L-Photo-methionine
L-Photo-methionine is a photo-reactive amino acid derivative of L-methionine that was synthetically formed in 2005.[1] Protein are long polymer chains of amino acids; which can range in various structures and sizes. Proteins can interact with each other (protein-protein interactions or PPI) and with these interactions, affects cellular interactions and pathways.[1] Such interactions; in viral fusion and in growth-factor signaling looked promising for antiviral or anti-cancer drugs, so research must be done to understand the interactions.[1] With that, research has begun to prove that proteins function in supramolecular complexes compared to isolated entities.[1] So, scientists Monika Suchanek, Anna Radzikowski, and Christoph Thiele researched that the direct way to study these interactions in the natural environment better was to create a new way of photo-cross-linking proteins; which led to the synthesis of L-photo-methionine and in that same study, L-photo-leucine.[1]
 | |
| Names | |
|---|---|
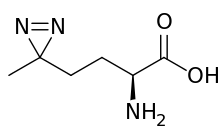
| IUPAC name
(S)-2-amino-4-(3-methyl-3H-diazirin-3-yl)butanoic acid | |
| Other names
L-2-amino-5,5-azi-hexanoic acid | |
| Identifiers | |
3D model (JSmol) |
|
| ChemSpider | |
PubChem CID |
|
| |
| |
| Properties | |
| C6H11N3O2 | |
| Molar mass | 157.173 g·mol−1 |
| 6 mg/mL | |
| Hazards | |
| GHS labelling: | |
| H315, H319, H335 | |
| P261, P305, P338, P351 | |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
Infobox references | |
Synthesis
Racemic Photo-Methionine is synthesized from 4,4'-azi-pentanal by the Strecker amino acid synthesis.[1] The L enantiomer is separated by enzymatic resolution of the acetamide.

Photoactivation of Methionine
As it was previously mentioned, L-Photo-Methionine can be used to study protein-protein interactions with the proteins in their native environment. How this is possible is how the amino acid behaves when exposed to UV light.[1] To prove that first the synthesis works, a radioactive carbon (14C) as added under its own synthesis to perform proper spectroscopic methods.
The Activation of Photo-Methionine
Because the previous synthesis had worked, photo-methionine is photo-reactive due to the diazirine ring. Once this ring has become exposed to UV light, nitrogen leaves as nitrogen gas (N2) and forms the highly reactive intermediate carbene.[1] Photo-activation of amino acids provide the ability of photo-cross-linking in proteins.[1] This type of cross-linking has three major advantages; there is greater specificity for this cross linking due to the short lived intermediates and that this amino acid is functional, and most importantly; not toxic (meaning it should not disrupt the protein's function or structure dramatically).[1] Research had found that this activation is the rate-limiting step; not the cross-linkage.[2]

A New Efficient Synthesis and Usage
Scientists Miquel Vila-Perello´, Matthew R. Pratt, Frej Tulin, and Tom W. Muir wanted to create an efficient synthesis as the original had required an enzymatic solution and had a low yield.[3] So, they started with L-glutamic acid with protecting groups on both the carboxylic acid (tert-butyl), and Boc on the amine. This synthesis will not undergo detail as the classic, but below is the full synthesis. To find the actual steps, look to the reference.[3]
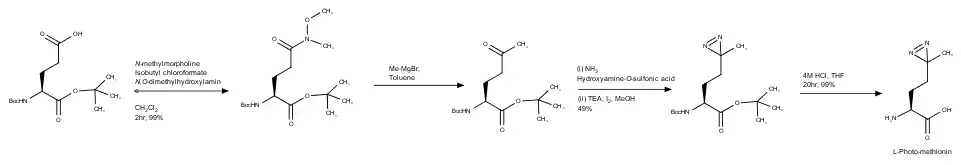
So, once they had synthesized L-photo-methionine, the yield was 32%, much higher (by six times) the original synthesis.[3] It was used then (with a protection group Fmoc on the amine) which that product underwent more synthetic steps to study if an amino-acid cross linker and a post-translational modification (PTM) could be introduced to the same protein site specifically to capture a covalent interaction of the amino-acid is dependent on the PTM.[3] PTM's regulate protein-protein interactions that have characteristics that are transient and substoichiometric; making these difficult to detect by standard methods.[3] So, in order to see if it would work, the MH2 domain of Smad2 was used because this signaling protein is known to form stable homo-trimers once they come into contact with receptor-phosphorylated serine residues.[3] Expression protein ligation (known as EPL) was used to synthesize to form Smad2-MH2-CSpSM-photo-Met (1). The product was studied with the cross-linker (photo-Met) against a control protein: HA-MH2-CSpSMpS (this lacks photo-methionine, 2) using SDS-PAGE and western blotting using anti-HA antibody.[3] 1 had generated two major cross-linked species that have molecular weight consistent with a dimer and trimer of Smad2-SH2. Without that cross-linker, the dimer and trimer were barely detected in the non-irradiated 1, and in 2 before and after UV irradiation.[3] Proving that l-photo-methionine can be used with EPL and could be used to determine a transient MH2-MH2 interaction that was dependent on a PTM.[3]
Usage of Photo-Methionine
Complex and Function of Membrane Protein in Cholesterol Homeostasis
As mentioned before, scientists Monika Suchanek, Anna Radzikowski, and Christoph Thiele wanted to study protein-protein interaction in their natural environment. Specifically, the membrane proteins (in a complex and are SCAP, Insig-1, and SREBP) that regulate cholesterol homeostasis so they wanted to know what their function was and the complex structure.[1] What they had found was that using this photo-reactive amino acid was incorporated efficiently into the protein by mammalian cells, but did not need to use modified tRNAs (transfer RNA's) or AARS's (aminoacyl tRNA syntheses) which that allowed the specific cross-linking needed.[1] This cross-linking could be determined by western blotting and they had discovered a direct interaction between Insig-1 and PGRMC1 (a progesterone-binding membrane protein).[1] All four of the membrane proteins are found in the endoplasmic reticulum and the complex responds to low cholesterol levels.[1] Cells (COS7) that had HA (hemagglutinin tagged PGRMC1) and Myc tagged Insig-1 were grown with and without photo-Met. In the presence of photo-Met, Insig-1 and SCAP had cross-linked with PGRMC1; specifically, Insig-1 cross-linked had a strong band.[1] The cross-linking was detected by immunoprecipitating detergent-extracts with an antibody to HA then the precipitant was tested for Insig-1 using western blotting with the antibody for Myc.[1] An identical band was found doing the reverse order of the detection; meaning Myc antibody was immunoprecipitated then followed by blotting with the HA antibody.[1] So, the method had proven to work that photo-Met could cross-link proteins, but the physiological implications of this cross-linking has yet to be determined.
Usage of Protein Nanoprobes to Study Protein-Protein Interactions with Mass Spectrometry
Protein was studied using a protein nanoprobe (that enables cross-linking) that introduced photo-methionine within the protein (during the recombinant expression) which lead to the protein keeping its reserved structure while having the ability to be mapped out for its interactions. The model was used as a region of contact surface that is involved in a well-known interaction (homodimerization) between two molecules of 14-3-3ζ protein. Once the photo-methionine is introduced and has become activated using UV-light, it can cross-link with no specificity (meaning no group) and the links have zero-length. High resolution mass spectrometry or MS can (even MS/MS) be used then to determine the cross-linked residues and the reaction radius; allowing the researchers to characterize and research the homodimerization of the protein. The usage of the high-resolution MS with photo-methionine has its advantages as it again allows the protein to be in its native state, there are reasonable time scales while using small quantities of the protein. There are also fewer limitations on the reaction specificity and restrictions using a photo-active cross-linker (photo-methionine) compared to chemical cross-linking. This method of combined photo-initiated cross-linking from the protein nanoprobe in tandem with MS could be useful to characterize not only homodimer formation but also oligomers and in theory; heteromers (such as the composition of the protein-protein mixture and its functionality).[4]
Ability to Study Cytochrome P450 Electron-Transport Chain using Photo-cytochrome b5
Cytochrome b5 was synthesized with photo-methionine to map the protein-protein interactions while also identifying its structure to study the mammalian mixed function oxidase system (also known as the MFO).[5] This system is located in the membrane of the endoplasmic reticulum and it is composed of cytochrome P450, NADPH: cytochrome P450 reductase, and cytochrome b5 along with NADH: cytochrome b5 reductase. Once the cytochrome b5 complex had photo-methionine incorporated (meaning photo-met was substituted in place of methionine and now photo-cyt b5), photo-cyt b5 and cytochrome P450 were put under UV-light and the products were able to be studied using SDS-Page; this method had shown three cross-links. The photo-methionine had proven successful in mapping photo-cyt b5 as the MALDI-TOF method shown three oligomers (from chymotryptic peptides) that were composed of photo-cyt b5 and cytochrome P450 in molecular weight ratio's of 1:1, 1:2, and 2:1. What makes photo-methionine here so useful in studying cytochrome P450 and cytochrome b5 is that this method not only mapped protein-protein interfaces not only in regions exposed to solvent, but also in the native environment; the membrane.[5] A typical cross-linking method can only work in solvent exposed regions, proving once again that photo-methionine is useful to map these protein-protein interactions with the protein in their native environment.[5]
Analysis of Nidogen-1 interacting with Laminin γ1
Laminin's are non-collagenous proteins found in basement membranes and form networks through non-covalent self-interactions. Nidogens (also known as entactins) are sulfated monomeric glycoproteins that are ubiquitously present in basement membranes of higher organisms. Nidogens help with the formation of the basement. With both laminins and nidogens present, both interact with each other to have a stoichiometry relationship of 1:1 in a complex. In order to study the short arm of laminin γ1, photo methionine introduced both to nidogen-1, laminin γ1 LEb2-4, and laminin γ1 short arm to see if this photo-cross linking method could map out the structure. MS/MS analysis was done before cross-linking to find only 13-25% of methionine's had been incorporated, but once UV-A-induced or another cross-linker, BS2G-mediated cross-linked (a homobifunctional cross-linker), the percentage of photo-methionine's had increased to 35%. Both cross-linkers had shown extra structural insight both computationally and experimentally to help with understanding the functions.[6]
Revelation of Dimeric Structure and Oligomeric Structure
Cyclooxygenase-2 (COX-2) and microsomal prostaglandin E2 synthase-1 (mPGES-1) structures were studied using both photo-methionine (photo-activatable) and bifunctional cross linkers. Photo-methionine used in COX-2 had shown just as the bifunctional cross-linker that there was a dimeric structure which this was consistent with the crystal structure of the enzyme. In mPGES-1, the human cells (A549) had been treated with disuccinimidyl suberate (chemical cross-linker) had yielded a dimer of 33kDa and a trimer of 45kDa while it was treated with photo-methionine had yielded a dimer of the same molecular weight (33kDa) and two putative trimers (50kDa and 55kDa). Once a mPGES-1 inhibitor (MF63) was introduced; this had inhibited the formation of the 50kDa and 55kDa complexes. The dimer and trimer yielded by the chemical cross-linker was not affected by the inhibitor. Yet, photo-methionine nor disuccinimidyl suberate had not shown any protein-protein interactions between COX-2 and mPGES-1 and this could be due to various reasons. For photo-methionine; one could be due to the low incorporation at this time as it was 0.7%. So, for mPGES-1; this has 152 amino acids so only one photo-methionine would be incorporated per monomer. That also means not one specific methionine would be replaced so that results in a heterogeneous population of mPGES-1 resulting in different cross-linking. Even though it did not show any protein-protein interactions, it could be used to detect inhibitor-induced protein conformational changes in the cell membranes on top of determining oligomeric structures.[7]
Escherichia coli cells
Photo-methionine can be used to label recombinant proteins in Escherichia coli cells; though methionine in general is a rare amino acid so that means it could only give limited structural data.[8] Nevertheless, photo-methionine was incorporated into Ca2+ regulating protein calmodulin (CaM that was 17-kDa) that has nine methionine's and studied via mass spectrometry (MS).[9] What makes this method different is the use of mineral salts medium instead of DMEM (Dulbecco's Modified Eagle's Limiting Medium) or dialyzed fetal bovine serum for the incorporation into the cells.[9] Using the mineral salt medium allowed the cells to be grown from the beginning in order to eliminate complicated steps with other protocols (incubating the cells in LB medium, followed by washing, and further incubating in the depleted medium), meaning that photo-methionine could be incorporated at the very beginning of the cell growth that had a high yield above 30%.[9] Photo-methionine had shown no damage during the cell growth process and once photo-activated by UV-A light, CaM had nine distinct cross-link sites once MS had determined there was peaks of photo-methionine labeled CaM.[9] Not only can photo-methionine be used for mapping 3D protein structure, studying protein-protein interactions, but now hydrophobic regions in the protein.[9]
References
- Suchanek, Monika; Radzikowska, Anna; Thiele, Christoph (April 2005). "Photo-leucine and photo-methionine allow identification of protein-protein interactions in living cells". Nature Methods. 2 (4): 261–268. doi:10.1038/nmeth752. ISSN 1548-7105. PMID 15782218. S2CID 205427639.
- Kölbel, Knut; Ihling, Christian H.; Sinz, Andrea (2012). "Analysis of Peptide Secondary Structures by Photoactivatable Amino Acid Analogues". Angewandte Chemie International Edition. 51 (50): 12602–12605. doi:10.1002/anie.201205308. ISSN 1521-3773. PMID 23109332.
- Vila-Perelló, Miquel; Pratt, Matthew R.; Tulin, Frej; Muir, Tom W. (2007-07-01). "Covalent Capture of Phospho-Dependent Protein Oligomerization by Site-Specific Incorporation of a Diazirine Photo-Cross-Linker". Journal of the American Chemical Society. 129 (26): 8068–8069. doi:10.1021/ja072013j. ISSN 0002-7863. PMC 3242410. PMID 17567014.
- Ptáčková, Renata; Ječmen, Tomáš; Novák, Petr; Hudeček, Jiří; Stiborová, Marie; Šulc, Miroslav (June 2014). "The Application of an Emerging Technique for Protein–Protein Interaction Interface Mapping: The Combination of Photo-Initiated Cross-Linking Protein Nanoprobes with Mass Spectrometry". International Journal of Molecular Sciences. 15 (6): 9224–9241. doi:10.3390/ijms15069224. PMC 4100091. PMID 24865487.
- Ječmen, Tomáš; Ptáčková, Renata; Černá, Věra; Dračínská, Helena; Hodek, Petr; Stiborová, Marie; Hudeček, Jiří; Šulc, Miroslav (2015-11-01). "Photo-initiated crosslinking extends mapping of the protein–protein interface to membrane-embedded portions of cytochromes P450 2B4 and b5". Methods. Mass Spectrometry-Based Structural Biology. 89: 128–137. doi:10.1016/j.ymeth.2015.07.015. ISSN 1046-2023. PMID 26235815.
- Lössl, Philip; Kölbel, Knut; Tänzler, Dirk; Nannemann, David; Ihling, Christian H.; Keller, Manuel V.; Schneider, Marian; Zaucke, Frank; Meiler, Jens; Sinz, Andrea (2014-11-11). "Analysis of Nidogen-1/Laminin γ1 Interaction by Cross-Linking, Mass Spectrometry, and Computational Modeling Reveals Multiple Binding Modes". PLOS ONE. 9 (11): e112886. Bibcode:2014PLoSO...9k2886L. doi:10.1371/journal.pone.0112886. ISSN 1932-6203. PMC 4227867. PMID 25387007.
- Hétu, Pierre-Olivier; Ouellet, Marc; Falgueyret, Jean-Pierre; Ramachandran, Chidambaram; Robichaud, Joel; Zamboni, Robert; Riendeau, Denis (2008-09-01). "Photo-crosslinking of proteins in intact cells reveals a dimeric structure of cyclooxygenase-2 and an inhibitor-sensitive oligomeric structure of microsomal prostaglandin E2 synthase-1". Archives of Biochemistry and Biophysics. 477 (1): 155–162. doi:10.1016/j.abb.2008.04.038. ISSN 0003-9861. PMID 18498757.
- Kohl, Bastian; Brüderlin, Mitchell; Ritz, Danilo; Schmidt, Alexander; Hiller, Sebastian (2020-08-07). "Protocol for High-Yield Production of Photo-Leucine-Labeled Proteins in Escherichia coli". Journal of Proteome Research. 19 (8): 3100–3108. doi:10.1021/acs.jproteome.0c00105. ISSN 1535-3893. PMID 32412763. S2CID 218658726.
- Piotrowski, Christine; Ihling, Christian H.; Sinz, Andrea (2015-11-01). "Extending the cross-linking/mass spectrometry strategy: Facile incorporation of photo-activatable amino acids into the model protein calmodulin in Escherichia coli cells". Methods. Mass Spectrometry-Based Structural Biology. 89: 121–127. doi:10.1016/j.ymeth.2015.02.012. ISSN 1046-2023. PMID 25726908.