Platensimycin
 | |
 | |
| Names | |
|---|---|
| IUPAC name
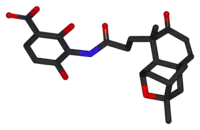
3-[[3-[(1R,3R,4R,5aR,9R,9aS)-1,4,5,8,9,9a-Hexahydro-3,9-dimethyl-8-oxo-3H-1,4:3,5a-dimethano-2-benzoxepin-9-yl]-1-oxopropyl]amino]-2,4-dihydroxy-benzoic acid | |
| Identifiers | |
CAS Number |
|
3D model (JSmol) |
|
| ChEBI | |
| ChEMBL | |
| ChemSpider | |
| DrugBank | |
PubChem CID |
|
| UNII | |
CompTox Dashboard (EPA) |
|
InChI
| |
SMILES
| |
| Properties | |
Chemical formula |
C24H27NO7 |
| Molar mass | 441.480 g·mol−1 |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
| Infobox references | |
Platensimycin, a metabolite of Streptomyces platensis, is an antibiotic, which act by blocking enzymes (β-ketoacyl-(acyl-carrier-protein (ACP)) synthase I/II (FabF/B)).[1]
History
Platensimycin was first isolated from a strain of Streptomyces platensis by workers at Merck.[2][3] Screens of 250,000 natural product extracts (83,000 strains in three growth conditions) led to the identification of a potent and selective small molecule from a strain of Streptomyces platensis recovered from a soil sample collected in South Africa. The identification process was carried out using a two-plate system in which control organisms were compared to cells expressing FabF antisense RNA. This method uses a combination of target-based whole-cell and biochemical assays, allowing compounds to be detected at concentrations that would be too low to detect using whole cell assays. The molecule they identified, platensimycin (C24H27NO7, relative molecular mass 441.47), comprises two distinct structural elements connected by an amide bond. The Merck group showed that platensimycin has potent, broad-spectrum Gram-positive activity in vitro and exhibits no cross-resistance to other key antibiotic-resistant bacteria including Methicillin-resistant Staphylococcus aureus(MRSA), vancomycin-intermediate S. aureus, vancomycin-resistant Enterococci, and linezolid-resistant and macrolide-resistant pathogens.
As confirmed by total synthesis of racemic platensimycin, its structure consists of a 3-amino-2,4-dihydroxybenzoic acid polar part linked through an amide bond to a lipophilic tetracyclic ketolide.[4]
Clinical use
Platensimycin is an experimental drug in preclinical trials involving MRSA in a mouse model. Platensimycin is an effective antibiotic in vivo when continuously administered to cells. Efficacy is reduced when administered by more conventional means.[5] Clinical trials have been delayed.[6] A variety of modifications have been investigated.[7][8] and increase the activity of platensimycin.
Biosynthesis
Biosynthesitic studies show that the benzoic ring is produced from pyruvate and acetate via the TCA cycle, while the C-17 tetracyclic enone acid core is produced from the non-mevalonate terpenoid pathway.
The tetracyclic enone isotope labeling pattern observed is consistent with the biosynthesis of the tetracycle via the non-mevalonate terpenoid pathway.[9][10] This pathway involves condensation of a thiamine-activated acetyl group arising from the decarboxylation of pyruvate and glyceraldehyde-3-phosphate followed by a transposition step. Since both pyruvate and glyceraldehyde-3-phosphate (also glycerol) are part of the glycolytic pathway, varying levels of incorporation are expected. Thus, the terpenoid building blocks, dimethylallyl diphosphate and isopentenyl diphosphate, synthesized by the non-mevalonate pathway utilizing pyruvate and glyceraldehyde-3-phosphate, condense to form the diterpenoid precursor geranylgeranyl diphosphate that cyclizes to an intermediate which is related to (or derived from) ent-kaurene.[11] Oxidative cleavage of the double bond of this intermediate would result in the loss of the terminal three carbons producing the C-17 tetracyclic enone acid unit. An N-acyltransferase reaction of tetracyclic enone and aminobenzoic acid would lead to platensimycin.
Mechanism of action
Platensimycin has shown good activity against a panel of Gram-positive bacteria, including various resistant strains.
Platensimycin works by inhibiting beta-ketoacyl synthases I/II (FabF/B), which are involved in the production of fatty acids required for bacterial cell membranes. It interferes with enzymes involved in the condensation steps in fatty acid biosynthesis,[12] which Gram-positive bacteria need to biosynthesise cell membranes. Other enzymes in this pathway have similarly been proven as antibiotic targets, such as FabI, the enoyl-ACP (acyl carrier protein) reductase, which is inhibited by isoniazid and related compounds and the antiseptic agent triclosan.[13]
One proposed mechanism of action is that, firstly, the thiol group of FabF Cys163 is activated through the dipole moment of helix N-alpha-3 which lowers the pKa.[14] The nucleophilicity of the cysteine is enhanced by an oxyanion hole formed with the backbone amides of Cys163 and Phe400.
The crystal structure complex with platensimycin employed a C163Q mutant, which gave a 50-fold increase in apparent binding. The Gln163 residue lies adjacent to the carboxylate of platensimycin but makes no specific hydrogen bond. The close proximity of the carboxylate of platensimycin (presumed to be an anion) to the anionic thiol of Cys163 in the wild type enzyme may suggest the reason behind the increase in binding of the C163Q mutant. The second set of residues worth considering comprises His303 and His340, which play a role in the decarboxylation mechanism of the malonyl moiety. In particular, His303 activates a structured water to attack the carboxylate of the incoming malonyl-ACP.[15] The crystal structure of FabF also demonstrates that His340 forms a hydrogen bond between the amide nitrogen of Leu342 and the N-delta- atom of the imidazole ring meaning that the lone pair must reside on this atom. In the platensimycin crystal structure the structured water adjacent to His303 is no longer present which may suggest an alternative electronic state for this residue. A strong possibility exists that His303 would present itself as a cation capable of forming an ionic interaction with the benzoic acid group of platensimycin.
References
- ↑ Rudolf, Jeffrey D.; Dong, Liao-Bin; Shen, Ben (2017). "Platensimycin and platencin: Inspirations for chemistry, biology, enzymology, and medicine". Biochemical Pharmacology. 133: 139–151. doi:10.1016/j.bcp.2016.11.013. PMC 5410400. PMID 27865713.
{{cite journal}}: CS1 maint: uses authors parameter (link) - ↑ Wang, Jun; Soisson, Stephen M.; Singh, Sheo; et al. (2006). "Platensimycin is a selective FabF inhibitor with potent antibiotic properties". Nature. 441 (7091): 358–61. Bibcode:2006Natur.441..358W. doi:10.1038/nature04784. PMID 16710421. S2CID 4329677. Retrieved 17 May 2020.
- ↑ Manallack, David T.; Crosby, I. T.; Khakham, Y; Capuano, B (2008). "Platensimycin: A Promising Antimicrobial Targeting Fatty Acid Synthesis". Current Medicinal Chemistry. 15 (7): 705–10. doi:10.2174/092986708783885255. PMID 18336284. Retrieved 17 May 2020.
- ↑ Nicolaou, K.C.; Li, Ang; Edmonds, David J. (2006). "Total Synthesis of Platensimycin". Angewandte Chemie International Edition. 45 (42): 7086–90. doi:10.1002/anie.200603892. PMID 17013803.
- ↑ Herath, Kithsiri B.; Attygalle, Athula B.; Singh, Sheo B. (2007). "Biosynthetic Studies of Platensimycin". Journal of the American Chemical Society. 129 (50): 15422–3. doi:10.1021/ja0758943. PMID 18034483.
- ↑ Smanski, Michael J.; Peterson, Ryan M.; Rajski, Scott R.; Shen, Ben (2009). "Engineered Streptomyces platensis Strains That Overproduce Antibiotics Platensimycin and Platencin". Antimicrobial Agents and Chemotherapy. 53 (4): 1299–1304. doi:10.1128/aac.01358-08. PMC 2663125. PMID 19164156.
- ↑ Krauss, Jurgen; Knorr, Veronika; Manhardt, Vera; Scheffels, Stephanie; Bracher, Franz (2008). "Synthesis of Platensimycin Analogues and Their Antibiotic Potency". Archiv der Pharmazie. 341 (6): 386–92. doi:10.1002/ardp.200700177. PMID 18442030. S2CID 3002538.
- ↑ Lu, X.; You, Q. (2010). "Recent advances on platensimycin: a potential antimicrobial agent". Current Medicinal Chemistry. 17 (12): 1139–55. doi:10.2174/092986710790827852. PMID 20158476.
- ↑ S W. White, J. Zheng, Y X M. Zhang, and C O. Rock, Annu. Rev. Biochem. 2005, 74, 791-831.
- ↑ S. Smith, A. Witkowski, and A K. Joshi, Prog. Lipid Res. 2003, 42, 289-317.
- ↑ W P. Revill, M J. Bibb, A K. Scheu, H J. Kieser, and D A. Hopwood, J. Bacteriol., 2001, 183, 3526-30.
- ↑ Häbich, Dieter; von Nussbaum, Franz (2006). "Platensimycin, a New Antibiotic and "Superbug Challenger" from Nature". ChemMedChem. 1 (9): 951–4. doi:10.1002/cmdc.200600145. PMID 16952137.
- ↑ Wright, H. Tonie; Reynolds, Kevin A. (2007). "Antibacterial targets in fatty acid biosynthesis". Current Opinion in Microbiology. 10 (5): 447–53. doi:10.1016/j.mib.2007.07.001. PMC 2271077. PMID 17707686.
- ↑ A C. Price, C O. Rock, S W. White, J. Bacteriol., 2003, 185, 4136-43.
- ↑ Y M. Zhang, J. Hurlbert, S W. White, C O. Rock, J. Biol. Chem. 2006, 281, 17390-99.