Section 19
RNA-Based Regulation
Book
Version 6
By Boundless
By Boundless
Boundless Microbiology
Microbiology
by Boundless
7 concepts
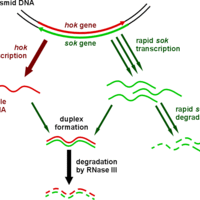
RNA Regulation and Antisense RNA
Antisense RNAs are single-stranded RNA molecules that can bind and inhibit specific mRNA translation to protein.

Attenuation
Attenuation is a mechanism utilized by bacteria to regulate unnecessary gene expression.
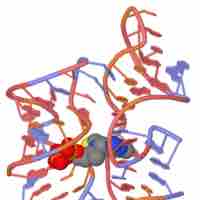
Riboswitches
Riboswitches are naturally occurring RNA molecules that can regulate gene expression.
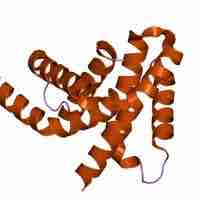
Regulation of Sigma Factor Activity
The sigma factor is responsible for proper transcriptional initiation.
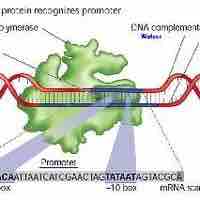
Regulation of Sigma Factor Translation
Sigma factors are proteins that regulate gene expression that are controlled at various levels, including at the translational level.
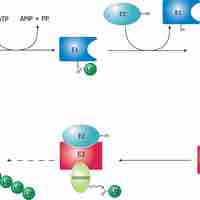
Proteolytic Degradation
Proteolytic degradation, or proteolysis, is a key factor that controls protein concentration and function.
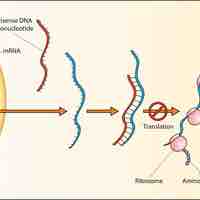
Small Regulatory RNAs
Small regulatory RNAs are non-coding RNA molecules that play a role in cellular processes such as activation or inhibition processes.