XPO5
Exportin-5 (XPO5) is a protein that, in humans, is encoded by the XPO5 gene.[5][6][7] In eukaryotic cells, the primary purpose of XPO5 is to export pre-microRNA (also known as pre-miRNA) out of the nucleus and into the cytoplasm, for further processing by the Dicer enzyme.[8][9][10][11] Once in the cytoplasm, the microRNA (also known as miRNA) can act as a gene silencer by regulating translation of mRNA. Although XPO5 is primarily involved in the transport of pre-miRNA, it has also been reported to transport tRNA.[12]
| XPO5 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | XPO5, exp5, exportin 5 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 607845 MGI: 1913789 HomoloGene: 69316 GeneCards: XPO5 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Much research on XPO5 is ongoing. miRNA is a prominent research topic due to its potential use as a therapeutic, with several miRNA-based drugs already in use.[13]
Mechanism
Binding to pre-miRNA
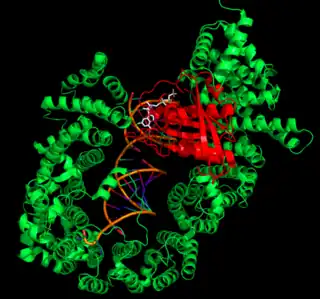
After RanGTP binds to XPO5, the XPO5-RanGTP complex forms a U-like structure to hold the pre-miRNA. The XPO5-RanGTP complex recognizes pre-miRNA by its two-nucleotide 3’ overhang—a sequence consisting of two bases at the 3’ end of the pre-miRNA that are not paired with other bases. This motif is unique to pre-miRNA, and by recognizing it XPO5 ensures specificity for transporting only pre-miRNA. On its own, pre-miRNA is in a “closed” conformation, with the 3’ overhang flipped up toward the RNA minor groove. However, upon binding to XPO5, the 3’ overhang is flipped downwards away from the rest of the pre-miRNA molecule into an “open” conformation. This helps the backbone phosphates of these two nucleotides form hydrogen bonds with many XPO5 residues, allowing XPO5 to recognize the RNA as pre-miRNA. Because these interactions involve only the RNA phosphate backbone, they are nonspecific and allow XPO5 to recognize and transport any pre-miRNA. The rest of the pre-miRNA stem binds to XPO5 via interactions between the negatively-charged phosphate backbone and several positively-charged interior XPO5 residues.[15]
XPO5 Ternary Complex Transport Mechanism
The combined structure of XPO5, RanGTP, and pre-miRNA is known as the ternary complex. Once the ternary complex is formed, it diffuses through a nuclear pore complex into the cytoplasm, transporting pre-miRNA into the cytoplasm in the process. Once in the cytoplasm, RanGAP hydrolyzes GTP to GDP, causing a conformational change that releases the pre-miRNA into the cytoplasm.[15]
Export out of the Nucleus
It has been suggested, through evidence provided by contour maps of water density, that the interior of XPO5 is hydrophilic, while the exterior of XPO5 is hydrophobic.[15] Therefore, this enhances the binding capabilities of XPO5 to the nuclear pore complex, allowing for transport of the ternary complex out of the nucleus.[15]
Potential oncogenic role
Recent evidence has shown higher levels of XPO5 in prostate cancer cell lines in-vitro, suggesting that altered XPO5 expression levels may have a role in cancer development. Suppressing XPO5 has also been found to be therapeutic in-vitro.[16] It has also been shown to function as an oncogene in colorectal cancer.[17]
References
- GRCh38: Ensembl release 89: ENSG00000124571 - Ensembl, May 2017
- GRCm38: Ensembl release 89: ENSMUSG00000067150 - Ensembl, May 2017
- "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- Brownawell AM, Macara IG (Jan 2002). "Exportin-5, a novel karyopherin, mediates nuclear export of double-stranded RNA binding proteins". The Journal of Cell Biology. 156 (1): 53–64. doi:10.1083/jcb.200110082. PMC 2173575. PMID 11777942.
- Bohnsack MT, Regener K, Schwappach B, Saffrich R, Paraskeva E, Hartmann E, Görlich D (Nov 2002). "Exp5 exports eEF1A via tRNA from nuclei and synergizes with other transport pathways to confine translation to the cytoplasm". The EMBO Journal. 21 (22): 6205–15. doi:10.1093/emboj/cdf613. PMC 137205. PMID 12426392.
- "Entrez Gene: XPO5 exportin 5".
- Yi R, Qin Y, Macara IG, Cullen BR (Dec 2003). "Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs". Genes & Development. 17 (24): 3011–6. doi:10.1101/gad.1158803. PMC 305252. PMID 14681208.
- Wilson RC, Doudna JA (2013). "Molecular mechanisms of RNA interference". Annual Review of Biophysics. 42: 217–39. doi:10.1146/annurev-biophys-083012-130404. PMC 5895182. PMID 23654304.
- Siomi H, Siomi MC (May 2010). "Posttranscriptional regulation of microRNA biogenesis in animals". Molecular Cell. 38 (3): 323–32. doi:10.1016/j.molcel.2010.03.013. PMID 20471939.
- Macias S, Cordiner RA, Cáceres JF (Aug 2013). "Cellular functions of the microprocessor". Biochemical Society Transactions. 41 (4): 838–43. doi:10.1042/BST20130011. hdl:1842/25877. PMID 23863141.
- Gupta, Asmita (2016). "Insights into the Structural Dynamics of Nucleocytoplasmic Transport of tRNA by Exportin-t". Biophysical Journal. 110 (6): 1264–1279. Bibcode:2016BpJ...110.1264G. doi:10.1016/j.bpj.2016.02.015. PMC 4816717. PMID 27028637.
- Christopher, Ajay Francis; Kaur, Raman Preet; Kaur, Gunpreet; Kaur, Amandeep; Gupta, Vikas; Bansal, Parveen (2016). "MicroRNA therapeutics: Discovering novel targets and developing specific therapy". Perspectives in Clinical Research. 7 (2): 68–74. doi:10.4103/2229-3485.179431. ISSN 2229-3485. PMC 4840794. PMID 27141472.
- Okada, Chimari; Yamashita, Eiki; Lee, Soo Jae; Shibata, Satoshi; Katahira, Jun; Nakagawa, Atsushi; Yoneda, Yoshihiro; Tsukihara, Tomitake (2009-11-27). "A high-resolution structure of the pre-microRNA nuclear export machinery". Science. 326 (5957): 1275–1279. Bibcode:2009Sci...326.1275O. doi:10.1126/science.1178705. ISSN 1095-9203. PMID 19965479. S2CID 206522317.
- Wang, Xia (2011). "Dynamic mechanisms for pre-miRNA binding and export by Exportin-5". RNA. 17 (8): 1516–1517. doi:10.1261/rna.2732611. PMC 3153975. PMID 21712399.
- Höti, Naseruddin; Yang, Shuang; Aiyetan, Paul; Kumar, Binod; Hu, Yingwei; Clark, David; Eroglu, Arife Unal; Shah, Punit; Johnson, Tamara (2017-09-04). "Overexpression of Exportin-5 Overrides the Inhibitory Effect of miRNAs Regulation Control and Stabilize Proteins via Posttranslation Modifications in Prostate Cancer". Neoplasia (New York, N.Y.). 19 (10): 817–829. doi:10.1016/j.neo.2017.07.008. ISSN 1522-8002. PMC 5587889. PMID 28881308.
- Shigeyasu, Kunitoshi; Okugawa, Yoshinaga; Toden, Shusuke; Boland, C. Richard; Goel, Ajay (2017-03-01). "Exportin-5 Functions as an Oncogene and a Potential Therapeutic Target in Colorectal Cancer". Clinical Cancer Research. 23 (5): 1312–1322. doi:10.1158/1078-0432.CCR-16-1023. ISSN 1557-3265. PMC 5435115. PMID 27553833.
Further reading
- Calado A, Treichel N, Müller EC, Otto A, Kutay U (Nov 2002). "Exportin-5-mediated nuclear export of eukaryotic elongation factor 1A and tRNA". The EMBO Journal. 21 (22): 6216–24. doi:10.1093/emboj/cdf620. PMC 137209. PMID 12426393.
- Gwizdek C, Ossareh-Nazari B, Brownawell AM, Doglio A, Bertrand E, Macara IG, Dargemont C (Feb 2003). "Exportin-5 mediates nuclear export of minihelix-containing RNAs". The Journal of Biological Chemistry. 278 (8): 5505–8. doi:10.1074/jbc.C200668200. PMID 12509441.
- Gwizdek C, Ossareh-Nazari B, Brownawell AM, Evers S, Macara IG, Dargemont C (Jan 2004). "Minihelix-containing RNAs mediate exportin-5-dependent nuclear export of the double-stranded RNA-binding protein ILF3". The Journal of Biological Chemistry. 279 (2): 884–91. doi:10.1074/jbc.M306808200. PMID 14570900.
- Lund E, Güttinger S, Calado A, Dahlberg JE, Kutay U (Jan 2004). "Nuclear export of microRNA precursors". Science. 303 (5654): 95–8. Bibcode:2004Sci...303...95L. doi:10.1126/science.1090599. PMID 14631048. S2CID 30217099.
- Bohnsack MT, Czaplinski K, Gorlich D (Feb 2004). "Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs". RNA. 10 (2): 185–91. doi:10.1261/rna.5167604. PMC 1370530. PMID 14730017.
- Chen T, Brownawell AM, Macara IG (Aug 2004). "Nucleocytoplasmic shuttling of JAZ, a new cargo protein for exportin-5". Molecular and Cellular Biology. 24 (15): 6608–19. doi:10.1128/MCB.24.15.6608-6619.2004. PMC 444848. PMID 15254228.
- Yi R, Doehle BP, Qin Y, Macara IG, Cullen BR (Feb 2005). "Overexpression of exportin 5 enhances RNA interference mediated by short hairpin RNAs and microRNAs". RNA. 11 (2): 220–6. doi:10.1261/rna.7233305. PMC 1370710. PMID 15613540.