Single-molecule experiment
A single-molecule experiment is an experiment that investigates the properties of individual molecules. Single-molecule studies may be contrasted with measurements on an ensemble or bulk collection of molecules, where the individual behavior of molecules cannot be distinguished, and only average characteristics can be measured. Since many measurement techniques in biology, chemistry, and physics are not sensitive enough to observe single molecules, single-molecule fluorescence techniques (that have emerged since the 1990s for probing various processes on the level of individual molecules) caused a lot of excitement, since these supplied many new details on the measured processes that were not accessible in the past. Indeed, since the 1990s, many techniques for probing individual molecules have been developed.[2]
The first single-molecule experiments were patch clamp experiments performed in the 1970s, but these were limited to studying ion channels. Today, systems investigated using single-molecule techniques include the movement of myosin on actin filaments in muscle tissue and the spectroscopic details of individual local environments in solids. Biological polymers' conformations have been measured using atomic force microscopy (AFM). Using force spectroscopy, single molecules (or pairs of interacting molecules), usually polymers, can be mechanically stretched, and their elastic response recorded in real time.
History
In the gas phase at ultralow pressures, single-molecule experiments have been around for decades, but in the condensed phase only since 1989 with the work by W. E. Moerner and Lothar Kador.[3] One year later, Michel Orrit and Jacky Bernard were able to show also the detection of the absorption of single molecules by their fluorescence.[4]
Many techniques have the ability to observe one molecule at a time, most notably mass spectrometry, where single ions are detected. In addition, one of the earliest means of detecting single molecules, came about in the field of ion channels with the development of the patch clamp technique by Erwin Neher and Bert Sakmann (who later went on to win the Nobel prize for their seminal contributions). However, the idea of measuring conductance to look at single molecules placed a serious limitation on the kind of systems which could be observed.
Fluorescence is a convenient means of observing one molecule at a time, mostly due to the sensitivity of commercial optical detectors, capable of counting single photons. However, spectroscopically, the observation of one molecule requires that the molecule is in an isolated environment and that it emits photons upon excitation, which owing to the technology to detect single photons by use of photomultiplier tubes (PMT) or avalanche photodiodes (APD), enables one to record photon emission events with great sensitivity and time resolution.
More recently, single-molecule fluorescence is the subject of intense interest for biological imaging, through the labeling of biomolecules such as proteins and nucleotides to study enzymatic function which cannot easily be studied on the bulk scale, due to subtle time-dependent movements in catalysis and structural reorganization. The most studied protein has been the class of myosin/actin enzymes found in muscle tissues. Through single-molecule techniques the step mechanism has been observed and characterized in many of these proteins.
In 1997, single-molecule detection was demonstrated with surface-enhanced Raman spectroscopy (SERS) by K. & H. Kneipp, Y. Wang, L.T. Perelman and others[5] at MIT and independently by S. Nie and S. R. Emory at Indiana University.[6] The MIT team used non-resonance Raman excitation and surface enhancement with silver nanoclusters to detect single cresyl violet molecules, while the team at Indiana University used resonance Raman excitation and surface enhancement with silver nanoparticles to detect single rhodamine 6G molecules.
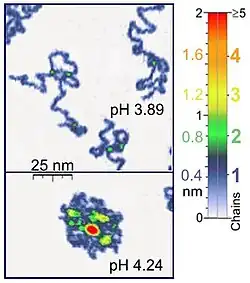
Nanomanipulators such as the atomic force microscope are also suited to single-molecule experiments of biological significance, since they work on the same length scale of most biological polymers. Besides, atomic force microscopy (AFM) is appropriate for the studies of synthetic polymer molecules. AFM provides a unique possibility of 3D visualization of polymer chains. For instance, AFM tapping mode is gentle enough for the recording of adsorbed polyelectrolyte molecules (for example, 0.4 nm thick chains of poly(2-vinylpyridine)) under liquid medium. The location of two-chain-superposition correspond in these experiments to twice the thickness of single chain (0.8 nm in the case of the mentioned example). At the application of proper scanning parameters, conformation of such molecules remain unchanged for hours that allows the performance of experiments under liquid media having various properties.[1] Furthermore, by controlling the force between the tip and the sample high resolution images can be obtained.[7][8] Optical tweezers have also been used to study and quantify DNA-protein interactions.[7][8]
About the experiments
Concept
Single-molecule fluorescence spectroscopy uses the fluorescence of a molecule for obtaining information on its environment, structure, and position. The technique affords the ability of obtaining information otherwise not available due to ensemble averaging (that is, a signal obtained when recording many molecules at the same time represents an average property of the molecules' dynamics). The results in many experiments of individual molecules are two-state trajectories.
Single-channel recording

As in the case of single molecule fluorescence spectroscopy, the technique known as single channel recording can be used to obtain specific kinetic information—in this case about ion channel function—that is not available when ensemble recording, such as whole-cell recording, is performed.[9] Specifically, ion channels alternate between conducting and non-conducting classes, which differ in conformation. Therefore, the functional state of ion channels can be directly measured with sufficiently sensitive electronics, provided that proper precautions are taken to minimize noise. In turn, each of these classes may be divided into one or more kinetic states with direct bearing on the underlying function of the ion channel. Performing these types of single molecule studies under systematically varying conditions (e.g. agonist concentration and structure, permeant ion and/or channel blocker, mutations in the ion channel amino acids), can provide information regarding the interconversion of various kinetic states of the ion channel. In a minimal model for an ion channel, there are two states: open and closed. However, other states are often needed in order to accurately represent the data, including multiple closed states as well as inactive and/or desensitized states, which are non-conducting states that can occur even in the presence of stimulus.[9]
Biomolecule labeling
Single fluorophores can be chemically attached to biomolecules, such as proteins or DNA, and the dynamics of individual molecules can be tracked by monitoring the fluorescent probe. Spatial movements within the Rayleigh limit can be tracked, along with changes in emission intensity and/or radiative lifetime, which often indicate changes in local environment. For instance, single-molecule labeling has yielded a vast quantity of information on how kinesin motor proteins move along microtubule strands in muscle cells. Single-molecule imaging in live cells reveals interesting information about protein dynamics under its physiological environment. Several biophysical parameters about protein dynamics can be quantified such as diffusion coefficient, mean squared displacements, residence time, the fraction of bound and unbound molecules, and target-search mechanism of protein binding to its target site in the live cell.[10]
Single-molecule fluorescence resonance energy transfer (FRET)
Main article smFRET.
In single-molecule fluorescence resonance energy transfer, the molecule is labeled in (at least) two places. A laser beam is focused on the molecule exciting the first probe. When this probe relaxes and emits a photon, it has a chance of exciting the other probe. The efficiency of the absorption of the photon emitted from the first probe in the second probe depends on the distance between these probes. Since the distance changes with time, this experiment probes the internal dynamics of the molecule.
Single-molecule experiments versus ensemble experiments
When looking at data related to individual molecules, one usually can construct propagators, and jumping time probability density functions, of the first order, the second order and so on, whereas from bulk experiments, one usually obtains the decay of a correlation function.[11] From the information contained in these unique functions (obtained from individual molecules), one can extract a relatively clear picture on the way the system behaves; e.g. its kinetic scheme, or its potential of activity, or its reduced dimensions form.[12][13] In particular, one can construct (many properties of) the reaction pathway of an enzyme when monitoring the activity of an individual enzyme.[14] Additionally, significant aspects regarding the analysis of single molecule data—such as fitting methods and tests for homogeneous populations—have been described by several authors.[9] On the other hand, there are several issues with the analysis of single molecule data including construction of a low noise environment and insulated pipet tips, filtering some of the remaining unwanted components (noise) found in recordings, and the length of time required for data analysis (pre-processing, unambiguous event detection, plotting data, fitting kinetic schemes, etc.).
Impact
Single-molecule techniques impacted optics, electronics, biology, and chemistry. In the biological sciences, the study of proteins and other complex biological machinery was limited to ensemble experiments that nearly made impossible the direct observation of their kinetics. For example, it was only after single molecule fluorescence microscopy was used to study kinesin-myosin pairs in muscle tissue that direct observation of the walking mechanisms were understood. These experiments, however, have for the most part been limited to in vitro studies, as useful techniques for live cell imaging have yet to be fully realized. The promise of single molecule in vivo imaging,[15] however, brings with it an enormous potential to directly observe bio-molecules in native processes. These techniques are often targeted for studies involving low-copy proteins, many of which are still being discovered. These techniques have also been extended to study areas of chemistry, including the mapping of heterogeneous surfaces.[16]
See also
- Single-molecule magnet
- Force spectroscopy
- Magnetic tweezers
- Optical tweezers
- Single-particle tracking
- Raman spectroscopy
- Scanning probe microscopy
- Electron microscopy
- Tethered particle motion (TPM)
- Super-resolution microscopy
- Voltage clamp
- Tunable resistive pulse sensing
- Single molecule real time sequencing
References
- Roiter V, Minko S (2005). "AFM Single Molecule Experiments at the Solid-Liquid Interface: In Situ Conformation of Adsorbed Flexible Polyelectrolyte Chains". Journal of the American Chemical Society. 127 (45): 15688–15689. doi:10.1021/ja0558239. PMID 16277495.
- Juette MF, Terry DS, Wasserman MR, Zhou Z, Altman RB, Zheng Q, Blanchard SC (June 2014). "The bright future of single-molecule fluorescence imaging". Curr Opin Chem Biol. 20: 103–111. doi:10.1016/j.cbpa.2014.05.010. PMC 4123530. PMID 24956235.
- W. E. Moerner and L. Kador, Optical detection and spectroscopy of single molecules in a solid, Phys. Rev. Lett. 62, 2535 - 2538 (1989)
- M. Orrit and J. Bernard, Single pentacene molecules detected by fluorescence excitation in a p-terphenyl crystal, Phys. Rev. Lett. 65, 2716–2719 (1990)
- Kneipp, Katrin; Wang, Yang; Kneipp, Harald; Perelman, Lev T.; Itzkan, Irving; Dasari, Ramachandra R.; Feld, Michael S. (1997-03-03). "Single Molecule Detection Using Surface-Enhanced Raman Scattering (SERS)". Physical Review Letters. 78 (9): 1667–1670. doi:10.1103/PhysRevLett.78.1667. ISSN 0031-9007.
- Nie, Shuming; Emory, Steven R. (1997-02-21). "Probing Single Molecules and Single Nanoparticles by Surface-Enhanced Raman Scattering". Science. 275 (5303): 1102–1106. doi:10.1126/science.275.5303.1102. ISSN 0036-8075.
- Murugesapillai D, et al. (2014). "DNA bridging and looping by HMO1 provides a mechanism for stabilizing nucleosome-free chromatin". Nucleic Acids Res. 42 (14): 8996–9004. doi:10.1093/nar/gku635. PMC 4132745. PMID 25063301.
- Murugesapillai D, et al. (2016). "Single-molecule studies of high-mobility group B architectural DNA bending proteins". Biophys Rev. 9 (1): 17–40. doi:10.1007/s12551-016-0236-4. PMC 5331113. PMID 28303166.
- B. Sakmann and E. Neher, Single-Channel Recording, ISBN 9780306414190 (1995).
- Podh, Nitesh Kumar; Paliwal, Sheetal; Dey, Partha; Das, Ayan; Morjaria, Shruti; Mehta, Gunjan (5 November 2021). "In-vivo Single-Molecule Imaging in Yeast: Applications and Challenges". Journal of Molecular Biology. 433 (22): 167250. doi:10.1016/j.jmb.2021.167250. PMID 34537238. S2CID 237573437.
- O. Flomenbom, J. Klafter, and A. Szabo, What can one learn from two-state single molecule trajectories? Archived January 14, 2012, at the Wayback Machine, Biophys. J. 88, 3780–3783 (2005); arXiv:q-bio/0502006
- O. Flomenbom, and R. J. Silbey, Utilizing the information content in two-state trajectories Archived January 14, 2012, at the Wayback Machine, Proc. Natl. Acad. Sci. USA 103, 10907–10910 (2006).
- O. Flomenbom, and R. J. Silbey, Toolbox for analyzing finite two-state trajectories, Phys. Rev. E 78, 066105 (2008); arXiv:0802.1520.
- O. Flomenbom, K. Velonia, D. Loos, et al., Stretched exponential decay and correlations in the catalytic activity of fluctuating single lipase molecules Archived January 14, 2012, at the Wayback Machine, Proc. Natl. Acad. Sci. US 102, 2368–2372 (2005).
- Zhan, Hong; Stanciauskas, Ramunas; Stigloher, Christian; Keomanee-Dizon, Kevin; Jospin, Maelle; Bessereau, Jean-Louis; Pinaud, Fabien (2014). "In vivo single-molecule imaging identifies altered dynamics of calcium channels in dystrophin-mutant C. elegans". Nature Communications. 5: ncomms5974. Bibcode:2014NatCo...5.4974Z. doi:10.1038/ncomms5974. PMC 4199201. PMID 25232639.
- Walder, R.; Nelson, N.; Schwartz, D. K. (2011). "Super-resolution surface mapping using the trajectories of molecular probes". Nature Communications. 2: 515. Bibcode:2011NatCo...2..515W. doi:10.1038/ncomms1530. PMID 22044994.