HIV integration
AIDS ("acquired immune deficiency syndrome") is caused by the human immunodeficiency virus (HIV). Individuals with HIV have what is referred to as a "HIV infection". When infected semen, vaginal secretions, or blood come in contact with the mucous membranes or broken skin of an uninfected person, HIV may be transferred to the uninfected person ("horizontal transfer"), causing another infection. Additionally, HIV can also be passed from infected pregnant women to their uninfected baby during pregnancy and/or delivery ("vertical transmission"), or via breastfeeding. As a result of HIV infection, a portion of these individuals will progress and go on to develop clinically significant AIDS.
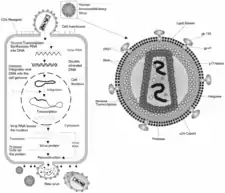
HIV is a retrovirus, which comprise a large and diverse family of RNA viruses that make a DNA copy of their RNA genome after infection of a host cell. An essential step in the replication cycle of HIV-1 and other retroviruses is the integration of this viral DNA into the host DNA. The RNA genome of progeny virions and the template for translation of viral proteins are made when the integrated viral DNA is transcribed.
Background Information: HIV-1 Integrase
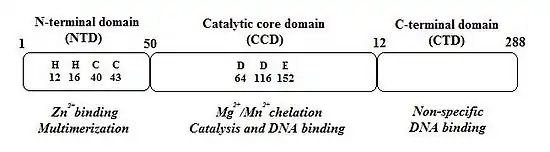
The integration of HIV DNA into the host DNA is a critical step in the HIV life cycle. Understanding the integration process will provide a framework for gaining insight into multiple potential sites of therapeutic intervention for HIV infection and AIDS. HIV’s enzyme for inserting the DNA version of its genome into the host cell DNA is called its "integrase". HIV-1 integrase catalyzes the “cut-and-paste” action of clipping the host DNA and joining the proviral genome to the clipped ends. This protein, which is 288 amino acids in length, contains three “domains”, in this order:
1. Amino (N)-terminal domain: Sometimes referred to as a "zinc finger", the N-terminal domain is composed of the conserved HHCC, His, and Cys residues, a motif that serves to bind zinc. The function of the N-terminal domain is not completely clear, but is thought to assist the integrase in forming multimers (fixed agglomerations of multiple integrase molecules).
2. The central catalytic domain (or "catalytic core"): The catalytic core encompasses the DDE catalytic triad of amino acids, or acid residues, that manage binding with a divalent metal (usually Mg2+ or Mn2−), forming the active catalytic site. In the case of HIV-1 integrase, the residues are Asp64, Asp116, and Glu152. This domain is also well conserved during evolution.
The HIV-1 catalytic domain appears dimeric in solution and in crystal structures. The vast surface area of the dimer interface indicates that it is biologically significant. The insertion sites on each strand of target DNA are separated by 5 base pairs, which parallel to approximately 15 Å for helical B-form DNA, implying that the catalytic domain (or the functional unit) of integrase should contain a pair of active sites separated by a like spacing. This said, the spacing among the active sites in the virtually spherical dimer is, however, apparently not very well-matched with the spacing among the insertion sites on the two strands of target DNA, as examination of crystal structures appears to reveal that the active sites in the dimers are separated by more than 30 Å when measured in a straight line through the proteins, and by an even greater distance when measured around the circumference of the dimer. Under the assumption that the dimer interface is preserved in the functional integrase multimer, at minimum a tetramer of integrase must be required for the complete integration reaction to proceed.
3. The Carboxy (C)- terminal domain : The C-terminal domain non-specifically binds DNA. Since the sites of integration into the target DNA are relatively non-specific, it is thought that this domain may work together in some fashion with the target DNA. Information retrieved from experiments with chimeric integrases show that recognition of the target site is controlled by the core domain. Cross-linking studies also suggest that the C-terminal domain works together with a subterminal region just inside the very ends of the viral DNA.

During the integration process, the HIV integrase enzyme performs two key catalytic reactions. First is the 3’ processing of the HIV DNA, followed by strand transfer of the HIV DNA into the host DNA. The integration of HIV DNA can occur either in dividing or resting cells, and the HIV integrase enzyme can exist in the form of a monomer, dimer, tetramer, and possibly even higher-order forms (such as octomers). Each HIV particle has an estimated 40 to 100 copies of the integrase enzyme.
Integrase functions are unique to retroviruses; human cells are not required to cut-and-paste pieces of DNA into the genome. For this reason, integrase inhibitors are prime targets for developing drug therapies for HIV infection and AIDS, since inhibition of integrase should not hamper the normal operations in human cells.
HIV Integration Mechanism
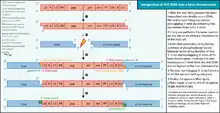
HIV integration is the insertion of HIV genetic material into the genome of the infected cell.[1] The process of HIV integration involves six sequential steps:
Binding of HIV Integrase to HIV DNA
The first step of the integration process occurs in the cytoplasm of the host cell following the completion of reverse transcription of the HIV RNA into c-DNA. This step involves the binding of integrase - most likely in the dimer form - to each end of the newly formed HIV c-DNA. The binding takes place at specific sequences in the long terminal repeat regions. The integrase-HIV DNA complex is part of an intracellular nucleoprotein particle known as the "preintegration complex" (PIC). This complex consists of linear HIV DNA, viral proteins, and host proteins. The viral proteins include integrase, nucleocapsid, matrix, viral protein R (Vpr), and reverse transcriptase. Several host proteins can also form part of this complex, although it is unclear whether some or all join the preintegration complex prior to nuclear transport.
Processing of the HIV DNA 3’ ends
In the second step of the integration process, which also takes place in the host cytoplasm, the integrase dimer cleaves the viral DNA at each 3’ end. This cleavage reaction removes GT dinucleotides on the 3’-side of a conserved CA dinucleotide region. The cleavage of the dinucleotide at each viral DNA 3’-end generates a dinucleotide 5’ "overhang" and a reactive intermediate that contains a 3’-hydroxyl group. This 3’ processing step is the first of two key catalytic reactions performed by the integrase enzyme, and it prepares the viral DNA for integration into the host DNA. In an alternative view of the DNA binding and 3’-processing reaction, the tetramer form of integrase (not the dimer) binds to the ends of the HIV DNA, and then cleaves the 3’ ends.
Translocation of HIV integrase to the host cell nucleus (Nuclear Translocation)
In the third step of the integration process, the preintegration complex is transported into the nucleus of the host cell, entering through one of the nuclear pore complexes.
Binding of the preintegration complex to the host DNA
Inside the nucleus, the host protein lens epithelium-derived growth factor/p75, commonly referred to in abbreviated form as LEDGF/p75, binds to the preintegration complex and the host DNA. The LEDGF/p75 serves as a tethering protein (or bridge) between the preintegration complex and the host DNA. The sequence of binding of the LEDGF/p75, the host DNA, and the preintegration complex remains unclear. In one version, the LEDGF/p75 binds first to the preintegration complex and then to the host DNA. On the other hand, LEDGF/p75 may bind first to the host DNA and then to the preintegration complex. Regardless of the sequence, it is believed that the presence of LEDGF/p75 results in the integrase dimers approaching each other to form a tetramer.
Transfer of HIV DNA into the host DNA (Strand Transfer)
The next step, the strand transfer reaction, takes place inside the host cell nucleus and involves the critical step of inserting the HIV DNA into a selected region of the host DNA. The region of insertion contains a weakly conserved palindromic sequence. This strand transfer reaction is initiated as the HIV integrase catalyzes the HIV DNA 3’-hydroxyl group attack on the host DNA. The attack by the HIV DNA occurs on opposite strands of the host DNA in a staggered fashion, typically 4-6 base pairs apart. This reaction leads to separation of the bonds in the host DNA base pairs located between the staggered cuts, and the joining of the HIV 3’-hydroxyl groups with the host DNA 5’ phosphate ends. At this point, the newly joined viral-host DNA region unfolds.
Repair of the gaps formed in the strand transfer process ("Gap Repair")


Following the strand transfer process, the HIV-DNA and host DNA junctions have unpaired regions of DNA, referred to as DNA "gaps". In addition, the two base pairs at the end of the 5’ region of the viral DNA remain unpaired after the strand transfer. The insertion of the new HIV DNA and the remaining gaps that flank the integration site is currently thought to induce a host cellular DNA damage response, but much of this mechanism remains speculative. The host DNA damage response is thought to be critical in the final step of integration, known as "gap repair" and may require at least three host enzymes - polymerase, nuclease, and ligase. In the first step of gap repair, it is thought that the polymerase enzyme(s) extend(s) the host DNA on each end and, thus, fill in the gaps. Next, it is possible that host nuclease activity removes the 5’ dinucleotide "flaps" on the HIV DNA. Lastly, it is thought that the DNA ligase enzymes join the remaining unbound segment of the HIV and host DNA strands. Currently this mechanism is largely validated experimentally and is an area under investigation. This gap repair process completes the integration of the HIV DNA into the host DNA, with the fully integrated HIV DNA now being referred to as "proviral DNA".
Role of splicing in HIV-1
Recent study suggested that HIV-1 prefers integration into highly spliced genes or genes with more introns.[2] This preference depends on the host chromatin binding protein LEDGF/p75 that interacts with many splicing factors.[2] Another study showed that HIV-1 preference for highly spliced genes depends on another host factor CPSF6, a poly adenylation factor.[3] Cancer genes with high number of introns are highly targeted by HIV-1.[2]
References
- Smith JA, Nunnari G, Preuss M, Pomerantz RJ, Daniel R (2007). "Pentoxifylline suppresses transduction by HIV-1-based vectors". Intervirology. 50 (5): 377–86. doi:10.1159/000109752. PMID 17938572. S2CID 34357038.
- Singh, Parmit Kumar; Plumb R., Matthew; Ferris, Andrea L.; Iben, James R.; Wu, Xiaolin; Fadel, Hind J.; Luke, Brian T.; Esnault, Caroline; Poeschla, Eric M. (2015). "LEDGF/p75 interacts with mRNA splicing factors and targets HIV-1 integration to highly spliced genes". Genes Dev. 29 (21): 2287–97. doi:10.1101/gad.267609.115. PMC 4647561. PMID 26545813.
- Sowd GA, Serrao E, Wang H, Wang W, Fadel HJ, Poeschla EM, et al. (2016). "A critical role for alternative polyadenylation factor CPSF6 in targeting HIV-1 integration to transcriptionally active chromatin". Proc Natl Acad Sci U S A. 113 (8): E1054–63. Bibcode:2016PNAS..113E1054S. doi:10.1073/pnas.1524213113. PMC 4776470. PMID 26858452.
Further reading
- Ciuffi Angela; Bushman Frederic D (2006). "Retroviral DNA integration: HIV and the role of LEDGF/p75". Trends in Genetics. 22 (7): 388–395. doi:10.1016/j.tig.2006.05.006. PMID 16730094.
- Brown; et al. (1987). "P.O. Brown, B. Bowerman, H.E. Varmus, J.M. Bishop, Correct integration of retroviral DNA in vitro". Cell. 49 (3): 347–356. doi:10.1016/0092-8674(87)90287-x. PMID 3032450. S2CID 35523639.
- Bushman; et al. (1990). "Retroviral DNA integration directed by HIV integration protein in vitro". Science. 28 (4976): 1555–1558. Bibcode:1990Sci...249.1555B. doi:10.1126/science.2171144. PMID 2171144.
- Ellison; et al. (1990). "V.H. Ellison, H. Abrams, T. Roe, J. Lifson, P.O. Brown, Human immunodeficiency virus integration in a cell-free system". J. Virol. 64 (6): 2711–2715. doi:10.1128/JVI.64.6.2711-2715.1990. PMC 249450. PMID 2335814.
- Engelman Alan; Mizuuchi Kiyoshi; Craigie Robert (1991). "HIV-1 DNA integration: Mechanism of viral DNA cleavage and DNA strand transfer". Cell. 67 (6): 1211–1221. doi:10.1016/0092-8674(91)90297-C. PMID 1760846. S2CID 1721232.
- Farnet C.M.; Haseltine W.A. (1990). "Integration of human immunodeficiency virus type 1 DNA in vitro". Proc. Natl. Acad. Sci. USA. 87 (11): 4164–4168. Bibcode:1990PNAS...87.4164F. doi:10.1073/pnas.87.11.4164. PMC 54068. PMID 2349226.
- Lander E.S.; Linton L.M.; Birren B.; Nusbaum C.; Zody M.C.; Baldwin J.; Devon K.; Dewar K.; Doyle M.; Fitzhugh W.; et al. (2001). "Initial sequencing and analysis of the human genome" (PDF). Nature. 409 (6822): 860–921. Bibcode:2001Natur.409..860L. doi:10.1038/35057062. PMID 11237011.
External links
- Stowell, Dan (2006). "preintegration complex". The Molecules of HIV - A Hypertextbook. Retrieved 2016-01-11.