodopsin phototrophy involves regulation of central metabolic pathways in marine planktonic bacteria". Proceedings of the National Academy of Sciences of the United States of America. 111 (35): E3650–E3658. Bibcode:2014PNAS..111E3650P. doi:10.1073/pnas.1402617111. PMC 4156726. PMID 25136122.
- Lombard, Fabien; Boss, Emmanuel; Waite, Anya M.; Vogt, Meike; Uitz, Julia; Stemmann, Lars; Sosik, Heidi M.; Schulz, Jan; Romagnan, Jean-Baptiste; Picheral, Marc; Pearlman, Jay (2019). "Globally Consistent Quantitative Observations of Planktonic Ecosystems". Frontiers in Marine Science. 6. doi:10.3389/fmars.2019.00196. ISSN 2296-7745.
- Posacka, A. M.; Maldonado, M. T. (2016-02-01). "Responses of Diverse Marine Heterotrophic Bacteria to Changing Copper Availability". American Geophysical Union. 2016: CT44B–0243. Bibcode:2016AGUOSCT44B0243P.
- Parker, Charles Thomas; Wigley, Sarah; Garrity, George M.; Taylor, Dorothea (2008). Parker, Charles Thomas; Garrity, George M (eds.). "Strain DSM 17200 (=KCTC 12391 =DSW-1) : Microbial Characteristics and Genomic Information". doi:10.1601/ex.9753.
{{cite journal}}: Cite journal requires|journal=(help) - Parker, Charles Thomas; Wigley, Sarah; Garrity, George M. (2009). Parker, Charles Thomas; Garrity, George M (eds.). "Taxonomy of the genus Dokdonia Yoon et al. 2005 emend. Yoon et al. 2012". NamesForLife. doi:10.1601/tx.9754.
- Choi S, Kang JW, Kim MS, Yoon JH, Seong CN (May 2018). "Dokdonia aurantiaca sp. nov., isolated from seaweed Zostera marina". International Journal of Systematic and Evolutionary Microbiology. 68 (5): 1697–1701. doi:10.1099/ijsem.0.002730. PMID 29570445.
- Yoon JH, Kang SJ, Park S, Oh TK (August 2012). "Reclassification of the three species of the genus Krokinobacter into the genus Dokdonia as Dokdonia genika comb. nov., Dokdonia diaphoros comb. nov. and Dokdonia eikasta comb. nov., and emended description of the genus Dokdonia Yoon et al. 2005". International Journal of Systematic and Evolutionary Microbiology. 62 (Pt 8): 1896–1901. doi:10.1099/ijs.0.035253-0. PMID 21984677.
- Park, Sooyeon; Won, Sung-Min; Yoon, Jung-Hoon (2018-09-01). "Dokdonia ponticola sp. nov., A Carrageenan-Degrading Bacterium of the Family Flavobacteriaceae Isolated from Seawater". Current Microbiology. 75 (9): 1126–1132. doi:10.1007/s00284-018-1496-y. ISSN 1432-0991. PMID 29761217. S2CID 46886165.
- Schoch CL, Ciufo S, Domrachev M, Hotton CL, Kannan S, Khovanskaya R, et al. (January 2020). "NCBI Taxonomy: a comprehensive update on curation, resources and tools". Database. 2020: baaa062. doi:10.1093/database/baaa062. PMC 7408187. PMID 32761142.
- González JM, Pinhassi J, Fernández-Gómez B, Coll-Lladó M, González-Velázquez M, Puigbò P, et al. (December 2011). "Genomics of the proteorhodopsin-containing marine flavobacterium Dokdonia sp. strain MED134". Applied and Environmental Microbiology. 77 (24): 8676–8686. Bibcode:2011ApEnM..77.8676G. doi:10.1128/AEM.06152-11. PMC 3233072. PMID 22003006.
- Bratlie, Marit S; Johansen, Jostein; Drabløs, Finn (2010-01-28). "Relationship between operon preference and functional properties of persistent genes in bacterial genomes". BMC Genomics. 11 (1): 71. doi:10.1186/1471-2164-11-71. ISSN 1471-2164. PMC 2837039. PMID 20109203.
- Dufayard, J.-F.; Duret, L.; Penel, S.; Gouy, M.; Rechenmann, F.; Perriere, G. (2005-02-15). "Tree pattern matching in phylogenetic trees: automatic search for orthologs or paralogs in homologous gene sequence databases". Bioinformatics. 21 (11): 2596–2603. doi:10.1093/bioinformatics/bti325. ISSN 1367-4803. PMID 15713731.
- Posacka AM, Semeniuk DM, Maldonado MT (2019-02-05). "Effects of Copper Availability on the Physiology of Marine Heterotrophic Bacteria". Frontiers in Marine Science. 5: 523. doi:10.3389/fmars.2018.00523. ISSN 2296-7745.
- Bogachev AV, Bertsova YV, Verkhovskaya ML, Mamedov MD, Skulachev VP (February 2016). "Real-time kinetics of electrogenic Na(+) transport by rhodopsin from the marine flavobacterium Dokdonia sp. PRO95". Scientific Reports. 6 (1): 21397. Bibcode:2016NatSR...621397B. doi:10.1038/srep21397. PMC 4749991. PMID 26864904.
- Spudich, John L.; Jung, Kwang-Hwan (2005-09-26), "Microbial Rhodopsins: Phylogenetic and Functional Diversity", Handbook of Photosensory Receptors, Weinheim, FRG: Wiley-VCH Verlag GmbH & Co. KGaA, pp. 1–23, doi:10.1002/352760510x.ch1, ISBN 9783527605101, retrieved 2022-04-03
- DeLong, Edward F.; Béjà, Oded (2010-04-27). "The Light-Driven Proton Pump Proteorhodopsin Enhances Bacterial Survival during Tough Times". PLOS Biology. 8 (4): e1000359. doi:10.1371/journal.pbio.1000359. ISSN 1545-7885. PMC 2860490. PMID 20436957.
- Riedel, Thomas; Gómez-Consarnau, Laura; Tomasch, Jürgen; Martin, Madeleine; Jarek, Michael; González, José M.; Spring, Stefan; Rohlfs, Meike; Brinkhoff, Thorsten; Cypionka, Heribert; Göker, Markus (2013-03-04). "Genomics and Physiology of a Marine Flavobacterium Encoding a Proteorhodopsin and a Xanthorhodopsin-Like Protein". PLOS ONE. 8 (3): e57487. Bibcode:2013PLoSO...857487R. doi:10.1371/journal.pone.0057487. ISSN 1932-6203. PMC 3587595. PMID 23526944.
- Gómez-Consarnau, Laura; Akram, Neelam; Lindell, Kristoffer; Pedersen, Anders; Neutze, Richard; Milton, Debra L.; González, José M.; Pinhassi, Jarone (2010-04-27). "Proteorhodopsin Phototrophy Promotes Survival of Marine Bacteria during Starvation". PLOS Biology. 8 (4): e1000358. doi:10.1371/journal.pbio.1000358. ISSN 1545-7885. PMC 2860489. PMID 20436956.
- Steindler, Laura; Schwalbach, Michael S.; Smith, Daniel P.; Chan, Francis; Giovannoni, Stephen J. (2011-05-09). "Energy Starved Candidatus Pelagibacter Ubique Substitutes Light-Mediated ATP Production for Endogenous Carbon Respiration". PLOS ONE. 6 (5): e19725. Bibcode:2011PLoSO...619725S. doi:10.1371/journal.pone.0019725. ISSN 1932-6203. PMC 3090418. PMID 21573025.
- Basili M, Quero GM, Giovannelli D, Manini E, Vignaroli C, Avio CG, et al. (2020-05-07). "Major Role of Surrounding Environment in Shaping Biofilm Community Composition on Marine Plastic Debris". Frontiers in Marine Science. 7: 262. doi:10.3389/fmars.2020.00262. ISSN 2296-7745.
- Burmølle M, Webb JS, Rao D, Hansen LH, Sørensen SJ, Kjelleberg S (June 2006). "Enhanced biofilm formation and increased resistance to antimicrobial agents and bacterial invasion are caused by synergistic interactions in multispecies biofilms". Applied and Environmental Microbiology. 72 (6): 3916–3923. Bibcode:2006ApEnM..72.3916B. doi:10.1128/AEM.03022-05. PMC 1489630. PMID 16751497.
- Koedooder, Coco; VanGeersdaële, Rémy; Guéneuguès, Audrey; Bouget, François-Yves; Obernosterer, Ingrid; Blain, Stéphane (2020-07-01). "The interplay between iron limitation, light and carbon in the proteorhodopsin-containing Photobacterium angustum S14". FEMS Microbiology Ecology. 96 (7): fiaa103. doi:10.1093/femsec/fiaa103. ISSN 0168-6496. PMID 32459302.
- Palovaara, Joakim; Akram, Neelam; Baltar, Federico; Bunse, Carina; Forsberg, Jeremy; Pedrós-Alió, Carlos; González, José M.; Pinhassi, Jarone (2014-08-18). "Stimulation of growth by proteorhodopsin phototrophy involves regulation of central metabolic pathways in marine planktonic bacteria". Proceedings of the National Academy of Sciences. 111 (35): E3650-8. Bibcode:2014PNAS..111E3650P. doi:10.1073/pnas.1402617111. ISSN 0027-8424. PMC 4156726. PMID 25136122.
- Posacka, Anna M.; Semeniuk, David M.; Maldonado, Maria T. (2019). "Effects of Copper Availability on the Physiology of Marine Heterotrophic Bacteria". Frontiers in Marine Science. 5. doi:10.3389/fmars.2018.00523. ISSN 2296-7745.
Further reading
- Anashkin, Viktor A.; Bertsova, Yulia V.; Mamedov, Adalyat M.; Mamedov, Mahir D.; Arutyunyan, Alexander M.; Baykov, Alexander A.; Bogachev, Alexander V. (2017-10-05). "Engineering a carotenoid-binding site in Dokdonia sp. PRO95 Na+-translocating rhodopsin by a single amino acid substitution". Photosynthesis Research. 136 (2): 161–169. doi:10.1007/s11120-017-0453-0. ISSN 0166-8595.
- Bunse, Carina; Israelsson, Stina; Baltar, Federico; Bertos-Fortis, Mireia; Fridolfsson, Emil; Legrand, Catherine; Lindehoff, Elin; Lindh, Markus V.; Martínez-García, Sandra; Pinhassi, Jarone (2019-01-17). "High Frequency Multi-Year Variability in Baltic Sea Microbial Plankton Stocks and Activities". Frontiers in Microbiology. 9. doi:10.3389/fmicb.2018.03296. ISSN 1664-302X.