6-Phosphogluconate dehydrogenase
6-Phosphogluconate dehydrogenase (6PGD) is an enzyme in the pentose phosphate pathway. It forms ribulose 5-phosphate from 6-phosphogluconate:
- 6-phospho-D-gluconate + NAD(P)+ D-Ribulose 5-phosphate + CO2 + NAD(P)H + H+
| 6PGD | |||||||||
|---|---|---|---|---|---|---|---|---|---|
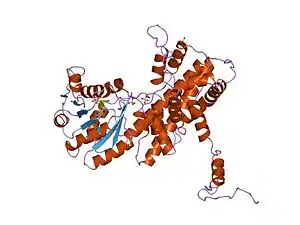 Crystallographic structure of sheep 6-phosphogluconate dehydrogenase complexed with adenosine 2'-monophosphate[1] | |||||||||
| Identifiers | |||||||||
| Symbol | 6PGD | ||||||||
| Pfam | PF00393 | ||||||||
| Pfam clan | CL0106 | ||||||||
| InterPro | IPR006114 | ||||||||
| PROSITE | PDOC00390 | ||||||||
| SCOP2 | 2pgd / SCOPe / SUPFAM | ||||||||
| |||||||||
| Phosphogluconate dehydrogenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
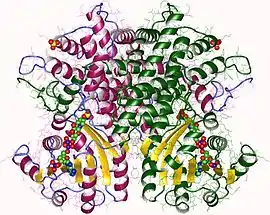 6-phosphogluconate dehydrogenase dimer, Ovis aries | |||||||||
| Identifiers | |||||||||
| EC no. | 1.1.1.44 | ||||||||
| CAS no. | 9001-82-5 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| phosphogluconate dehydrogenase | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | PGD | ||||||
| NCBI gene | 5226 | ||||||
| HGNC | 8891 | ||||||
| OMIM | 172200 | ||||||
| RefSeq | NM_002631 | ||||||
| UniProt | P52209 | ||||||
| Other data | |||||||
| EC number | 1.1.1.44 | ||||||
| Locus | Chr. 1 p36.3-36.13 | ||||||
| |||||||
It is an oxidative carboxylase that catalyses the decarboxylating reduction of 6-phosphogluconate into ribulose 5-phosphate in the presence of NADP. This reaction is a component of the hexose mono-phosphate shunt and pentose phosphate pathways (PPP).[2][3] Prokaryotic and eukaryotic 6PGD are proteins of about 470 amino acids whose sequences are highly conserved.[4] The protein is a homodimer in which the monomers act independently:[3] each contains a large, mainly alpha-helical domain and a smaller beta-alpha-beta domain, containing a mixed parallel and anti-parallel 6-stranded beta sheet.[3] NADP is bound in a cleft in the small domain, the substrate binding in an adjacent pocket.[3]
Biotechnological significance
Recently, 6PGD was demonstrated to catalyze also the reverse reaction (i.e. reductive carboxylation) in vivo.[5] Experiments using Escherichia coli selection strains revealed that this reaction was efficient enough to support the formation of biomass based solely on CO2 and pentose sugars. In the future, this property could be exploited for synthetic carbon fixation routes.
Clinical significance
Mutations within the gene coding this enzyme result in 6-phosphogluconate dehydrogenase deficiency, an autosomal hereditary disease affecting the red blood cells.
As a possible drug target
6PGD is involved in cancer cell metabolism so 6PGD inhibitors have been sought.[6]
See also
- Parietin, a 6PGD inhibitor
References
- PDB: 1PGQ; Adams MJ, Ellis GH, Gover S, Naylor CE, Phillips C (July 1994). "Crystallographic study of coenzyme, coenzyme analogue and substrate binding in 6-phosphogluconate dehydrogenase: implications for NADP specificity and the enzyme mechanism". Structure. 2 (7): 651–68. doi:10.1016/s0969-2126(00)00066-6. PMID 7922042.
- Broedel SE, Wolf RE (July 1990). "Genetic tagging, cloning, and DNA sequence of the Synechococcus sp. strain PCC 7942 gene (gnd) encoding 6-phosphogluconate dehydrogenase". J. Bacteriol. 172 (7): 4023–31. doi:10.1128/jb.172.7.4023-4031.1990. PMC 213388. PMID 2113917.
- Adams MJ, Archibald IG, Bugg CE, Carne A, Gover S, Helliwell JR, Pickersgill RW, White SW (1983). "The three dimensional structure of sheep liver 6-phosphogluconate dehydrogenase at 2.6 A resolution". EMBO J. 2 (6): 1009–14. doi:10.1002/j.1460-2075.1983.tb01535.x. PMC 555222. PMID 6641716.
- Reizer A, Deutscher J, Saier MH, Reizer J (May 1991). "Analysis of the gluconate (gnt) operon of Bacillus subtilis". Mol. Microbiol. 5 (5): 1081–9. doi:10.1111/j.1365-2958.1991.tb01880.x. PMID 1659648. S2CID 2006623.
- Satanowski A, Dronsella B, Noor E, Vögeli B, He H, Wichmann P, et al. (November 2020). "Awakening a latent carbon fixation cycle in Escherichia coli". Nature Communications. 11 (1): 5812. doi:10.1038/s41467-020-19564-5. PMC 7669889. PMID 33199707.
- 6-Phosphogluconate dehydrogenase links oxidative PPP, lipogenesis and tumour growth by inhibiting LKB1–AMPK signalling. 2015
Further reading
- Frampton EW, Wood WA (October 1961). "Carbohydrate oxidation by Pseudomonas fluorescens VI. Conversion of 2-keto-6-phosphogluconate to pyruvate". The Journal of Biological Chemistry. 236: 2571–7. doi:10.1016/S0021-9258(19)61700-X. PMID 13894458.