T7 phage
Bacteriophage T7 (or the T7 phage) is a bacteriophage, a virus that infects bacteria. It infects most strains of Escherichia coli and relies on these hosts to propagate. Bacteriophage T7 has a lytic life cycle, meaning that it destroys the cell it infects. It also possesses several properties that make it an ideal phage for experimentation: its purification and concentration have produced consistent values in chemical analyses;[2] it can be rendered noninfectious by exposure to UV light;[3] and it can be used in phage display to clone RNA binding proteins.[3]
| Escherichia virus T7 | |
|---|---|
 | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Duplodnaviria |
| Kingdom: | Heunggongvirae |
| Phylum: | Uroviricota |
| Class: | Caudoviricetes |
| Order: | Caudovirales |
| Family: | Autographiviridae |
| Genus: | Teseptimavirus |
| Species: | Escherichia virus T7 |
Discovery
In a 1945 study by Demerec and Fano,[4] T7 was used to describe one of the seven phage types (T1 to T7) that grow lytically on Escherichia coli; [5] although all seven phages were numbered arbitrarily, phages with odd numbers, or T-odd phages, were later discovered to share morphological and biochemical features that distinguish them from T-even phages.[6] Before being physically referred to as T7, the phage was used in prior experiments. German-American biophysicist Max Delbrück worked with the same virus in the late 1930s, calling it phage δ, and French-Canadian microbiologist Félix d'Herelle likely studied its close relative in the 1920s.[7][5]
Hosts
T7 grows on rough strains of Escherichia coli (i.e. those without full-length O-antigen polysaccharide on their surface) and some other enteric bacteria, but close relatives also infect smooth and even capsulated strains.[8] E. coli is more resistant to T7 than to some other similar phages.
Virion structure

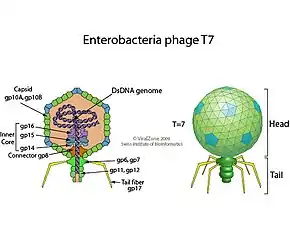
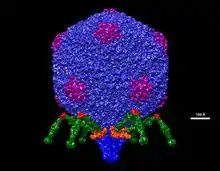
The virus has complex structural symmetry, with a capsid of the phage that is icosahedral (twenty faces) with an inner diameter of 55 nm and a tail 19 nm in diameter and 28.5 nm long attached to the capsid.[9] The ejection of proteins from the capsid upon infection causes the virus to change structure when it enters the cell.[10]
Genome
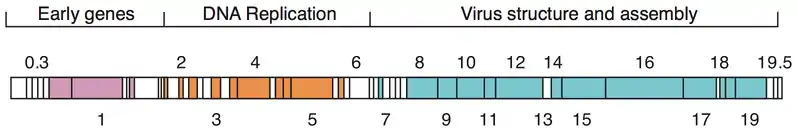
The genome of phage T7[11] was among the first completely sequenced genomes and was published in 1983.[12] The head of the phage particle contains the roughly 40 kbp dsDNA genome which encodes 55 proteins.[13] The genome features numerous overlapping genes [14] that were partially removed through 'refactoring' the genome to produce T7.1.[15]
Life cycle
T7 has a life cycle of 17 min at 37˚C, i.e. the time from infection to the lysis of the host cell when new phage are released. Due to the short latent period, most physiological studies are conducted at 30˚C where infected cells lyse after 30 min. However, high-fitness strains of T7 have been isolated with a latent period of only ~11 min at 37˚C growing under optimal conditions in rich media results. This adapted phage can undergo an effective expansion of its population by more than 1013 in one hour of growth.[17]
Infection of host bacteria
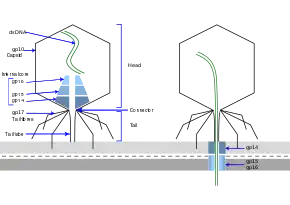

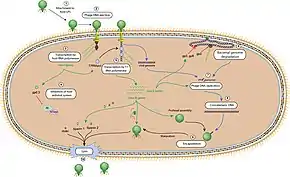
The T7 phage recognizes certain receptors on the surface of E.coli cells, and binds to the cell surface by its viral tail fibers. In some strains of T7, the tail fibers are replaced with tail-spikes that degrade the O- or K-antigens on the cell surface by way of enzymatic activity.
The adsorbtion and penetration process use lysozymes to create an opening within the peptidoglycan layer of the bacterial cell wall, allowing transfer of the viral DNA into the bacterium. The short, stubby tail of the T7-like phage is too short to span the cell envelope and, in order to eject the phage genome into the cell at the initiation of infection, virion proteins must first make a channel from the tip of the tail into the cell cytoplasm.[18] The phage also releases five proteins needed to begin replication of the viral genome and cleave the host genome.[19] T7 bacteriophage has been evolved to override several of the host bacteria's defenses including the peptidoglycan cell wall and the CRISPR system.[19] Once the T7 phage has inserted the viral genome, the process of DNA replication of the host genome is halted and replication of viral genome begins.
Under optimal conditions, the T7 phage can complete the lytic process within 25 minutes, leading to the death of the E. coli host cell. At the time of lysis, the virus can produce over 100 progeny.[19]
Components
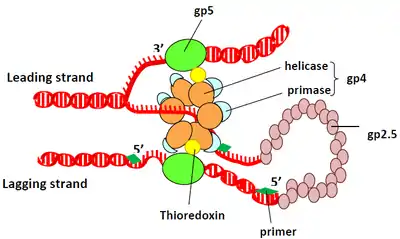
Gp5 (encoded by gene gp5) is T7 phage's DNA polymerase. T7 polymerase uses E. coli's endogenous thioredoxin, a REDOX protein, as a sliding DNA clamp during phage DNA replication (though thioredoxin normally has a different function). The sliding clamp functions to hold the polymerase onto the DNA, which increases the rate of synthesis.[20]
DNA replication and repair
Phage T7 has the simplest known DNA replisome, consisting of a helicase and primase that reside in a single polypeptide chain that forms a hexamer in the presence of DNA and ATP or dTTP. T7 DNA polymerase, assisted by E. coli thioredoxin, performs both leading and lagging-strand DNA synthesis.
In phage T7, DNA double-strand breaks are likely repaired by insertion of a patch of donor DNA into a gap at the break site.[21] This repair of double-strand breaks is facilitated by the gene 2.5 protein that promotes the annealing of homologous complementary strands of DNA.[22]
Replicative intermediates
The replicating intracellular DNA of phage T7, when stretched out after cell lysis, is usually longer than the mature phage chromosome (11 to 15 µM) and can occur in the form of highly concatenated linear strands up to 66 times the length of the mature phage chromosome.[23] The replicating DNA can also be seen in the form of coiled ring structures that appear to correspond to multiply looped DNA configurations in which superhelical twists, necessary for compaction of the DNA, were relieved by strand nicking upon cell lysis.
Applications in molecular biology
The T7 promoter sequence is used extensively in molecular biology due to its extremely high affinity for T7 RNA polymerase and thus high level of expression.[3][2]
T7 has been used as a model in synthetic biology. Chan et al. (2005) "refactored" the genome of T7, replacing approximately 12 kbp of its genome with engineered DNA.[15] The engineered DNA was designed to be easier to work with in a number of ways: individual functional elements were separated by restriction endonuclease sites for simple modification, and overlapping protein coding domains were separated and, where necessary, modified by single base pair silent mutations. T7 has been tested on human osteosarcoma to treat tumor cells.
References
- Dr. Victor Padilla-Sanchez, PhD (2021-07-10), Bacteriophage T7 Structural Model at Atomic Resolution., doi:10.5281/zenodo.5133295, retrieved 2021-07-24
- Studier, F. William (1969-11-01). "The genetics and physiology of bacteriophage T7". Virology. 39 (3): 562–574. doi:10.1016/0042-6822(69)90104-4. ISSN 0042-6822. PMID 4902069.
- Teesalu, Tambet; Sugahara, Kazuki N.; Ruoslahti, Erkki (2012-01-01), "Chapter two - Mapping of Vascular ZIP Codes by Phage Display", in Wittrup, K. Dane; Verdine, Gregory L. (eds.), Protein Engineering for Therapeutics, Part B, Methods in Enzymology, vol. 503, Academic Press, pp. 35–56, doi:10.1016/B978-0-12-396962-0.00002-1, PMID 22230564, retrieved 2019-11-18
- Demerec M, Fano U (March 1945). "Bacteriophage-Resistant Mutants in Escherichia Coli". Genetics. 30 (2): 119–36. doi:10.1093/genetics/30.2.119. PMC 1209279. PMID 17247150.
- Lobocka M, Szybalski, eds. (2012-12-31). Bacteriophages. Academic Press. pp. 226–. ISBN 978-0-12-394788-8.
- Cammack, Richard; Atwood, Teresa; Campbell, Peter; Parish, Howard; Smith, Anthony; Vella, Frank; Stirling, John, eds. (2006). Oxford Dictionary of Biochemistry and Molecular Biology. Oxford University Press. doi:10.1093/acref/9780198529170.001.0001. ISBN 9780191727641.
- d’Herelle, F. (1926). The Bacteriophage and Its Behavior. Baltimore, MD: Williams & Wilkins
- Molineux, I. J. (2006). Chapter 20: The T7 group. In: The Bacteriophages (R. Calendar, ed.), pp. 277. Oxford University Press, Oxford.
- "Teseptimavirus ~ ViralZone page". viralzone.expasy.org. Retrieved 2019-11-18.
- Molineux, Ian J.; Panja, Debabrata (March 2013). "Popping the cork: mechanisms of phage genome ejection". Nature Reviews Microbiology. 11 (3): 194–204. doi:10.1038/nrmicro2988. ISSN 1740-1534. PMID 23385786. S2CID 205498472.
- "Genome of bacteriophage T7". 9 September 2004. Retrieved 18 May 2011.
- Dunn, J. J.; Studier, F. W. (1983). "Complete nucleotide sequence of bacteriophage T7 DNA and the locations of T7 genetic elements". Journal of Molecular Biology. 166 (4): 477–535. doi:10.1016/S0022-2836(83)80282-4. PMID 6864790.
- "Uniprot: reference proteome of bacteriophage T7".
- Wright, Bradley W.; Molloy, Mark P.; Jaschke, Paul R. (2021-10-05). "Overlapping genes in natural and engineered genomes". Nature Reviews Genetics. 23 (3): 154–168. doi:10.1038/s41576-021-00417-w. ISSN 1471-0064. PMC 8490965. PMID 34611352.
- Chan LY, Kosuri S, Endy D (2005). "Refactoring bacteriophage T7". Molecular Systems Biology. 1: E1–E10. doi:10.1038/msb4100025. PMC 1681472. PMID 16729053.
- Häuser, R; Blasche, S; Dokland, T; Haggård-Ljungquist, E; von Brunn, A; Salas, M; Casjens, S; Molineux, I; Uetz, P (2012). Bacteriophage protein-protein interactions. Advances in Virus Research. Vol. 83. pp. 219–98. doi:10.1016/B978-0-12-394438-2.00006-2. ISBN 9780123944382. PMC 3461333. PMID 22748812.
- Heineman, R. H.; Bull, J. J. (2007). "Testing Optimality with Experimental Evolution: Lysis Time in a Bacteriophage". Evolution. 61 (7): 1695–1709. doi:10.1111/j.1558-5646.2007.00132.x. PMC 1974807. PMID 17598749.
- Chang, C. Y.; Kemp, P; Molineux, I. J. (2010). "Gp15 and gp16 cooperate in translocating bacteriophage T7 DNA into the infected cell". Virology. 398 (2): 176–86. doi:10.1016/j.virol.2009.12.002. PMC 2825023. PMID 20036409.
- "New Details about Bacteriophage T7-Host Interactions". Archived from the original on 2011-08-17.
- Jeffery, Constance J. (1999-01-01). "Moonlighting proteins". Trends in Biochemical Sciences. 24 (1): 8–11. doi:10.1016/S0968-0004(98)01335-8. ISSN 0968-0004. PMID 10087914.
- Lai YT, Masker W. Repair of double-strand breaks by incorporation of a molecule of homologous DNA. Mol Microbiol. 2000 Apr;36(2):437-46. PMID 10792729
- Yu M, Masker W. T7 single strand DNA binding protein but not T7 helicase is required for DNA double strand break repair. J Bacteriol. 2001 Mar;183(6):1862-9. PMID 11222583
- Bernstein C, Bernstein H. Coiled rings of DNA released from cells infected with bacteriophages T7 or T4 or from uninfected Escherichia coli. J Virol. 1974 Jun;13(6):1346-55. doi: 10.1128/JVI.13.6.1346-1355.1974. PMID 4598784; PMCID: PMC355455.
External links
- T7+Phage at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- ViralZone
- New Details about T7-host interactions. Microbe Magazine
- T7 reference proteome in Uniprot