Polyproline helix
A polyproline helix is a type of protein secondary structure which occurs in proteins comprising repeating proline residues.[1] A left-handed polyproline II helix (PPII, poly-Pro II, κ-helix[2]) is formed when sequential residues all adopt (φ,ψ) backbone dihedral angles of roughly (-75°, 150°) and have trans isomers of their peptide bonds. This PPII conformation is also common in proteins and polypeptides with other amino acids apart from proline. Similarly, a more compact right-handed polyproline I helix (PPI, poly-Pro I) is formed when sequential residues all adopt (φ,ψ) backbone dihedral angles of roughly (-75°, 160°) and have cis isomers of their peptide bonds. Of the twenty common naturally occurring amino acids, only proline is likely to adopt the cis isomer of the peptide bond, specifically the X-Pro peptide bond; steric and electronic factors heavily favor the trans isomer in most other peptide bonds. However, peptide bonds that replace proline with another N-substituted amino acid (such as sarcosine) are also likely to adopt the cis isomer.
Polyproline II helix
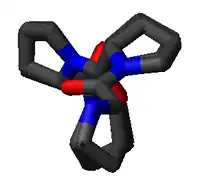
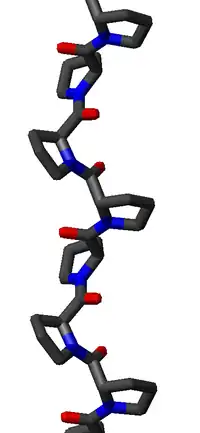
The PPII helix is defined by (φ,ψ) backbone dihedral angles of roughly (-75°, 150°) and trans isomers of the peptide bonds. The rotation angle Ω per residue of any polypeptide helix with trans isomers is given by the equation
Substitution of the poly-Pro II (φ,ψ) dihedral angles into this equation yields almost exactly Ω = -120°, i.e., the PPII helix is a left-handed helix (since Ω is negative) with three residues per turn (360°/120° = 3). The rise per residue is approximately 3.1 Å. This structure is somewhat similar to that adopted in the fibrous protein collagen, which is composed mainly of proline, hydroxyproline, and glycine. PPII helices are specifically bound by SH3 domains; this binding is important for many protein-protein interactions and even for interactions between the domains of a single protein.
The PPII helix is relatively open and has no internal hydrogen bonding, as opposed to the more common helical secondary structures, the alpha helix and its relatives the 310 helix and the pi helix, as well as the β-helix. The amide nitrogen and oxygen atoms are too far apart (approximately 3.8 Å) and oriented incorrectly for hydrogen bonding. Moreover, these atoms are both H-bond acceptors in proline; there is no H-bond donor due to the cyclic side chain.
The PPII backbone dihedral angles (-75°, 150°) are observed frequently in proteins, even for amino acids other than proline.[3] The Ramachandran plot is highly populated in the PPII region, comparably to the beta sheet region around (-135°, 135°). For example, the PPII backbone dihedral angles are often observed in turns, most commonly in the first residue of a type II β-turn. The "mirror image" PPII backbone dihedral angles (75°, -150°) are rarely seen, except in polymers of the achiral amino acid glycine. The analog of the poly-Pro II helix in poly-glycine is called the poly-Gly II helix. Some proteins, such as the antifreeze protein of Hypogastrura harveyi consist of bundles of glycine-rich polyglycine II helices.[4] This remarkable protein, whose 3D structure is known,[5] has unique NMR spectra and is stabilized by dimerization and 28 Cα-H··O=C hydrogen bonds.[6] The PPII helix is not common in transmembrane proteins, and this secondary structure does not traverse lipid membranes in natural conditions. In 2018, a group of researcher from Germany constructed and experimentally observed the first transmembrane PPII helix formed by specifically designed artificial peptides.[7][8]
Polyproline I helix
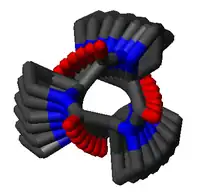

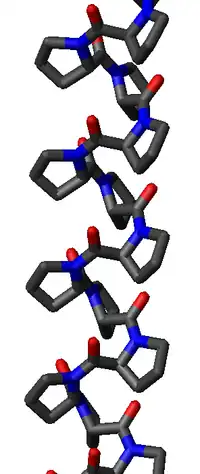
The poly-Pro I helix is much denser than the PPII helix due to the cis isomers of its peptide bonds. It is also rarer than the PPII conformation because the cis isomer is higher in energy than the trans. Its typical dihedral angles (-75°, 160°) are close, but not identical to, those of the PPII helix. However, the PPI helix is a right-handed helix and more tightly wound, with roughly 3.3 residues per turn (rather than 3). The rise per residue in the PPI helix is also much smaller, roughly 1.9 Å. Again, there is no internal hydrogen bonding in the poly-Pro I helix, both because an H-bond donor atom is lacking and because the amide nitrogen and oxygen atoms are too distant (roughly 3.8 Å again) and oriented incorrectly.
Structural properties
Traditionally, PPII has been considered to be relatively rigid and used as a "molecular ruler" in structural biology, e.g., to calibrate FRET efficiency measurements. However, subsequent experimental and theoretical studies have called into question this picture of a polyproline peptide as a "rigid rod".[9][10] Further studies using terahertz spectroscopy and density functional theory calculations highlighted that polyproline is in fact much less rigid than originally thought.[11] Interconversions between the PPII and PPI helix forms of poly-proline are slow, due to the high activation energy of X-Pro cis-trans isomerization (Ea ≈ 20 kcal/mol); however, this interconversion may be catalyzed by specific isomerases known as prolyl isomerases or PPIases. The interconversion between the PPII and PPI helices involve the cis-trans peptide bond isomerization along the whole peptide chain. Studies based on ion-mobility spectrometry revealed existence of a defined set of intermediates along this process.[12]
References
- Adzhubei, Alexei A.; Sternberg, Michael J.E.; Makarov, Alexander A. (2013). "Polyproline-II Helix in Proteins: Structure and Function". Journal of Molecular Biology. 425 (12): 2100–2132. doi:10.1016/j.jmb.2013.03.018. ISSN 0022-2836. PMID 23507311.
- "DSSP". pdb-redo.eu (in Dutch). Retrieved 2023-07-24.
- Adzhubei, Alexei A.; Sternberg, Michael J.E. (1993). "Left-handed Polyproline II Helices Commonly Occur in Globular Proteins". Journal of Molecular Biology. 229 (2): 472–493. doi:10.1006/jmbi.1993.1047. ISSN 0022-2836. PMID 8429558.
- Davies, Peter L.; Graham, Laurie A. (2005-10-21). "Glycine-Rich Antifreeze Proteins from Snow Fleas". Science. 310 (5747): 461. doi:10.1126/science.1115145. ISSN 0036-8075. PMID 16239469.
- Pentelute, Brad L.; Gates, Zachary P.; Tereshko, Valentina; Dashnau, Jennifer L.; Vanderkooi, Jane M.; Kossiakoff, Anthony A.; Kent, Stephen B. H. (2008-07-01). "X-ray Structure of Snow Flea Antifreeze Protein Determined by Racemic Crystallization of Synthetic Protein Enantiomers". Journal of the American Chemical Society. 130 (30): 9695–9701. doi:10.1021/ja8013538. ISSN 0002-7863. PMC 2719301. PMID 18598029.
- Treviño, Miguel Ángel; Pantoja-Uceda, David; Menéndez, Margarita; Gomez, M. Victoria; Mompeán, Miguel; Laurents, Douglas V. (2018-11-15). "The Singular NMR Fingerprint of a Polyproline II Helical Bundle". Journal of the American Chemical Society. 140 (49): 16988–17000. doi:10.1021/jacs.8b05261. PMID 30430829.
- Kubyshkin, Vladimir; Grage, Stephan L.; Bürck, Jochen; Ulrich, Anne S.; Budisa, Nediljko (2018). "Transmembrane Polyproline Helix". The Journal of Physical Chemistry Letters. 9 (9): 2170–2174. doi:10.1021/acs.jpclett.8b00829. PMID 29638132.
- Kubyshkin, Vladimir; Grage, Stephan L.; Ulrich, Anne S.; Budisa, Nediljko (2019). "Bilayer thickness determines the alignment of model polyproline helices in lipid membranes". Physical Chemistry Chemical Physics. 21 (40): 22396–22408. Bibcode:2019PCCP...2122396K. doi:10.1039/c9cp02996f. PMID 31577299.
- S. Doose, H. Neuweiler, H. Barsch, and M. Sauer, Proc. Natl. Acad. Sci. USA. 104, 17400 (2007)
- M. Moradi, V. Babin, C. Roland, T. A. Darden, and C. Sagui, Proc. Natl. Acad. Sci. USA. 106, 20746 (2009)
- M. T. Ruggiero, J. Sibik, J. A. Zeitler, and T. M. Korter, Agnew. Chemie. Int. Ed. 55, 6877 (2016)
- El-Baba, Tarick J.; Fuller, Daniel R.; hales, David A.; Russel, David H.; Clemmer, David E. (2019). "Solvent Mediation of Peptide Conformations: Polyproline Structures in Water, Methanol, Ethanol, and 1-Propanol as Determined by Ion Mobility Spectrometry-Mass Spectrometry". Journal of the American Society for Mass Spectrometry. 30 (1): 77–84. Bibcode:2019JASMS..30...77E. doi:10.1007/s13361-018-2034-7. PMC 6503664. PMID 30069641.
