Hectochlorin
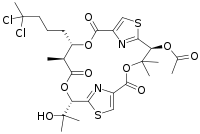
Hectochlorin is a lipopeptide that exhibits potent antifungal activity against C. albicans and a number of plants pathogens, as well as inhibiting growth of human cell lines by hyperpolymerization of actin.[1] It was originally isolated from the filamentous cyanobacterium Moorea producens JHB, collected from Hector Bay, Jamaica, 1996,[2] which is a strain also known for being the producer of other two potent biomolecules named Jamaicamide A[3] and Cryptomaldamide.[4] Due to its activity against plants pathogens, synthetic efforts elucidated the compound’s total synthesis in 2002.[5] Moorea species are normally the main component of the dietary of some sea hares, which concentrate the cyanobacterial metabolites as a mechanism of defense from predators. Therefore, in 2005, hectochlorin was re-isolated from the Thai sea hare Bursatella leachii, along with a new analogue, deacetylhectochlorin.[6] Another reisolation of hectochlorin was reported in 2013, from another Moorea producens strain (RS05), isolated from the Red Sea, surprising in a non-tropical environment, as opposed to the other Moorea strains isolated before. The predicted biosynthesis of hectochlorin was published in 2007 and consists in a hybrid NRPS-PKS, with a hexanoic acid as start unit that becomes halogenated twice in the position 5, producing fairly rare gem-dichloro group, that along with two 2,3-dihydroxyisovaleric acid (DHIV) units compose a very interesting bioactive molecule.
 | |
| Identifiers | |
|---|---|
3D model (JSmol) |
|
| ChEMBL | |
| ChemSpider | |
PubChem CID |
|
| |
| |
| Properties | |
| C27H34Cl2N2O9S2 | |
| Molar mass | 665.59 g·mol−1 |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
Infobox references | |
Biosynthesis
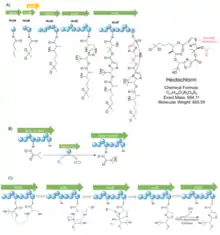
The biosynthetic gene cluster (BGC) is composed of eight genes (Figure 1A), of which seven are directly related to the synthesis of the molecule (hctA-B and hctD-H, in green) and one is predicted to encode a transposase (hctC, in yellow), that tends to be related to the mobility of the gene and not the synthesis of molecule features. The cluster is also flanked by other 5 ORF (Open reading rrames), including three hypothetical proteins, a homing endonuclease and a reverse transcriptase of unknown function regarding biosynthesis mechanistic of the molecule.
Biosynthesis of hectochlorin starts with hctA (Figure 1A), responsible for the start unit, which has 53% similarity to an Acyl-ACP synthetase in Fischerella and is predicted to generate a hexanoic acid that starts the hectochlorin molecule. This hexanoic acid gets halogenated twice at the fifth carbon by the gene hctB, generating the gem-dichloro group in 5,5-dichlorohexanoic acid. For such, hctB has one halogenation domain in the N-terminus, which is 47% similar to a halogenase at Microcystis aeruginosa and also contains all conserved residues for the binding of Fe2+/2-oxoglutarate co-factor. In the C-terminus, one ACP domain is present and it is fairly homologous to several others ACP domain at cyanobacteria, like Curacin A, Jamaicamide. As mentioned before, hctC is a transposase of unknown function and it is not directly related in the molecule synthesis. Next, this 5,5-dichlorohexanoic acquires one KS extension by hctD. This gene consists of a single KS module with a minimal configuration (KS-AT-CP) plus one KR and cMT, producing a 7,7-dichloro-3-hydroxy-2-methyl-octanoic acid. HctE consists of a bi-modular NRPS, of which the first module incorporates an isovaleric acid and the second module incorporates a heterocyclic cysteine. In the first module, the adenlyation-domain (A domain) has a mutation substituting a conserved aspartate residue (Asp235) that is related to interactions with the amino group of the cognate amino acid, therefore, incorporating isovaleric acid instead an amino acid. This hydroxyl group that substitutes an amino group condensates with the previous carbonyl from the KS extension. The second module has 67% identity to CurF and BarG (from Curacin A and Barbamide BGCs) and is predicted to adenylate and heterocyclize a cysteine, as well as oxidize it by FMN-dependent oxidase present in between adenlyation conserved motifs, catalyzing the formation of a thiazole ring (Figure 1C). HctF gene has remarkable similarity to hctE, although, there are two main differences: the iso-valeric acid incorporated does not condensate by the hydroxyl group that substitute an amino group, instead, it condensates be the hydroxyl group in the side chain created by a P450 oxidation (Figure 1B); hctF has an extra thioesterase domain that converts thioester bond to ester bond and catalyze the attack from the C-terminal free hydroxyl group to this newly formed ester, cyclizing the molecule. The final genes hctG and hctH probably encode two P450 that oxidize the side chain of both isovaleric acids (Figure 1D). Lastly, a post-NRPS modification happens in the free hydroxyl group from the second isovaleric acid, adding an acetyl group. This addition is not predicted in the current biosynthesis, although the absence of this post-NRPS modification would produce the analogue previously mentioned, deacetylhectochlorin.
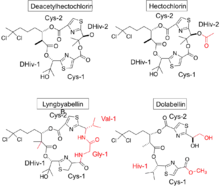
Currently in the MarinLit database, other compounds (besides deacetylhectochlorin and hectochlorin) that contain this fairly unusual gem-dichloro group are lyngbyabellin A-N, 27-deoxylyngbyabellin A and dolabellin, all of those synthesized from Moorea species. Most lyngbyabellins also contain DHIV in their structure, as well as dolabellin. Figure 2 compares the structures of deacetylhectochlorin, hectochlorin, lyngbyabellin B and dolabellin. This figure illustrates the similarities (black) and differences (red) between those compounds compared to deacetylhectochlorin. All of those compounds (but hectochlorin) do not have a proposed biosynthesis published.
References
- Ramaswamy, A. V., Sorrels, C. M. & Gerwick, W. H. Cloning and biochemical characterization of the hectochlorin biosynthetic gene cluster from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 70, 1977–1986 (2007).
- Marquez, B. L. et al. Structure and absolute stereochemistry of hectochlorin, a potent stimulator of actin assembly. J. Nat. Prod. 65, 866–871 (2002).
- Marquez, B. L. et al. Structure and absolute stereochemistry of hectochlorin, a potent stimulator of actin assembly. J. Nat. Prod. 65, 866–871 (2002).
- Kinnel RB et al; A Maldiisotopic Approach to Discover Natural Products: Cryptomaldamide, a Hybrid Tripeptide from the Marine Cyanobacterium Moorea producens. J Nat Prod. 2017 May 26;80(5):1514-1521. doi: 10.1021/acs.jnatprod.7b00019.
- Cetusic, J. R. P., Green, F. R., Graupner, P. R. & Oliver, M. P. Total Synthesis of Hectochlorin. Org. Lett. 4, 1307–1310 (2002).
- Suntornchashwej, S., Chaichit, N., Isobe, M. & Suwanborirux, K. Hectochlorin and morpholine derivatives from the Thai sea hare, Bursatella leachii. J. Nat. Prod. 68, 951–955 (2005).