CRACD-like protein
CRACD-like protein. previously known as KIAA1211L is a protein that in humans is encoded by the CRACDL gene. It is highly expressed in the cerebral cortex of the brain.[5] Furthermore, it is localized to the microtubules and the centrosomes and is subcellularly located in the nucleus.[6][7] Finally, CRACDL is associated with certain mental disorders and various cancers.[8][9]
| CRACDL | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | CRACDL, C2orf55, KIAA1211-like, KIAA1211 like, KIAA1211L, CRACD like | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | MGI: 1919347 HomoloGene: 19208 GeneCards: CRACDL | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Gene
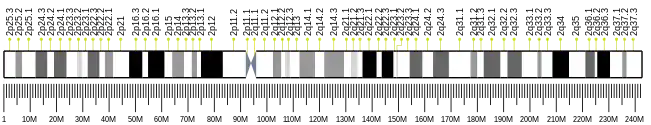
| Chromosome | 2 (2q.11.2)[10] |
| Location | 98,793,846 bp from pter to 98,936,259 bp from pter[10] |
| Size | 142,414 bases[10] |
| Accession Number | NM_207362[11] |
| Also Known As | KIAA1211 Like
C2orf55 Chromosome 2 Open Reading Frame 55[10] |
CRACDL is a protein-coding gene.[10] The table above presents the gene's alias, location, size and accession number.
Protein

| Amino Acid Length | 962 [10] |
| Molecular Weight | 102 kda[12] |
| Isoelectric Point | 8[12] |
| Accession Number | NP_997245.2[11] |
| Also Known As | Uncharacterized Protein KIAA1211-like[10]
Uncharacterized Protein C2orf55[10] Hypothetical Protein LOC343990[13] |
The table above presents the protein's alias, size, and accession number. The CRACD-L protein is proline rich and asparagine, isoleucine, phenylalanine, and tyrosine poor.[12]
Domains and motifs
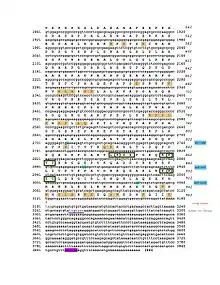
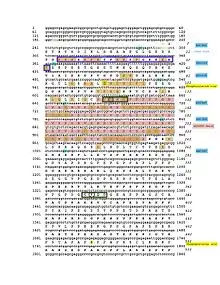
The CRACD-L protein has one domain called the DUF4592 motif and spans amino acids 131–239.[14] This domain is highly conserved among the CRACDL orthologs. The DUF4592 motif is depicted in both the conceptual translation and schematic figures.
Post translational modifications
CRACDL is phosphorylated at the Ser92 and Ser490 amino acids.[15] The KIAA1211L protein is also predicted to have five different SUMOylation sites located at Lys134, Lys375, Lys866, Lys874, and Lys914.[16] Both the phosphorylated sites and the SUMOylation sites are depicted in the conceptual translation and schematic figures.
Secondary structure
The CRACD-L protein predicted secondary structure is composed of 50% alpha helixes, 8.9% beta sheets, and 17.9% turns.[17] The high number of turns is consistent with the fact that CRACD-L is proline rich.[12]
Subcellular location
The CRACD-L protein is predicted to be located in the nucleus.[7] The orthologs, including the elephant shark, horse, rock dove, and chimp, are also predicted to be located in the nucleus.[7] The nuclear location signal is located on amino acids 25-43 which is depicted in both the conceptual translation and schematic figures. .[7] This signal is conserved throughout the orthologs. Additionally, this location (amino acids 24-43) is positively charged, probably due to the high amount of lysine at this location.[12] Finally, it is predicted that CRACD-L is mainly localized to the microtubules and centrosome and sometimes localized to the cytokinetic bridge.[6]
Expression
The gene is highly expressed in the cerebral cortex of the brain.[5] The CRACD-L protein is located in many different tissue types, including the brain, the hippocampus, the lung, breast carcinoma, the islets of Langerhans, the pancreas, the kidney, and 38 other tissues.[18] Additionally, it is expressed an average amount compared to other human proteins.[19]
Regulation of transcription
The promoter region of CRACDL is approximately 1340 base pairs with various predicted transcription factors.[20] The glial cells missing homolog 1 and the oligodendrocyte lineage transcription factors are notable because CRACDL is highly expressed in the brain.[20][5] Furthermore, the Estrogen-related receptor alpha is also a notable transcription factor due to CRACDL's low expression levels when estrogen receptors are knocked down.[21][20] Furthermore, CRACDL is predicted to be SUMOylated.[16] The 3' UTR of CRACDL is predicted to be a targeted by miRNA-132, which is depicted in the conceptual translation figure.[22]
Function
Interacting proteins
Glycogen Synthase Kinase 3 Beta (GSK3B)
GSK3B is a protein kinase that regulates transcription factors and microtubules.[23] As such, it phosphorylates proteins, decreasing their ability to bind and stabilize microtubules.[23] The proteins it phosphorylates are the principle components of neurofibrillary tangles in Alzheimer disease.[23] The protein is needed for the establishment of neuronal polarity and axon outgrowth and phosphorylates proteins in neuroblastoma cells.[23] Furthermore, it is associated with bipolar disease and is active in breast cancer cells.[23][24]
As such, the predicted interaction between CRACDL and GSK3B is likely because CRACDL is highly expressed in the brain, associated with bipolar disorder and breast cancer, and is localized on the microtubules.[5][6][8][9] The interaction between GSK3B and CRACDL was predicted using anti bait coimmunoprecipitation, pull down, tandem affinity purification, fluorescence polarization spectroscopy, protein kinases assay, two hybrid, and confocal microscopy experiments.[25]
CRACD-L protein is also predicted to interact with Alpha-synuclein (SNCA), E3 Ubiquitin-Protein Ligase Mdm2 (MDM2), Serine/Threonine-Protein Kinase PAK 1 (PAK 1), and DNA Replication Factor Cdt1 (CDT1).[25]
Homology
Paralogs
KIAA1211 is the paralog to KIAA1211L. KIAA1211 is located on chromosome 4 and has 1233 amino acids.[26] Its percent identity to KIAA1211L is 21%.[27] The KIAA1211 has an ortholog in the bacteria Proteus vulgarism, indicating the paralog duplicated 4290 million years ago, before KIAA1211L.[28][29]
Orthologs
Below is the table of various KIAA1211L orthologs. It includes closely, intermediately, and distantly related orthologs. The most distant ortholog is the elephant shark, indicating KIAA1211L duplicated 473 MYA. The amino acids conserved among all the KIAA1211L orthologs are depicted in the conceptual translation.
| Species[30] | NCBI Accession #[30] | Date of Divergence[31] | Sequence Identity[12] | Sequence Similarity[32] |
|---|---|---|---|---|
| Pan troglodytes (Chimpanzee) | XP_515643.2 | 6.65 MYA | 99.1% | 99.3% |
| Octodon negus (Degu) | XP_004633240.1 | 90 MYA | 65.9% | 73.1% |
| Panthera pardus (Leopard) | XP_019312964.1 | 96 MYA | 67.8% | 73.3% |
| Anas platyrhynchos (Mallard Duck) | XP_012949224.1 | 312 MYA | 41.2% | 52.40% |
| Pygoscelis adeliae (Adélie penguin) | XP_009321834.1 | 312 MYA | 38.5% | 51.6% |
| Python bivittatus (Burmese python) | XP_007428826 | 312 MYA | 34.2% | 46.3% |
| Nanorana parker (High Himalaya frog) | XP_018418330.1 | 352 MYA | 32.1% | 43.7% |
| Callorhinchus milii (Elephant Shark) | XP_007889338.1 | 473 MYA | 30.5% | 42.4% |
Phylogeny
The CRACDL gene is similar and conserved in mammals, birds, reptiles, amphibians, and fish. It is not conserved in bacteria, archaea, protists, plants, fungus, trichoplax, and invertebrates.
Citations
- GRCh38: Ensembl release 89: ENSG00000196872 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000026090 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "KIAA1211L KIAA1211 like [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-04-23.
- "Cell atlas - KIAA1211L - The Human Protein Atlas". www.proteinatlas.org. Retrieved 2017-04-23.
- "GenScript Protein Subcellular Location Prediction Tool".
- Spurrell CH (2013). Identifying New Genes for Inherited Breast Cancer by Exome Sequencing (Doctor of Philosophy thesis). University of Washington.
- Iwamoto K, Kakiuchi C, Bundo M, Ikeda K, Kato T (April 2004). "Molecular characterization of bipolar disorder by comparing gene expression profiles of postmortem brains of major mental disorders". Molecular Psychiatry. 9 (4): 406–16. doi:10.1038/sj.mp.4001437. PMID 14743183.
- Database, GeneCards Human Gene. "KIAA1211L Gene - GeneCards | K121L Protein | K121L Antibody". www.genecards.org. Retrieved 2017-02-24.
- "Homo sapiens KIAA1211 like (KIAA1211L), mRNA - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-04-23.
- Workbench, NCSA Biology. "SDSC Biology Workbench". workbench.sdsc.edu. Retrieved 2017-04-23.
- "Genatlas sheet". genatlas.medecine.univ-paris5.fr. Retrieved 2017-04-23.
- "Pfam: Family: DUF4592 (PF15262)". pfam.xfam.org. Retrieved 2017-04-23.
- "Homo sapiens KIAA1211 like (KIAA1211L), mRNA - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-04-23.
- "SUMOplot™ Analysis Program | Abgent". www.abgent.com. Retrieved 2017-04-23.
- Kumar, Prof. T. Ashok. "BioGem.Org - Ashok Kumar's Bioinformatics Portal... | Home". www.biogem.org. Retrieved 2017-04-23.
- "Tissue expression of KIAA1211L - Summary - The Human Protein Atlas". www.proteinatlas.org. Retrieved 2017-04-23.
- "KIAA1211L protein abundance in PaxDb". pax-db.org. Retrieved 2017-04-23.
- "Genomatix - NGS Data Analysis & Personalized Medicine". www.genomatix.de. Retrieved 2017-05-07.
- Al Saleh S, Al Mulla F, Luqmani YA (2011). "Estrogen receptor silencing induces epithelial to mesenchymal transition in human breast cancer cells". PLOS ONE. 6 (6): e20610. Bibcode:2011PLoSO...620610A. doi:10.1371/journal.pone.0020610. PMC 3119661. PMID 21713035.
- Alvarez-Saavedra, M (2010). "MicroRNA-132-Dependent Post-Transcriptional Regulation of Clock Entrainment Physiology Via Modulation of Chromatin Remodeling and Translational Control Gene Targets". University of Ottawa.
- "GSK3B - Glycogen synthase kinase-3 beta - Homo sapiens (Human) - GSK3B gene & protein". www.uniprot.org. Retrieved 2017-04-23.
- Database, GeneCards Human Gene. "GSK3B Gene - GeneCards | GSK3B Protein | GSK3B Antibody". www.genecards.org. Retrieved 2017-04-23.
- "IntAct". www.ebi.ac.uk. Retrieved 2017-04-23.
- Database, GeneCards Human Gene. "KIAA1211 Gene - GeneCards | K1211 Protein | K1211 Antibody". www.genecards.org. Retrieved 2017-04-23.
- Myers EW, Miller W (March 1988). "Optimal alignments in linear space". Computer Applications in the Biosciences. 4 (1): 11–7. doi:10.1093/bioinformatics/4.1.11. PMID 3382986.
- "kiaa1211l KIAA1211 like [Callorhinchus milii (elephant shark)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-02-24.
- "TimeTree :: The Timescale of Life". www.timetree.org. Retrieved 2017-02-24.
- "BLAST: Basic Local Alignment Search Tool". blast.ncbi.nlm.nih.gov. Retrieved 2017-04-23.
- "TimeTree :: The Timescale of Life". www.timetree.org. Retrieved 2017-04-23.
- EMBL-EBI. "EMBOSS Needle < Pairwise Sequence Alignment < EMBL-EBI". www.ebi.ac.uk. Retrieved 2017-04-23.