YjdF RNA motif
The yjdF RNA motif is a conserved RNA structure identified using bioinformatics.[1] Most yjdF RNAs are located in bacteria classified within the phylum Bacillota. A yjdF RNA is found in the presumed 5' untranslated region (5' UTR) of the yjdF gene in Bacillus subtilis, and almost all yjdF RNAs are found in the 5' UTRs of homologs of this gene. The function of the yjdF gene is unknown, but the protein that it is predicted to encode is classified by the Pfam Database as DUF2992.[2]
| Protein of unknown function (DUF2992) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | DUF2992 | ||||||||
| Pfam | PF11208 | ||||||||
| |||||||||
| yjdF RNA | |
|---|---|
| Identifiers | |
| Symbol | yjdF |
| Rfam | RF01764 |
| Other data | |
| RNA type | Cis-reg; |
| Domain(s) | Bacteria; |
| SO | SO:0005836 |
| PDB structures | PDBe |
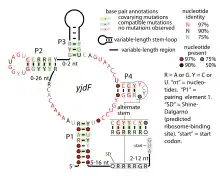
The moderately complex secondary structure and nucleotide positions conserved in the yjdF RNA motif and the fact that it is likely to be present in 5' UTRs led to the hypothesis that yjdF RNAs function as riboswitches. One yjdF RNA is found instead in the presumed 5' UTR of genes encoding enzymes involved in the synthesis of nicotinamide adenine dinucleotide (NAD+). These RNAs lack a region of the otherwise conserved yjdF RNA motif and might be non-functional, or conform to a distinct structure that achieves the same function. If yjdF RNAs are riboswitches, this other RNA might bind a distinct small molecule ligand.
Experiments support the conclusion that the yjdF RNA in B. subtilis is indeed in the 5' UTR of the yjdF gene, and that the yjdF genes are transcribed separately from the upstream manPA operon.[1] The biochemical function of yjdF RNAs remains unknown.
yjdF RNAs appear to function as riboswitches that sense azaaromatic compounds, although the precise compound or set of compounds that is sensed by this riboswitch in the cell remains unclear.[3]
References
- Weinberg Z, Wang JX, Bogue J, et al. (March 2010). "Comparative genomics reveals 104 candidate structured RNAs from bacteria, archaea and their metagenomes". Genome Biol. 11 (3): R31. doi:10.1186/gb-2010-11-3-r31. PMC 2864571. PMID 20230605.
- "DUF2992 (PF11208)". InterPro.
- Li S, Hwang XY, Stav S, Breaker RR (2016). "The yjdF riboswitch candidate regulates gene expression by binding diverse azaaromatic compounds". RNA. 22 (4): 530–41. doi:10.1261/rna.054890.115. PMC 4793209. PMID 26843526.