Caulobacter crescentus
Caulobacter crescentus is a Gram-negative, oligotrophic bacterium widely distributed in fresh water lakes and streams. The taxon is more properly known as Caulobacter vibrioides (Henrici and Johnson 1935).[1]
| Caulobacter crescentus | |
|---|---|
 | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Alphaproteobacteria |
| Order: | Caulobacterales |
| Family: | Caulobacteraceae |
| Genus: | Caulobacter |
| Species: | C. crescentus |
| Binomial name | |
| Caulobacter crescentus Poindexter 1964 | |
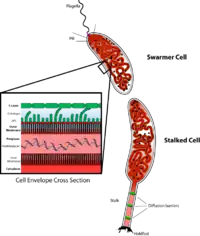
C. crescentus is an important model organism for studying the regulation of the cell cycle, asymmetric cell division, and cellular differentiation. Caulobacter daughter cells have two very different forms. One daughter is a mobile "swarmer" cell that has a single flagellum at one cell pole that provides swimming motility for chemotaxis. The other daughter, called the "stalked" cell, has a tubular stalk structure protruding from one pole that has an adhesive holdfast material on its end, with which the stalked cell can adhere to surfaces. Swarmer cells differentiate into stalked cells after a short period of motility. Chromosome replication and cell division only occurs in the stalked cell stage.
C. crescentus derives its name from its crescent shape, which is caused by the protein crescentin. It is an interesting organism to study because it inhabits nutrient-poor aquatic environments. Their ability to thrive in low levels of nutrients is facilitated by its dimorphic developmental cycle. The swarmer cell has a flagellum that protrudes from a single pole and is unable to initiate DNA replication unless differentiated into a stalked cell. The differentiation process includes a morphological transition characterized by ejection of its flagellum and growth of a stalk at the same pole. Stalked cells can elongate and replicate their DNA while growing a flagellum at the opposite pole, giving rise to a pre-divisional cell. Although the precise function of stalks is still being investigated, it is likely that the stalks are involved in the uptake of nutrients in nutrient-limited conditions.[2] Its use as a model originated with developmental biologist Lucy Shapiro.[3][4]
Strains
In the laboratory, researchers distinguish between C. crescentus strain CB15 (the strain originally isolated from a freshwater lake) and NA1000 (the primary experimental strain). In strain NA1000, which was derived from CB15 in the 1970s,[5] the stalked and predivisional cells can be physically separated in the laboratory from new swarmer cells, while cell types from strain CB15 cannot be physically separated. The isolated swarmer cells can then be grown as a synchronized cell culture. Detailed study of the molecular development of these cells as they progress through the cell cycle has enabled researchers to understand Caulobacter cell cycle regulation in great detail. Due to this capacity to be physically synchronized, strain NA1000 has become the predominant experimental Caulobacter strain throughout the world. Additional phenotypic differences between the two strains have subsequently accumulated due to selective pressures on the NA1000 strain in the laboratory environment. The genetic basis of the phenotypic differences between the two strains results from coding, regulatory, and insertion/deletion polymorphisms at five chromosomal loci.[6] C. crescentus is synonymous with Caulobacter vibrioides.[1]
Genomics
The Caulobacter CB15 genome has 4,016,942 base pairs in a single circular chromosome encoding 3,767 genes.[7] The genome contains multiple clusters of genes encoding proteins essential for survival in a nutrient-poor habitat. Included are those involved in chemotaxis, outer membrane channel function, degradation of aromatic ring compounds, and the breakdown of plant-derived carbon sources, in addition to many extracytoplasmic function sigma factors, providing the organism with the ability to respond to a wide range of environmental fluctuations. In 2010, the Caulobacter NA1000 strain was sequenced and all differences with the CB15 "wild type" strain were identified.[6]
Role of the swarmer cell stage
The Caulobacter stalked cell stage provides a fitness advantage by anchoring the cell to surfaces to form biofilms and or to exploit nutrient sources. Generally, the bacterial species that divides fastest will be most effective at exploiting resources and effectively occupying ecological niches. Yet, Caulobacter has the swarmer cell stage that results in slower population growth. The swarmer cell is thought to provide cell dispersal, so that the organism constantly seeks out new environments. This may be particularly useful in severely nutrient-limited environments when the scant resources available can be depleted very quickly. Many, perhaps most, of the swarmer daughter cells will not find a productive environment, but the obligate dispersal stage must increase the reproductive fitness of the species as a whole.
Cell cycle
The Caulobacter cell cycle regulatory system controls many modular subsystems that organize the progression of cell growth and reproduction. A control system constructed using biochemical and genetic logic circuitry organizes the timing of initiation of each of these subsystems. The central feature of the cell cycle regulation is a cyclical genetic circuit—a cell cycle engine—that is centered around the successive interactions of five master regulatory proteins: DnaA, GcrA, CtrA, SciP, and CcrM whose roles were worked out by the laboratories of Lucy Shapiro and Harley McAdams.[8][9][10] These five proteins directly control the timing of expression of over 200 genes. The five master regulatory proteins are synthesized and then eliminated from the cell one after the other over the course of the cell cycle. Several additional cell signaling pathways are also essential to the proper functioning of this cell cycle engine. The principal role of these signaling pathways is to ensure reliable production and elimination of the CtrA protein from the cell at just the right times in the cell cycle.
An essential feature of the Caulobacter cell cycle is that the chromosome is replicated once and only once per cell cycle. This is in contrast to the E. coli cell cycle where there can be overlapping rounds of chromosome replication simultaneously underway. The opposing roles of the Caulobacter DnaA and CtrA proteins are essential to the tight control of Caulobacter chromosome replication.[11] The DnaA protein acts at the origin of replication to initiate the replication of the chromosome. The CtrA protein, in contrast, acts to block initiation of replication, so it must be removed from the cell before chromosome replication can begin. Multiple additional regulatory pathways integral to cell cycle regulation and involving both phospho signaling pathways and regulated control of protein proteolysis[12] act to assure that DnaA and CtrA are present in the cell just exactly when needed.
Each process activated by the proteins of the cell cycle engine involve a cascade of many reactions. The longest subsystem cascade is DNA replication. In Caulobacter cells, replication of the chromosome involves about 2 million DNA synthesis reactions for each arm of the chromosome over 40 to 80 min depending on conditions. While the average time for each individual synthesis reaction can be estimated from the observed average total time to replicate the chromosome, the actual reaction time for each reaction varies widely around the average rate. This leads to a significant and inevitable cell-to-cell variation time to complete replication of the chromosome. There is similar random variation in the rates of progression of all the other subsystem reaction cascades. The net effect is that the time to complete the cell cycle varies widely over the cells in a population even when they all are growing in identical environmental conditions. Cell cycle regulation includes feedback signals that pace progression of the cell cycle engine to match progress of events at the regulatory subsystem level in each particular cell. This control system organization, with a controller (the cell cycle engine) driving a complex system, with modulation by feedback signals from the controlled system creates a closed loop control system.
The rate of progression of the cell cycle is further adjusted by additional signals arising from cellular sensors that monitor environmental conditions (for example, nutrient levels and the oxygen level) or the internal cell status (for example, presence of DNA damage).[13]
Evolutionary conservation of the cell cycle control system
The control circuitry that directs and paces Caulobacter cell cycle progression involves the entire cell operating as an integrated system. The control circuitry monitors the environment and the internal state of the cell, including the cell topology, as it orchestrates activation of cell cycle subsystems and Caulobacter crescentus asymmetric cell division. The proteins of the Caulobacter cell cycle control system and its internal organization are co-conserved across many alphaproteobacteria species, but there are great differences in the regulatory apparatus' functionality and peripheral connectivity to other cellular subsystems from species to species.[14][15] The Caulobacter cell cycle control system has been exquisitely optimized by evolutionary selection as a total system for robust operation in the face of internal stochastic noise and environmental uncertainty.
The bacterial cell's control system has a hierarchical organization.[16] The signaling and the control subsystem interfaces with the environment by means of sensory modules largely located on the cell surface. The genetic network logic responds to signals received from the environment and from internal cell status sensors to adapt the cell to current conditions. A major function of the top level control is to ensure that the operations involved in the cell cycle occur in the proper temporal order. In Caulobacter, this is accomplished by the genetic regulatory circuit composed of five master regulators and an associated phospho-signaling network. The phosphosignaling network monitors the state of progression of the cell cycle and plays an essential role in accomplishing asymmetric cell division. The cell cycle control system manages the time and place of the initiation of chromosome replication and cytokinesis as well as the development of polar organelles. Underlying all these operations are the mechanisms for production of protein and structural components and energy production. The “housekeeping” metabolic and catabolic subsystems provide the energy and the molecular raw materials for protein synthesis, cell wall construction and other operations of the cell. The housekeeping functions are coupled bidirectionally to the cell cycle control system. However, they can adapt, somewhat independently of the cell cycle control logic, to changing composition and levels of the available nutrient sources.
The proteins of the Caulobacter cell cycle control system are widely co-conserved across the alphaproteobacteria, but the ultimate function of this regulatory system varies widely in different species. These evolutionary changes reflect enormous differences between the individual species in fitness strategies and ecological niches. For example, Agrobacterium tumefaciens is a plant pathogen, Brucella abortus is an animal pathogen, and Sinorhizobium meliloti is a soil bacterium that invades, and becomes a symbiont in, plant root nodules that fix nitrogen yet most of the proteins of the Caulobacter cell cycle control are also found in these species. The specific coupling between the protein components of the cell cycle control network and the downstream readout of the circuit differ from species to species. The pattern is that the internal functionality of the network circuitry is conserved, but the coupling at the “edges” of the regulatory apparatus to the proteins controlling specific cellular functions differs widely among the different species.
The evolution of stalk positioning in the Caulobacter clade

Caulobacter crescentus is a member of a group of bacteria that possess the stalk structure, a tubular extension from the cell body. However, the positioning of the stalk is not necessarily conserved at the pole of the cell body in different closely related species. Specifically, research has shown that not only the position of the stalk can change, but the number can vary as well in the closely related genus Asticcacaulis.[17][18] SpmX, a polarly localized protein in Caulobacter crescentus, has been shown to manipulate stalk positioning in these Asticcacaulis species.[17] Presumably, It does so by a gain of function after protein expansion from around 400 amino acids in Caulobacter crescentus to more than 800 amino acids in Asticcacaulis species.
Caulobacter aging
Caulobacter was the first asymmetric bacterium shown to age. Reproductive senescence was measured as the decline in the number of progeny produced over time.[19][20] On the basis of experimental evolution studies in C. crescentus, Ackermann et al.[19] suggested that aging is probably a fundamental property of all cellular organisms. A similar phenomenon has since been described in the bacterium Escherichia coli, which gives rise to morphologically similar daughter cells.[21]
Cell polarity regulation
In C. crescentus, cell polarity is readily apparent by the assembly of polar organelles and by the polarization of the division plane, which results in the generation of stalked progeny that are longer than swarmer progeny. The formation of new cell poles at division implies that cell polarity must be re-established in the stalked progeny and reversed in the swarmer progeny.[22]
The C. crescentus life cycle is governed by regulators such as TipN, a cell cycle protein. Yale University's data strongly suggest a model in which TipN regulates the orientation of the polarity axis by providing a positional cue from the preceding cell cycle. In this model TipN specifies the site of the most recent division by identifying the new pole. The cell uses this positional information as a source of intracellular asymmetry to establish and maintain the orientation of the polarity axis, which is crucial for polar morphogenesis and division. Recruitment of TipN to the nascent poles at the end of the division cycle redefines the identity of the poles and resets the correct polarity in both future daughter cells (with a polarity reversal in the swarmer cell).[22] The cell cycle–regulated synthesis and removal of these polarly localized structures have provided a rich playground for the identification of landmark proteins important for their proper localization.[23] TipN has two transmembrane regions in the N-terminal region and a large C-terminal coiled-coil domain. TipN homologues are present in other alpha-proteobacteria. TipN localizes to the new pole in both daughter cells after division and relocalizes to the cell division site in the late predivisional cell. Therefore, both daughter cells have TipN at the new pole after division.[23]
The landmark protein TipN is essential for the proper placement of the flagellum. [24] Mutants lacking TipN make serious mistakes in development. Instead of making a single flagellum at the correct cell pole , the cell makes multiple flagella at various locations, even on the stalk.[22]
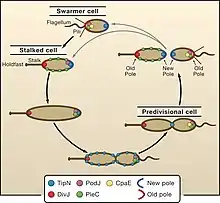
Cell development involves many such proteins working together. Fig#1 shows how TipN interact with two other polar proteins : the flagellar marker PodJ , and the stalk marker DivJ. [25]
References
- Abraham, Wolf-Rainer; Carsten Strömpl; Holger Meyer; Sabine Lindholst; Edward R. B. Moore; Ruprecht Christ; Marc Vancanneyt; B. J. Tindali; Antonio Bennasar; John Smit; Michael Tesar (1999). "Phylogeny and polyphasic taxonomy of Caulobacter species. Proposal of Maricaulis gen. nov. with Maricaulis maris (Poindexter) comb. nov. as the type species, and emended description of the genera Brevundirnonas and Caulobacter". International Journal of Systematic Bacteriology. 49 (3): 1053–73. doi:10.1099/00207713-49-3-1053. PMID 10425763.
- Ausmees, Nora; Kuhn, Jeffrey R.; Jacobs-Wagner, Christine (December 2003). "The bacterial cytoskeleton: an intermediate filament-like function in cell shape". Cell. 115 (6): 705–13. doi:10.1016/S0092-8674(03)00935-8. PMID 14675535. S2CID 14459851.
- Conger, Krista (March 31, 2009). "Top Canadian Prize Goes to Stanford Scientist Lucy Shapiro for Bringing Cell Biology into Three Dimensions". Business Wire. Retrieved 14 May 2015.
- "2014 Lucy Shapiro". Greengard Prize. 2014. Retrieved 14 May 2015.
- Poindexter, JS (Sep 1964). "Biological Properties and Classification of the Caulobacter Group". Microbiol. Mol. Biol. Rev. 28 (3): 231–95. doi:10.1128/mmbr.28.3.231-295.1964. PMC 441226. PMID 14220656.
- Marks ME; Castro-Rojas CM; Teiling C; et al. (July 2010). "The Genetic Basis of Laboratory Adaptation in Caulobacter crescentus". J. Bacteriol. 192 (14): 3678–88. doi:10.1128/JB.00255-10. PMC 2897358. PMID 20472802.
- Nierman, WC; Feldblyum, TV; Laub, MT; Paulsen, IT; Nelson, KE; Eisen, JA; Heidelberg, JF; Alley, MR; Ohta, N; Maddock, JR; Potocka, I; Nelson, WC; Newton, A; Stephens, C; Phadke, ND; Ely, B; DeBoy, RT; Dodson, RJ; Durkin, AS; Gwinn, ML; Haft, DH; Kolonay, JF; Smit, J; Craven, MB; Khouri, H; Shetty, J; Berry, K; Utterback, T; Tran, K; Wolf, A; Vamathevan, J; Ermolaeva, M; White, O; Salzberg, SL; Venter, JC; Shapiro, L; Fraser, CM (Mar 27, 2001). "Complete genome sequence of Caulobacter crescentus". Proceedings of the National Academy of Sciences of the United States of America. 98 (7): 4136–41. Bibcode:2001PNAS...98.4136N. doi:10.1073/pnas.061029298. PMC 31192. PMID 11259647.
- McAdams, HH; Shapiro, L (Dec 17, 2009). "System-level design of bacterial cell cycle control". FEBS Letters. 583 (24): 3984–91. doi:10.1016/j.febslet.2009.09.030. PMC 2795017. PMID 19766635.
- Collier, J; Shapiro, L (Aug 2007). "Spatial complexity and control of a bacterial cell cycle". Current Opinion in Biotechnology. 18 (4): 333–40. doi:10.1016/j.copbio.2007.07.007. PMC 2716793. PMID 17709236.
- Tan, M. H.; Kozdon, J. B.; Shen, X.; Shapiro, L.; McAdams, H. H. (2010). "An essential transcription factor, SciP, enhances robustness of Caulobacter cell cycle regulation". Proceedings of the National Academy of Sciences. 107 (44): 18985–990. Bibcode:2010PNAS..10718985T. doi:10.1073/pnas.1014395107. PMC 2973855. PMID 20956288.
- Collier, J; Murray, SR; Shapiro, L (Jan 25, 2006). "DnaA couples DNA replication and the expression of two cell cycle master regulators". The EMBO Journal. 25 (2): 346–56. doi:10.1038/sj.emboj.7600927. PMC 1383511. PMID 16395331.
- Jenal, U (Nov 2009). "The role of proteolysis in the Caulobacter crescentus cell cycle and development". Research in Microbiology. 160 (9): 687–95. doi:10.1016/j.resmic.2009.09.006. PMID 19781638.
- Shen, X; Collier, J; Dill, D; Shapiro, L; Horowitz, M; McAdams, HH (Aug 12, 2008). "Architecture and inherent robustness of a bacterial cell-cycle control system". Proceedings of the National Academy of Sciences of the United States of America. 105 (32): 11340–45. Bibcode:2008PNAS..10511340S. doi:10.1073/pnas.0805258105. PMC 2516238. PMID 18685108.
- McAdams, Harley H.; Shapiro, Lucy (2011). "The Architecture and Conservation Pattern of Whole-Cell Control Circuitry". Journal of Molecular Biology. 409 (1): 28–35. doi:10.1016/j.jmb.2011.02.041. PMC 3108490. PMID 21371478.
- Brilli, Matteo; Fondi, Marco; Fani, Renato; Mengoni, Alessio; Ferri, Lorenzo; Bazzicalupo, Marco; Biondi, Emanuele G. (2010). "The diversity and evolution of cell cycle regulation in alpha-proteobacteria: A comparative genomic analysis". BMC Systems Biology. 4: 52. doi:10.1186/1752-0509-4-52. PMC 2877005. PMID 20426835.
- McAdams, HH; Shapiro, L. (May 2011). "The architecture and conservation pattern of whole-cell control circuitry". J Mol Biol. 409 (1): 28–35. doi:10.1016/j.jmb.2011.02.041. PMC 3108490. PMID 21371478.
- Jiang, Chao; Brown, Pamela J.B.; Ducret, Adrien; Brun1, Yves V. (2014-02-27). "Sequential evolution of bacterial morphology by co-option of a developmental regulator". Nature. 506 (7489): 489–93. Bibcode:2014Natur.506..489J. doi:10.1038/nature12900. ISSN 0028-0836. PMC 4035126. PMID 24463524.
- Jiang, Chao; Caccamo, Paul D.; Brun, Yves V. (April 2015). "Mechanisms of bacterial morphogenesis: evolutionary cell biology approaches provide new insights". BioEssays. 37 (4): 413–25. doi:10.1002/bies.201400098. ISSN 1521-1878. PMC 4368449. PMID 25664446.
- Ackermann, Martin; Stephen C. Stearns; Urs Jenal (2003). "Senescence in a bacterium with asymmetric division". Science. 300 (5627): 1920. doi:10.1126/science.1083532. PMID 12817142. S2CID 34770745.
- Ackermann, Martin; Alexandra Schauerte; Stephen C. Stearns; Urs Jenal (2007). "Experimental evolution of aging in a bacterium". BMC Evolutionary Biology. 7: 126. doi:10.1186/1471-2148-7-126. PMC 2174458. PMID 17662151.
- Stewart, Eric J.; Richard Madden; Gregory Paul; Francois Taddei (2005). "Aging and Death in an Organism That Reproduces by Morphologically Symmetric Division". PLOS Biology. 3 (2): e45. doi:10.1371/journal.pbio.0030045. PMC 546039. PMID 15685293.
- H, Lam; Wb, Schofield; C, Jacobs-Wagner (2006-03-10). "A Landmark Protein Essential for Establishing and Perpetuating the Polarity of a Bacterial Cell". Cell. 124 (5): 1011–23. doi:10.1016/j.cell.2005.12.040. PMID 16530047. S2CID 14200442.
- Treuner-Lange, Anke; Søgaard-Andersen, Lotte (2014-07-07). "Regulation of cell polarity in bacteria". Journal of Cell Biology. 206 (1): 7–17. doi:10.1083/jcb.201403136. ISSN 0021-9525. PMC 4085708. PMID 25002676.
- Huitema, Edgar; Pritchard, Sean; Matteson, David; Radhakrishnan, Sunish Kumar; Viollier, Patrick H. (2006-03-10). "Bacterial Birth Scar Proteins Mark Future Flagellum Assembly Site". Cell. 124 (5): 1025–37. doi:10.1016/j.cell.2006.01.019. ISSN 0092-8674. PMID 16530048. S2CID 15574493.
- Lawler, Melanie L.; Brun, Yves V. (2006-03-10). "A Molecular Beacon Defines Bacterial Cell Asymmetry". Cell. 124 (5): 891–93. doi:10.1016/j.cell.2006.02.027. ISSN 0092-8674. PMID 16530036.