Brine pool
A brine pool, sometimes called an underwater lake, deepwater or brine lake, is a volume of brine collected in a seafloor depression. The pools are dense bodies of water that have a salinity that is three to eight times greater than the surrounding ocean. Brine pools are commonly found below polar sea ice and in the deep ocean. Those below sea ice form through a process called brine rejection.[1] For deep-sea brine pools, salt is necessary to increase the salinity gradient. The salt can come from one of two processes: the dissolution of large salt deposits through salt tectonics[2] or geothermally heated brine issued from tectonic spreading centers.[3]



The brine often contains high concentrations of hydrogen sulfide and methane, which provide energy to chemosynthetic organisms that live near the pool. These creatures are often extremophiles and symbionts.[4][5] Deep-sea and polar brine pools are toxic to marine animals due to their high salinity and anoxic properties,[1] which can ultimately lead to toxic shock and possibly death.
Characteristics
Brine pools are sometimes called sea floor "lakes" because the dense brine does not easily mix with overlying seawater creating a distinct interface between water masses. The pools range in area from less than 1 square metre (11 sq ft) to as large as the 120-square-kilometre (46 sq mi) Orca Basin.[2] The high salinity raises the density of the brine, which creates a surface and shoreline for the pool. Depending on concentration, some minerals such as baryte (barium sulfate) precipitate out of the brine and form crystalline crusts around the edge of the pool.[6]
Because of the brine's high density and lack of mixing currents in the deep ocean, brine pools often become anoxic and deadly to respiring organisms.[7] Brine pools supporting chemosynthetic activity, however, form life on the pool's shores where bacteria and their symbionts grow near the highest concentrations of nutrient release.[8] Patchy, reddish layers can be observed floating above the dense brine interface due to high densities of halophilic archaea that are supported by these environments.[9] These shores are complex environments with significant shifts in salinity, oxygen concentration, pH, and temperature over a relatively small vertical scale. These transitions provide a variety of environmental niches.[10][11]
Formation
Brine pools are created through three primary methods: brine rejection below sea ice, dissolution of salts into bottom water through salt tectonics, and geothermal heating of brine at tectonic boundaries and hot spots.
- Brine rejection: When sea water freezes, salts do not fit into the crystalline structure of ice so the salts are expelled. The expelled salts form a cold, dense, brine that sinks below the sea ice to the sea floor. Brine rejection on an oceanic scale is associated with the formation of North Atlantic Deep Water (NADW) and Antarctic Bottom Water (AAW) that play a large role in global thermohaline circulation (THC). On a localized scale, that rejected brine collects in seafloor depressions forming a brine pool. In the absence of mixing, the brine will become anoxic in a matter of weeks.[1]
- Salt tectonics: During the Middle Jurassic period, the Gulf of Mexico was a shallow sea that dried out, producing a thick layer of salt and seawater derived minerals up to 8 km thick. When the Gulf refilled with water, the salt layer was preserved from dissolution by sediments accumulating over the salt. Subsequent sedimentation layers became so heavy that they began to deform and move the more malleable salt layer below. In some places, the salt layer now protrudes at or near the seafloor where it can interact with seawater. Where seawater comes in contact with the salt, the deposits dissolve and form brines. The location of these surfacing Jurassic period salt deposits is also associated with methane releases giving deep ocean brine pools their chemical characteristics.[2]
- Geothermal heating: At earth's oceanic tectonic spreading centers, plates are moving apart, allowing new magma to rise and cool. This process is involved in creating new sea floor. These mid-ocean ridges allow seawater to seep downward into fractures where they come in contact with and dissolve minerals. In the Red Sea for example, Red Sea Deep Water (RSDW) seeps into the fissures created at the tectonic boundary. The water dissolves salts from deposits created in the Miocene epoch much like the Jurassic period deposits in the Gulf of Mexico. The resulting brine is then superheated in the hydrothermal active zone over the magma chamber. The heated brine rises to the seafloor where it cools and settles in depressions as brine pools. The location of these pools is also associated with methane, hydrogen sulfide, and other chemical releases setting the stage for chemosynthetic activity.[3]
Support of life
Due to the methods of their formation and lack of mixing, brine pools are anoxic and deadly to aerobic organisms, including most eukaryotes and multicellular organisms. When an organism enters a brine pool, they attempt to "breathe" the environment and experience cerebral hypoxia due to the lack of oxygen and toxic shock from the hyper-salinity. If organisms cannot surface long enough to retreat to the rim, they quickly die.[12] When observed by submarines or remotely operated underwater vehicle (ROV), brine pools are found to be eerily littered with dead fish, crabs, amphipods, and various organisms that ventured too far into the brine. Dead organisms are then preserved in the brine for years without decay due to the anoxic nature of the pool preventing decay and creating a fish "graveyard".[8]
Despite the harsh conditions, life in the form of macrofauna such as bivalves can be found in a thin area along the rim of a brine pool. A novel genus and species of bivalves, known as Apachecorbula muriatica, have been found along the edge of the "Valdiva Deep" brine pool in the Red Sea.[13] There have also been recorded instances of macrofauna brine pools at the seawater interface. Inactive sulfur chimneys have been found with affiliated epifauna such as polychaetes and hydroids. In fauna like gastropods, capitellid polychaetes, and top snails have also been found to be associated with brine pools in the Red Sea. Such species typically feed on microbial symbionts or bacterial and detritus films.[14]
While organisms can typically flourish on the outskirts of a brine pool, they are not always safe from harm here. One possible reason for this is that underwater landslides can impact brine pools and cause waves of hypersaline brine to spill out into surrounding basins, thus negatively affecting the biological communities which live there.[15]
Despite their inhospitable nature, brine pools can also provide a home, allowing organisms to flourish. Deep-sea brine pools often coincide with cold seep activity allowing for chemosynthetic life to thrive. Methane and hydrogen sulfide released by the seep is processed by bacteria, which have a symbiotic relationship with organisms such as seep mussels.[16] The seep mussels create two distinct zones. The inner zone, which is at the edge of the pool, provides the best physiological conditions and allows for maximum growth. The outer zone is near the transition between the mussel bed and the surrounding seafloor, and this area provides the worst conditions causing these mussels to have lower maximum sizes and densities.[17] This ecosystem is dependent on chemical energy and, relative to almost all other life on Earth, has no dependence on energy from the Sun.[18]
An important part of the study of extreme environments such as brine pools is the function and survival of microbes. Microbes help support the larger biological community around environments like brine pools and are key to understanding the survival of other extremophiles. Biofilms contribute to the creation of microbes and are considered the foundation by which other micro-organisms can survive in extreme environments. The research into the growth and function of artificial extremophile biofilms has been slow due to the difficulty in recreating the extreme deep-sea environments they are found in.[19]
Microbial diversity and community composition
Microbial composition
Red Sea brine pool microbiology is among the most intensively studied using metagenomics and amplicon sequencing.
Metagenomic analysis
Metagenomic analysis is a powerful approach for characterizing microbial communities in a variety of environments. Previously, genetic analysis required having the microorganisms in culture, which is problematic since most microorganisms in nature have not been cultivated.[20] Metagenomics overcomes these problems by allowing researchers to directly sample and analyze and genetically characterize microbial communities sampled from the desired environment.[21] Metagenomic analyses has revealed previously uncharacterized microbial communities in multiple brine pools.[22] Common procedures for characterizing marine microbial communities by metagenomic analysis includes sampling, filtration and extraction, DNA sequencing, and comparison to databases.
Main clades
The taxonomic makeup of the main microbial communities found at Atlantis II and Discovery without including minor or unknown species to avoid ambiguity is summarized in the following list that is based on the data from primary articles.[23][24][25]
| Domain | Microbes |
|---|---|
| Bacteria | [Order] Actinomycetales[25] |
| [Family] Microbacteriaceae[23] | |
| [Genus] Bacillus[24] | |
| [Phylum] Bacteroidetes[25] | |
| [Class] Flavobacteria[23] | |
| [Phylum] Candidatus division od1[23] | |
| [Phylum] Chloroflexi[25] | |
| [Class] Anaerolineae[23] | |
| [Class] SAR202 clade[23] | |
| [Phylum] Cyanobacteria[23] | |
| [Phylum] Deinococcota[25] | |
| [Genus] Meiothermus[24] | |
| [Class] Deferribacteres[23] | |
| [Phylum] Firmicutes[25] | |
| [Order] Thermotoaea[25] | |
| [Class] Candidatus Scalindua[23] | |
| [Class] Candidatus Brocadiales l[23] | |
| [Class] Alphaproteobacteria[23][25] | |
| [Genus] Phyllobacterium[24] | |
| [Genus] Afipia[24] | |
| [Genus] Bradyrhizobium[24] | |
| [Order] SAR11(Pelagibacterales)[23] | |
| [Class] Betaproteobacteria[23][25] | |
| [Genus] Rhodoferax[24] | |
| [Genus] Malikia[24] | |
| [Genus] Cupriavidus[24] | |
| [Genus] Ralstonia[24] | |
| [Class] Deltaproteobacteria[23][25] | |
| [Class] Gammaproteobacteria[25] | |
| [Order] Alteromonadales[23] | |
| [Order] Oceanospirillales[23] | |
| [Genus] Acinetobacter[24] | |
| [Genus] Alkanindiges[24] | |
| [Genus] Stenotrophomonas[24] | |
| Archaea | [Class] Group c3[23] |
| [Class] Marine benthic group a (MBG-A)[23] | |
| [Class] Marine benthic group b (MBG-B)[23] | |
| [Class] Marine group I (MGI)[23] | |
| [Class] Misc crenarchaeotic group[23] | |
| [Class] Psl12[23] | |
| [Order] Desulfurococcales[25] | |
| [Order] Sulfolobales[25] | |
| [Order] Thermoproteales[25] | |
| [Phylum] Euryarchaeota[25] | |
| [Class] Archaeoglobi[23] | |
| [Order] Archaeoglobales[25] | |
| [Order] Halobacteria[23] | |
| [Order] Halobacteriales[25] | |
| [Class] Methanomicrobia[23] | |
| [Order] Methanobacteriales[25] | |
| [Order] Methanocellales[25] | |
| [Order] Methanococcales[25] | |
| [Order] Methanomicrobiales[25] | |
| [Order] Methanopyrales[25] | |
| [Order] Methanosarcinales[25] | |
| [Class] Thermoplasmata[23] | |
| [Genus] Candidatus Korarchaeum cryptofilum[25] | |
| [Genus] Candidatus Caldiarchaeum[25] |
Environmental challenges and adaptations
The lack of mixing with the water column in combination with high salinity, anoxia, extremes in water temperature and hydrostatic pressure results in microbial assemblages that are specific to these environments.[26]
Challenges
The high salinity levels present challenges for the retention of water by cells and consequent effects on cell turgor and functioning.[26] Brine pools also exert ionic, kosmotropic and chaotropic effects on the cells, which also causes additional challenges for the organisms to survive these extreme environments.[27][28]
In addition, the lack of oxygen increases the difficulty of organisms to yield energy, as oxygen is the most energy-yielding electron acceptor.[29]
Adaptations
Organisms developed different strategies to solve the challenges imposed by high levels of salinity. In order to decrease the risk of chaotropic effects on the cells, halophilic archaea have a "salt-in" approach and "compatible-solute" strategy, which increases intracellular ionic concentration (mostly K+) to decrease the osmotic pressure; Thus, these organisms have adapted their entire metabolic machinery to maintain salt concentration inside of their cells.[30]
In some brine pools, high water temperatures and hydrostatic pressures, result in piezophilic microorganisms that synthesize thermoprotective molecules (e.g. hydroxyketone) to prevent the denaturation of proteins and decrease the risk of desiccation.[31][32][33][34]
Another important adaptation is the use of alternative electron acceptors to yield energy, such as iron, manganese,[35] sulfate, elemental sulfur,[36] carbon dioxide, nitrite, and nitrate.[37]
Animals have also been found living in these anaerobic brine pools, such as the first known metazoan from these environments described by Danovaro et al. (2010).[38] Many other taxa that from these extreme environments are still uncharacterized.[39][40]
Nutrient cycling in brine pools
Chemical composition and metabolic significance
As the name suggests, brine pools, or deep hypersaline anoxic basins (DHABs), are characterized by a very high salt concentration and anoxic conditions. Sodium, chloride, magnesium, potassium, and calcium ion concentration are all extremely high in brine pools. And, due to low mixing rates between the above seawater and the brine water, brine-pool water becomes anoxic within the first ten centimeters or so.[41] While there are large variations in the geochemical composition of individual pools,[41] as well as extreme chemical stratification within the same pool,[42] conserved chemical trends are present. Deeper layers of DHABs will be saltier, hotter, more acidic and more anaerobic than the preceding layers.[43][44] The concentration of heavy metals (Fe, Mn, Si, Cu) and certain nutrients (NO2−, NH4+, NO3−, and PO4−) will tend to increase with depth, while the concentration of SO4− and both organic and inorganic carbon decrease with depth.[42] While these trends are all observed to some capacity in DHABs, the intensity and distance over which these trends take effect can vary in depth from one meter, to tens of meters.[41]
The heavy stratification within DHABs has led to increased microbial metabolic diversity and varying cell concentrations between layers. The majority of cell biomass has been found at the interfaces between the distinct chemical layers (with the highest concentrations of cells located at the brine-surface interface).[45] Microbes exploit the sharp chemical gradients between the layers to make their metabolisms more thermodynamically favorable.[46]
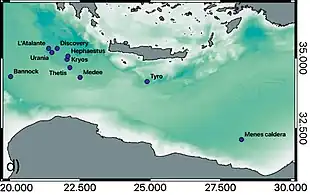
Four heavily studied DHABs are Urania, Bannock, L'Atalante, and Discovery. All four of these brine pools are located in the Mediterranean Sea, yet they all exhibit distinct chemical properties: Urania has the highest concentration of sulfuric acid observed (at c. 16 mM)—compared to normal sea water (2.6×10−6 mM or the next highest [HS-] in the Bannock basin (c. 3 mM).[47][45] Discovery has an extremely low concentration of Na+ (68 mM) and an extremely high concentration of Mg2+ (4,995 mM)—compared to the surrounding seawater with concentrations of 528 mM and 60 mM respectively.[45][48] The L'Atalante basin has a high SO42- concentration compared to the other pools. This extreme variability in environmental conditions leads to each brine pool having a unique metabolic composition.
Carbon cycling
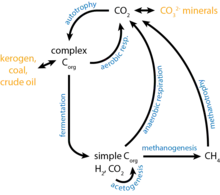
While it was initially thought that particulate organic matter (POM) was an important source of carbon for DHABs, due to their depth, the concentration of POM reaching the pools is not significant as originally thought.[41] The majority of fixed carbon is now thought to come from autotrophy, specifically methanogenesis. Direct measurements of methane production in DHABs have provided extensive molecular evidence of methanogenesis in these environments.[45] Proteomic analysis further support the presence of methanogenesis by identifying the enzyme RuBisCo in various DHABs.[49] Interestingly, it has been suggested that instead of CO2 or acetoclastic methanogenesis, prokaryotes in DHABs use methylotrophic methanogenesis as it allows for a higher energy yield[50] and the intermediates can be used for osmoprotectants.[51]
Nitrogen cycling

One of the key metabolic features of DHABs is the dissimilatory reduction of nitrogen. This is predominantly due to the thermodynamic favorability of nitrogen based metabolisms in anaerobic environments. In Bannock basin and L'Atalante basin, anammox and denitrification pathways have been identified using a combination of transcriptomics and direct isotope tracking.[52] Other DHABs have been analyzed for anammox pathways using metatranscriptomic techniques with little positive results, which may be due to the limitations of transcriptomic sensitivity. In deeper DHAB layers, nitrogen fixation and ammonium assimilation has been observed. These reductive pathways require a lot of energy and are mainly performed by methanogens to synthesize osmoprotectants.[53]
Sulfur cycling
Due to the high concentration of sulfate (especially in the Uranian Basin), sulfate reduction is extremely important in the biogeochemical cycling of DHABs. The highest rates of sulfate reduction tend to be found in the deepest DHAB layers, where redox potential is lowest.[46] Sulfate reducing bacteria have been found in the brines of Kebrit deep, Nereus Deep, Erba deep, Atlantis II deep, and Discovery Deep.[54] Oxidative sulfur pathways help close the biogeochemical sulfur loops within the DHABs. There are three main sulfur oxidizing pathways which are likely found in DHABs:
- sulfur-oxidizing multienzyme complex which can oxidize sulfide or thiosulfate to sulfate (w/ elemental sulfur or sulfite as an intermediate).
- a sulfide/quinone complex which oxidizes hydrogen sulfide to elemental sulfur.
- polysulfide reductase, which reduces precipitate sulfur to sulfide.
A combination between the second and third pathway would allow for increased energetic yield.[55] In addition, some novel groups have been isolated from saline lakes which can anaerobically respire sulfur using acetate, pyruvate, formate, or hydrogen as a sole electron donors.[56]
Microbial symbiosis
There is a high concentration of bacteria present in brine pools that serve essential roles for the ecosystem, such as being part of symbiotic relationships or acting as a food source for several organisms in this habitat. Examples include tubeworms and clams having a symbiotic relationship with many of these bacteria to convert chemical energy from hydrogen sulfide, and in exchange providing them food to allow reproduction and development;[57] or mussels providing a safe habitat for bacteria that feed on methane while thriving due to the chemosynthetic, carbon-fixing symbionts that are inhabiting their gill tissues.[58] Thus, these symbiotic relationships with bacteria allow organisms to be abundant and have high biomass in these harsher environments.[59]
Bacteria can also act as epibiotic symbiont, which were found to play an important role in the adaptations of microorganisms to these environments, such as organisms from the flagellated group Euglenozoa that have been thriving in brine pools due to this relationship.[60]
Examples
- Afifi[61]
- Atlantis II[62]
- Conrad[63]
- Discovery[64]
- Kebrit[65]
- Kryos [66]
- L'Atalante basin
- Orca Basin
- Shaban[67]
Future uses
One major idea involves harnessing the salinity of brine pools to use as a power source. This would be done using an osmotic engine which draws the high salinity top water through the engine and pushes it down due to osmotic pressure. This would cause the brackish stream (which is less dense and has a lighter salinity) to be propelled away from the engine via buoyancy. The energy created by this exchange can be harnessed using a turbine to create a power output.[7]
It is possible to study liquid brine in order to harness its electrical conductivity to study if liquid water is present on Mars.[68] A HABIT (Habitability: Brines, Irradiation, and Temperature) instrument will be part of a 2020 campaign to monitor changing conditions on Mars. This device will include a BOTTLE (Brine Observation Transition to Liquid Experiment) experiment to quantify the formation of transient liquid brine as well as observe its stability over time under non-equilibrium conditions.[68]
A third idea involves using microorganisms in deep-sea brine pools to form natural product drugs.[69] These microorganisms are important sources of bioactive molecules against various diseases due to the extreme environment they inhabit, giving potential to an increasing number of drugs in clinical trials.[70] In particular, a novel finding in a study used microorganisms from the Red Sea brine pools as potential anticancer drugs.[71][72][73]
Deep sea brine pools have also been a large interest in bioprospecting in the hope that unlikely environments might serve as sources of biomedical breakthroughs due to unexplored biodiversity. Some areas have been found to host antibacterial and anticancer activities in biosynthetic clusters.[74] Other novel antibiotic resistance enzymes have been found that are useful in various biomedical and industrial applications.[75]
References
- Kvitek, Rikk (February 1998). "Black pools of death: Hypoxic, brine-filled ice gouge depressions become lethal traps for benthic organisms in a shallow Arctic embayment". Marine Ecology Progress Series. 162: 1–10. Bibcode:1998MEPS..162....1K. doi:10.3354/meps162001 – via ResearchGate.
- "NOAA Ocean Explorer: Gulf of Mexico 2002". oceanexplorer.noaa.gov. Retrieved 2020-09-28.
- Salem, Mohamed (2017-06-01). "Study of Conrad and Shaban deep brines, Red Sea, using bathymetric, parasound and seismic surveys". NRIAG Journal of Astronomy and Geophysics. 6 (1): 90–96. Bibcode:2017JAsGe...6...90S. doi:10.1016/j.nrjag.2017.04.003. S2CID 132353952.
- Extremophile life near brine pools Archived November 10, 2006, at the Wayback Machine
- Eder, W; Jahnke, LL; Schmidt, M; Huber, R (July 2001). "Microbial diversity of the brine-seawater interface of the Kebrit Deep, Red Sea, studied via 16S rRNA gene sequences and cultivation methods". Appl. Environ. Microbiol. 67 (7): 3077–85. Bibcode:2001ApEnM..67.3077E. doi:10.1128/AEM.67.7.3077-3085.2001. PMC 92984. PMID 11425725.
- Stevens, Eric W. (October 2013). Barite crusts from a brine pool in the Gulf of Mexico and the role of sulfur oxidizing bacteria in the precipitation of barite (BaSO4) (Thesis). hdl:11299/162398.
- Arias, Francisco J.; Heras, Salvador De Las (2019). "On the feasibility of ocean brine pool power stations". International Journal of Energy Research. 43 (15): 9049–9054. doi:10.1002/er.4708. hdl:2117/170786. ISSN 1099-114X. S2CID 208754788.
- "Brine Pools: The Underwater Lakes of Despair". Amusing Planet. Retrieved 2020-09-28.
- DasSarma, Shiladitya; DasSarma, Priya (2012-03-15). "Halophiles". eLS. Chichester, UK: John Wiley & Sons, Ltd. doi:10.1002/9780470015902.a0000394.pub3. ISBN 978-0-470-01617-6. Retrieved 2020-11-02.
- Antunes, André; Olsson-Francis, Karen; McGenity, Terry J. (2020), "Exploring Deep-Sea Brines as Potential Terrestrial Analogues of Oceans in the Icy Moons of the Outer Solar System", Astrobiology: Current, Evolving, and Emerging Perspectives, Caister Academic Press, vol. 38, pp. 123–162, doi:10.21775/9781912530304.06, ISBN 978-1-912530-30-4, PMID 31967579
- Bougouffa, S.; Yang, J. K.; Lee, O. O.; Wang, Y.; Batang, Z.; Al-Suwailem, A.; Qian, P. Y. (June 2013). "Distinctive Microbial Community Structure in Highly Stratified Deep-Sea Brine Water Columns". Applied and Environmental Microbiology. 79 (11): 3425–3437. Bibcode:2013ApEnM..79.3425B. doi:10.1128/AEM.00254-13. ISSN 0099-2240. PMC 3648036. PMID 23542623.
- Frazer, Jennifer. "Playing in a Deep-Sea Brine Pool Is Fun, as Long as You're an ROV [Video]". Scientific American Blog Network. Retrieved 2020-10-30.
- Oliver, P. Graham; Vestheim, Hege; Antunes, André; Kaartvedt, Stein (May 2015). "Systematics, functional morphology and distribution of a bivalve (Apachecorbula muriatica gen. et sp. nov.) from the rim of the 'Valdivia Deep' brine pool in the Red Sea". Journal of the Marine Biological Association of the United Kingdom. 95 (3): 523–535. doi:10.1017/S0025315414001234. hdl:1822/39421. ISSN 0025-3154. S2CID 55354847.
- Vestheim, Hege; Kaartvedt, Stein (2015-02-26). "A deep sea community at the Kebrit brine pool in the Red Sea". Marine Biodiversity. 46 (1): 59–65. doi:10.1007/s12526-015-0321-0. ISSN 1867-1616. S2CID 16122787.
- Sawyer, Derek E.; Mason, R. Alan; Cook, Ann E.; Portnov, Alexey (2019-01-15). "Submarine Landslides Induce Massive Waves in Subsea Brine Pools". Scientific Reports. 9 (1): 128. Bibcode:2019NatSR...9..128S. doi:10.1038/s41598-018-36781-7. ISSN 2045-2322. PMC 6333809. PMID 30644410. S2CID 58010364.
- Hourdez, Stéphane; Frederick, Lee-Ann; Schernecke, Andrea; Fisher, Charles R. (2001). "Functional respiratory anatomy of a deep-sea orbiniid polychaete from the Brine Pool NR-1 in the Gulf of Mexico". Invertebrate Biology. 120 (1): 29–40. doi:10.1111/j.1744-7410.2001.tb00023.x. ISSN 1744-7410.
- Smith, Emily B.; Scott, Kathleen M.; Nix, Erica R.; Korte, Carol; Fisher, Charles R. (September 2000). "Growth and Condition of Seep Mussels (Bathymodiolus Childressi) at a Gulf of Mexico Brine Pool". Ecology. 81 (9): 2392–2403. doi:10.1890/0012-9658(2000)081[2392:GACOSM]2.0.CO;2. ISSN 0012-9658.
- World Wildlife Fund. "Deep sea ecology: hydrothermal vents and cold seeps". March 23, 2006. Accessed October 3, 2007.
- Zhang, Weipeng; Wang, Yong; Bougouffa, Salim; Tian, Renmao; Cao, Huiluo; Li, Yongxin; Cai, Lin; Wong, Yue Him; Zhang, Gen; Zhou, Guowei; Zhang, Xixiang (2015). "Synchronized dynamics of bacterial niche-specific functions during biofilm development in a cold seep brine pool". Environmental Microbiology. 17 (10): 4089–4104. doi:10.1111/1462-2920.12978. hdl:10754/561085. ISSN 1462-2920. PMID 26171930.
- Hugenholtz, Philip; Goebel, Brett M.; Pace, Norman R. (1998-09-15). "Impact of Culture-Independent Studies on the Emerging Phylogenetic View of Bacterial Diversity". Journal of Bacteriology. 180 (18): 4765–4774. doi:10.1128/JB.180.18.4765-4774.1998. ISSN 0021-9193. PMC 107498. PMID 9733676.
- Simon, Carola; Daniel, Rolf (2011-02-15). "Metagenomic Analyses: Past and Future Trends". Applied and Environmental Microbiology. 77 (4): 1153–1161. Bibcode:2011ApEnM..77.1153S. doi:10.1128/AEM.02345-10. ISSN 0099-2240. PMC 3067235. PMID 21169428.
- Behzad, Hayedeh; Ibarra, Martin Augusto; Mineta, Katsuhiko; Gojobori, Takashi (2016). "Metagenomic studies of the Red Sea". Gene. 576 (2): 717–723. doi:10.1016/j.gene.2015.10.034. hdl:10754/581498. PMID 26526132.
- Bougouffa, S.; Yang, J. K.; Lee, O. O.; Wang, Y.; Batang, Z.; Al-Suwailem, A.; Qian, P. Y. (May 2013). "Distinctive Microbial Community Structure in Highly Stratified Deep-Sea Brine Water Columns". Applied and Environmental Microbiology. 79 (11): 3425–3437. Bibcode:2013ApEnM..79.3425B. doi:10.1128/AEM.00254-13. ISSN 0099-2240. PMC 3648036. PMID 23542623.
- Wang, Yong; Yang, Jiangke; Lee, On On; Dash, Swagatika; Lau, Stanley C K; Al-Suwailem, Abdulaziz; Wong, Tim Y H; Danchin, Antoine; Qian, Pei-Yuan (2011). "Hydrothermally generated aromatic compounds are consumed by bacteria colonizing in Atlantis II Deep of the Red Sea". The ISME Journal. 5 (10): 1652–1659. doi:10.1038/ismej.2011.42. ISSN 1751-7362. PMC 3176515. PMID 21525946.
- Wang, Yong; Cao, Huiluo; Zhang, Guishan; Bougouffa, Salim; Lee, On On; Al-Suwailem, Abdulaziz; Qian, Pei-Yuan (2013). "Autotrophic Microbe Metagenomes and Metabolic Pathways Differentiate Adjacent Red Sea Brine Pools". Scientific Reports. 3 (1): 1748. Bibcode:2013NatSR...3E1748W. doi:10.1038/srep01748. ISSN 2045-2322. PMC 3638166. PMID 23624511.
- Mei, Yunjun; Liu, Huan; Zhang, Shunxi; Yang, Ming; Hu, Chun; Zhang, Jian; Shen, Ping; Chen, Xiangdong (2017-09-19). "Effects of salinity on the cellular physiological responses of Natrinema sp. J7-2". PLOS ONE. 12 (9): e0184974. Bibcode:2017PLoSO..1284974M. doi:10.1371/journal.pone.0184974. ISSN 1932-6203. PMC 5604999. PMID 28926633.
- Hallsworth, John E.; Yakimov, Michail M.; Golyshin, Peter N.; Gillion, Jenny L. M.; D'Auria, Giuseppe; de Lima Alves, Flavia; La Cono, Violetta; Genovese, Maria; McKew, Boyd A.; Hayes, Sandra L.; Harris, Gayle (2007). "Limits of life in MgCl2-containing environments: chaotropicity defines the window". Environmental Microbiology. 9 (3): 801–813. doi:10.1111/j.1462-2920.2006.01212.x. ISSN 1462-2912. PMID 17298378.
- Hallsworth, John E.; Heim, Sabina; Timmis, Kenneth N. (2003-11-27). "Chaotropic solutes cause water stress in Pseudomonas putida". Environmental Microbiology. 5 (12): 1270–1280. doi:10.1111/j.1462-2920.2003.00478.x. ISSN 1462-2912. PMID 14641573.
- Oren, Aharon (2010-11-05). "Thermodynamic limits to microbial life at high salt concentrations". Environmental Microbiology. 13 (8): 1908–1923. doi:10.1111/j.1462-2920.2010.02365.x. ISSN 1462-2912. PMID 21054738.
- Gunde-Cimerman, Nina; Plemenitaš, Ana; Oren, Aharon (2018-05-01). "Strategies of adaptation of microorganisms of the three domains of life to high salt concentrations". FEMS Microbiology Reviews. 42 (3): 353–375. doi:10.1093/femsre/fuy009. ISSN 0168-6445. PMID 29529204.
- Tanne, Christoph; Golovina, Elena A.; Hoekstra, Folkert A.; Meffert, Andrea; Galinski, Erwin A. (2014-04-04). "Glass-forming property of hydroxyectoine is the cause of its superior function as a desiccation protectant". Frontiers in Microbiology. 5: 150. doi:10.3389/fmicb.2014.00150. ISSN 1664-302X. PMC 3983491. PMID 24772110.
- Kamanda Ngugi, David; Blom, Jochen; Alam, Intikhab; Rashid, Mamoon; Ba-Alawi, Wail; Zhang, Guishan; Hikmawan, Tyas; Guan, Yue; Antunes, Andre; Siam, Rania; El Dorry, Hamza (2014-08-08). "Comparative genomics reveals adaptations of a halotolerant thaumarchaeon in the interfaces of brine pools in the Red Sea". The ISME Journal. 9 (2): 396–411. doi:10.1038/ismej.2014.137. ISSN 1751-7362. PMC 4303633. PMID 25105904.
- Ngugi, David Kamanda; Blom, Jochen; Stepanauskas, Ramunas; Stingl, Ulrich (2015-12-11). "Diversification and niche adaptations of Nitrospina-like bacteria in the polyextreme interfaces of Red Sea brines". The ISME Journal. 10 (6): 1383–1399. doi:10.1038/ismej.2015.214. ISSN 1751-7362. PMC 5029188. PMID 26657763.
- Kato, C.; Qureshi, M. (1999). "Pressure response in deep-sea piezophilic bacteria". Journal of Molecular Microbiology and Biotechnology. 1 (1): 87–92. doi:10.1023/A:1008989800098. PMID 10941789. S2CID 32898991.
- Van Cappellen, Philippe; Viollier, Eric; Roychoudhury, Alakendra; Clark, Lauren; Ingall, Ellery; Lowe, Kristine; Dichristina, Thomas (1998-08-21). "Biogeochemical Cycles of Manganese and Iron at the Oxic−Anoxic Transition of a Stratified Marine Basin (Orca Basin, Gulf of Mexico)". Environmental Science & Technology. 32 (19): 2931–2939. Bibcode:1998EnST...32.2931V. doi:10.1021/es980307m. ISSN 0013-936X.
- Guan, Yue; Hikmawan, Tyas; Antunes, André; Ngugi, David; Stingl, Ulrich (2015). "Diversity of methanogens and sulfate-reducing bacteria in the interfaces of five deep-sea anoxic brines of the Red Sea". Research in Microbiology. 166 (9): 688–699. doi:10.1016/j.resmic.2015.07.002. ISSN 0923-2508. PMID 26192212.
- Borin, Sara; Mapelli, Francesca; Rolli, Eleonora; Song, Bongkeun; Tobias, Craig; Schmid, Markus C.; De Lange, Gert J.; Reichart, Gert J.; Schouten, Stefan; Jetten, Mike; Daffonchio, Daniele (2013-01-23). "Anammox bacterial populations in deep marine hypersaline gradient systems". Extremophiles. 17 (2): 289–299. doi:10.1007/s00792-013-0516-x. hdl:2066/111535. ISSN 1431-0651. PMID 23340764. S2CID 12353717.
- Reinhardt, Danovaro, Roberto Dell'Anno, Antonio Pusceddu, Antonio Gambi, Cristina Heiner, Iben Møbjerg Kristensen (2010-04-06). The first metazoa living in permanently anoxic conditions. BioMed Central Ltd. OCLC 808847572.
{{cite book}}: CS1 maint: multiple names: authors list (link) - Neves, Ricardo Cardoso; Gambi, Cristina; Danovaro, Roberto; Kristensen, Reinhardt Møbjerg (2014-08-08). "Spinoloricus cinziae (Phylum Loricifera), a new species from a hypersaline anoxic deep basin in the Mediterranean Sea". Systematics and Biodiversity. 12 (4): 489–502. doi:10.1080/14772000.2014.943820. ISSN 1477-2000. S2CID 84682701.
- Danovaro, Roberto; Gambi, Cristina; Dell’Anno, Antonio; Corinaldesi, Cinzia; Pusceddu, Antonio; Neves, Ricardo Cardoso; Kristensen, Reinhardt Møbjerg (2016-06-07). "The challenge of proving the existence of metazoan life in permanently anoxic deep-sea sediments". BMC Biology. 14 (1): 43. doi:10.1186/s12915-016-0263-4. ISSN 1741-7007. PMC 4895820. PMID 27267928.
- Merlino, Giuseppe; Barozzi, Alan; Michoud, Grégoire; Ngugi, David Kamanda; Daffonchio, Daniele (2018-07-01). "Microbial ecology of deep-sea hypersaline anoxic basins". FEMS Microbiology Ecology. 94 (7): fiy085. doi:10.1093/femsec/fiy085. ISSN 0168-6496. PMID 29905791.
- Bougouffa, S.; Yang, J. K.; Lee, O. O.; Wang, Y.; Batang, Z.; Al-Suwailem, A.; Qian, P. Y. (May 2013). "Distinctive Microbial Community Structure in Highly Stratified Deep-Sea Brine Water Columns". Applied and Environmental Microbiology. 79 (11): 3425–3437. Bibcode:2013ApEnM..79.3425B. doi:10.1128/AEM.00254-13. ISSN 0099-2240. PMC 3648036. PMID 23542623.
- Anschutz, Pierre; Blanc, Gérard (1996-07-01). "Heat and salt fluxes in the Atlantis II Deep (Red Sea)". Earth and Planetary Science Letters. 142 (1): 147–159. Bibcode:1996E&PSL.142..147A. doi:10.1016/0012-821X(96)00098-2. ISSN 0012-821X.
- De Lange, G. J; Middelburg, J. J; Van der Weijden, C. H; Catalano, G; Luther, G. W; Hydes, D. J; Woittiez, J. R. W; Klinkhammer, G. P (1990-12-10). "Composition of anoxic hypersaline brines in the Tyro and Bannock Basins, eastern Mediterranean". Marine Chemistry. Anoxic Brines of the Mediterranean Sea. 31 (1): 63–88. doi:10.1016/0304-4203(90)90031-7. ISSN 0304-4203.
- van der Wielen, Paul W. J. J.; Bolhuis, Henk; Borin, Sara; Daffonchio, Daniele; Corselli, Cesare; Giuliano, Laura; D'Auria, Giuseppe; de Lange, Gert J.; Huebner, Andreas; Varnavas, Sotirios P.; Thomson, John (2005-01-07). "The Enigma of Prokaryotic Life in Deep Hypersaline Anoxic Basins". Science. 307 (5706): 121–123. Bibcode:2005Sci...307..121V. doi:10.1126/science.1103569. ISSN 0036-8075. PMID 15637281. S2CID 206507712.
- Borin, Sara; Brusetti, Lorenzo; Mapelli, Francesca; D'Auria, Giuseppe; Brusa, Tullio; Marzorati, Massimo; Rizzi, Aurora; Yakimov, Michail; Marty, Danielle; De Lange, Gert J.; Van der Wielen, Paul (2009-06-09). "Sulfur cycling and methanogenesis primarily drive microbial colonization of the highly sulfidic Urania deep hypersaline basin". Proceedings of the National Academy of Sciences of the United States of America. 106 (23): 9151–9156. Bibcode:2009PNAS..106.9151B. doi:10.1073/pnas.0811984106. ISSN 0027-8424. PMC 2685740. PMID 19470485.
- La Cono, Violetta; Smedile, Francesco; Bortoluzzi, Giovanni; Arcadi, Erika; Maimone, Giovanna; Messina, Enzo; Borghini, Mireno; Oliveri, Elvira; Mazzola, Salvatore; L'Haridon, Stephan; Toffin, Laurent (August 2011). "Unveiling microbial life in new deep-sea hypersaline Lake Thetis. Part I: Prokaryotes and environmental settings". Environmental Microbiology. 13 (8): 2250–2268. doi:10.1111/j.1462-2920.2011.02478.x. ISSN 1462-2920. PMID 21518212.
- Yakimov, Michail M.; La Cono, Violetta; Spada, Gina L.; Bortoluzzi, Giovanni; Messina, Enzo; Smedile, Francesco; Arcadi, Erika; Borghini, Mireno; Ferrer, Manuel; Schmitt-Kopplin, Phillippe; Hertkorn, Norbert (2015-01-26). "Microbial community of the deep-sea brine LakeKryosseawater-brine interface is active below the chaotropicity limit of life as revealed by recovery of mRNA". Environmental Microbiology. 17 (2): 364–382. doi:10.1111/1462-2920.12587. ISSN 1462-2912. PMID 25622758. S2CID 31463948.
- Van Der Wielen, Paul W. J. J. (2006-06-01). "Diversity of ribulose-1,5-bisphosphate carboxylase/oxygenase large-subunit genes in the MgCl2-dominated deep hypersaline anoxic basin discovery". FEMS Microbiology Letters. 259 (2): 326–331. doi:10.1111/j.1574-6968.2006.00284.x. ISSN 0378-1097. PMID 16734797.
- Oren, Aharon (June 1999). "Bioenergetic Aspects of Halophilism". Microbiology and Molecular Biology Reviews. 63 (2): 334–348. doi:10.1128/MMBR.63.2.334-348.1999. ISSN 1092-2172. PMC 98969. PMID 10357854.
- Yakimov, Michail M.; La Cono, Violetta; Slepak, Vladlen Z.; La Spada, Gina; Arcadi, Erika; Messina, Enzo; Borghini, Mireno; Monticelli, Luis S.; Rojo, David; Barbas, Coral; Golyshina, Olga V. (2013-12-19). "Microbial life in the Lake Medee, the largest deep-sea salt-saturated formation". Scientific Reports. 3 (1): 3554. Bibcode:2013NatSR...3E3554Y. doi:10.1038/srep03554. ISSN 2045-2322. PMC 3867751. PMID 24352146.
- Borin, Sara; Mapelli, Francesca; Rolli, Eleonora; Song, Bongkeun; Tobias, Craig; Schmid, Markus C.; De Lange, Gert J.; Reichart, Gert J.; Schouten, Stefan; Jetten, Mike; Daffonchio, Daniele (March 2013). "Anammox bacterial populations in deep marine hypersaline gradient systems". Extremophiles: Life Under Extreme Conditions. 17 (2): 289–299. doi:10.1007/s00792-013-0516-x. hdl:2066/111535. ISSN 1433-4909. PMID 23340764. S2CID 12353717.
- Pachiadaki, Maria G.; Yakimov, Michail M.; LaCono, Violetta; Leadbetter, Edward; Edgcomb, Virginia (June 2014). "Unveiling microbial activities along the halocline of Thetis, a deep-sea hypersaline anoxic basin". The ISME Journal. 8 (12): 2478–2489. doi:10.1038/ismej.2014.100. ISSN 1751-7370. PMC 4260694. PMID 24950109.
- Guan, Yue; Hikmawan, Tyas; Antunes, André; Ngugi, David; Stingl, Ulrich (2015-11-01). "Diversity of methanogens and sulfate-reducing bacteria in the interfaces of five deep-sea anoxic brines of the Red Sea". Research in Microbiology. Deep Sea Microbiology. 166 (9): 688–699. doi:10.1016/j.resmic.2015.07.002. ISSN 0923-2508. PMID 26192212.
- Friedrich, Cornelius G.; Rother, Dagmar; Bardischewsky, Frank; Quentmeier, Armin; Fischer, Jörg (July 2001). "Oxidation of Reduced Inorganic Sulfur Compounds by Bacteria: Emergence of a Common Mechanism?". Applied and Environmental Microbiology. 67 (7): 2873–2882. Bibcode:2001ApEnM..67.2873F. doi:10.1128/AEM.67.7.2873-2882.2001. ISSN 0099-2240. PMC 92956. PMID 11425697.
- Sorokin, Dimitry Y; Messina, Enzo; Smedile, Francesco; Roman, Pawel; Damsté, Jaap S Sinninghe; Ciordia, Sergio; Mena, Maria Carmen; Ferrer, Manuel; Golyshin, Peter N; Kublanov, Ilya V; Samarov, Nazar I (January 2017). "Discovery of anaerobic lithoheterotrophic haloarchaea, ubiquitous in hypersaline habitats". The ISME Journal. 11 (5): 1245–1260. doi:10.1038/ismej.2016.203. ISSN 1751-7362. PMC 5437934. PMID 28106880.
- "NOAA Ocean Explorer: Gulf of Mexico 2002". oceanexplorer.noaa.gov. Retrieved 2022-03-29.
- Duperron, Sébastien; Lorion, Julien; Samadi, Sarah; Gros, Olivier; Gaill, Françoise (2009). "Symbioses between deep-sea mussels (Mytilidae: Bathymodiolinae) and chemosynthetic bacteria: diversity, function and evolution". Comptes Rendus Biologies. 332 (2–3): 298–310. doi:10.1016/j.crvi.2008.08.003. PMID 19281960.
- Nougué, Odrade; Gallet, Romain; Chevin, Luis-Miguel; Lenormand, Thomas (2015). "Niche Limits of Symbiotic Gut Microbiota Constrain the Salinity Tolerance of Brine Shrimp". The American Naturalist. 186 (3): 390–403. doi:10.1086/682370. ISSN 0003-0147. PMID 26655356. S2CID 44744283.
- Wang, Yong; Zhang, Wei Peng; Cao, Hui Luo; Shek, Chun Shum; Tian, Ren Mao; Wong, Yue Him; Batang, Zenon; Al-Suwailem, Abdulaziz; Qian, Pei-Yuan (2014). "Diversity and distribution of eukaryotic microbes in and around a brine pool adjacent to the Thuwal cold seeps in the Red Sea". Frontiers in Microbiology. 5: 37. doi:10.3389/fmicb.2014.00037. ISSN 1664-302X. PMC 3922051. PMID 24575081.
- Duarte, Carlos M.; Røstad, Anders; Michoud, Grégoire; Barozzi, Alan; Merlino, Giuseppe; Delgado-Huertas, Antonio; Hession, Brian C.; Mallon, Francis L.; Afifi, Abdulakader M.; Daffonchio, Daniele (2020-01-22). "Discovery of Afifi, the shallowest and southernmost brine pool reported in the Red Sea". Scientific Reports. 10 (1): 910. Bibcode:2020NatSR..10..910D. doi:10.1038/s41598-020-57416-w. ISSN 2045-2322. PMC 6976674. PMID 31969577. S2CID 210844928.
- Wang, Yong; Yang, Jiang Ke; Lee, On On; Li, Tie Gang; Al-Suwailem, Abdulaziz; Danchin, Antoine; Qian, Pei-Yuan (2011-12-21). "Bacterial Niche-Specific Genome Expansion Is Coupled with Highly Frequent Gene Disruptions in Deep-Sea Sediments". PLOS ONE. 6 (12): e29149. Bibcode:2011PLoSO...629149W. doi:10.1371/journal.pone.0029149. ISSN 1932-6203. PMC 3244439. PMID 22216192.
- Salem, Mohamed (2017-06-01). "Study of Conrad and Shaban deep brines, Red Sea, using bathymetric, parasound and seismic surveys". NRIAG Journal of Astronomy and Geophysics. 6 (1): 90–96. Bibcode:2017JAsGe...6...90S. doi:10.1016/j.nrjag.2017.04.003. ISSN 2090-9977. S2CID 132353952.
- Siam, Rania; Mustafa, Ghada A.; Sharaf, Hazem; Moustafa, Ahmed; Ramadan, Adham R.; Antunes, Andre; Bajic, Vladimir B.; Stingl, Uli; Marsis, Nardine G. R.; Coolen, Marco J. L.; Sogin, Mitchell (2012-08-20). "Unique Prokaryotic Consortia in Geochemically Distinct Sediments from Red Sea Atlantis II and Discovery Deep Brine Pools". PLOS ONE. 7 (8): e42872. Bibcode:2012PLoSO...742872S. doi:10.1371/journal.pone.0042872. ISSN 1932-6203. PMC 3423430. PMID 22916172.
- Abdallah, Rehab Z.; Adel, Mustafa; Ouf, Amged; Sayed, Ahmed; Ghazy, Mohamed A.; Alam, Intikhab; Essack, Magbubah; Lafi, Feras F.; Bajic, Vladimir B.; El-Dorry, Hamza; Siam, Rania (2014). "Aerobic methanotrophic communities at the Red Sea brine-seawater interface". Frontiers in Microbiology. 5: 487. doi:10.3389/fmicb.2014.00487. ISSN 1664-302X. PMC 4172156. PMID 25295031.
- Yakimov, Michail M.; Cono, Violetta La; Spada, Gina L.; Bortoluzzi, Giovanni; Messina, Enzo; Smedile, Francesco; Arcadi, Erika; Borghini, Mireno; Ferrer, Manuel; Schmitt‐Kopplin, Phillippe; Hertkorn, Norbert (2015). "Microbial community of the deep-sea brine Lake Kryos seawater–brine interface is active below the chaotropicity limit of life as revealed by recovery of mRNA". Environmental Microbiology. 17 (2): 364–382. doi:10.1111/1462-2920.12587. ISSN 1462-2920. PMID 25622758. S2CID 31463948.
- Salem, Mohamed (2017-06-01). "Study of Conrad and Shaban deep brines, Red Sea, using bathymetric, parasound and seismic surveys". NRIAG Journal of Astronomy and Geophysics. 6 (1): 90–96. Bibcode:2017JAsGe...6...90S. doi:10.1016/j.nrjag.2017.04.003. ISSN 2090-9977. S2CID 132353952.
- Nazarious, Miracle Israel; Ramachandran, Abhilash Vakkada; Zorzano, Maria-Paz; Martin-Torres, Javier (2019-09-01). "Calibration and preliminary tests of the Brine Observation Transition To Liquid Experiment on HABIT/ExoMars 2020 for demonstration of liquid water stability on Mars". Acta Astronautica. 162: 497–510. Bibcode:2019AcAau.162..497N. doi:10.1016/j.actaastro.2019.06.026. ISSN 0094-5765.
- Li, Dehai; Wang, Fengping; Xiao, Xiang; Zeng, Xiang; Gu, Qian-Qun; Zhu, Weiming (May 2007). "A new cytotoxic phenazine derivative from a deep sea bacterium Bacillus sp". Archives of Pharmacal Research. 30 (5): 552–555. doi:10.1007/BF02977647. ISSN 0253-6269. PMID 17615672. S2CID 10515104.
- Ziko, Laila; Saqr, Al-Hussein A.; Ouf, Amged; Gimpel, Matthias; Aziz, Ramy K.; Neubauer, Peter; Siam, Rania (2019-03-18). "Antibacterial and anticancer activities of orphan biosynthetic gene clusters from Atlantis II Red Sea brine pool". Microbial Cell Factories. 18 (1): 56. doi:10.1186/s12934-019-1103-3. ISSN 1475-2859. PMC 6423787. PMID 30885206.
- Craig, H. (1966-12-23). "Isotopic Composition and Origin of the Red Sea and Salton Sea Geothermal Brines". Science. 154 (3756): 1544–1548. Bibcode:1966Sci...154.1544C. doi:10.1126/science.154.3756.1544. ISSN 0036-8075. PMID 17807292. S2CID 40574864.
- Sagar, Sunil; Esau, Luke; Hikmawan, Tyas; Antunes, Andre; Holtermann, Karie; Stingl, Ulrich; Bajic, Vladimir B.; Kaur, Mandeep (2013-02-06). "Cytotoxic and apoptotic evaluations of marine bacteria isolated from brine-seawater interface of the Red Sea". BMC Complementary and Alternative Medicine. 13 (1): 29. doi:10.1186/1472-6882-13-29. ISSN 1472-6882. PMC 3598566. PMID 23388148.
- Grötzinger, Stefan Wolfgang; Alam, Intikhab; Alawi, Wail Ba; Bajic, Vladimir B.; Stingl, Ulrich; Eppinger, Jörg (2014). "Mining a database of single amplified genomes from Red Sea brine pool extremophiles—improving reliability of gene function prediction using a profile and pattern matching algorithm (PPMA)". Frontiers in Microbiology. 5: 134. doi:10.3389/fmicb.2014.00134. ISSN 1664-302X. PMC 3985023. PMID 24778629.
- Ziko, Laila; Saqr, Al-Hussein A.; Ouf, Amged; Gimpel, Matthias; Aziz, Ramy K.; Neubauer, Peter; Siam, Rania (2019-03-18). "Antibacterial and anticancer activities of orphan biosynthetic gene clusters from Atlantis II Red Sea brine pool". Microbial Cell Factories. 18 (1): 56. doi:10.1186/s12934-019-1103-3. ISSN 1475-2859. PMC 6423787. PMID 30885206.
- Elbehery, Ali H. A.; Leak, David J.; Siam, Rania (2016-12-22). "Novel thermostable antibiotic resistance enzymes from the Atlantis II Deep Red Sea brine pool". Microbial Biotechnology. 10 (1): 189–202. doi:10.1111/1751-7915.12468. ISSN 1751-7915. PMC 5270753. PMID 28004885.
Further reading
- Boetius, A.; Joye, S. (2009-06-18). "Thriving in Salt". Science. 324 (5934): 1523–1525. doi:10.1126/science.1172979. ISSN 0036-8075.
- Eder, W., Jahnke, L. L., Schmidt, M., & Huber, R. (2001). Microbial Diversity of the Brine-Seawater Interface of the Kebrit Deep, Red Sea, Studied via 16S rRNA Gene Sequences and Cultivation Methods. Applied and Environmental Microbiology, 67(7), 3077–3085. doi:10.1128/aem.67.7.3077-3085.2001
- Guan, Y., Hikmawan, T., Antunes, A., Ngugi, D., & Stingl, U. (2015). Diversity of methanogens and sulfate-reducing bacteria in the interfaces of five deep-sea anoxic brines of the Red Sea. Research in Microbiology, 166(9), 688–699. doi:10.1016/j.resmic.2015.07.002
- Hartmann, M., Scholten, J., Stoffers, P., & Wehner, F. (1998). Hydrographic structure of brine-filled deeps in the Red Sea—new results from the Shaban, Kebrit, Atlantis II, and Discovery Deep. Marine Geology, 144(4), 311–330. doi:10.1016/s0025-3227(97)00055-8
- Patowary, K. (2018, November 7). Brine Pools: The Underwater Lakes of Despair. Retrieved October 28, 2020, from https://www.amusingplanet.com/2018/11/brine-pools-lakes-under-ocean.html
- US Department of Commerce, N. (n.d.). Gulf of Mexico 2002. Retrieved October 28, 2020, from https://oceanexplorer.noaa.gov/explorations/02mexico/welcome.html
- Wankel, S. D., Joye, S. B., Samarkin, V. A., Shah, S. R., Friederich, G., Melas-Kyriazi, J., & Girguis, P. R. (2010). New constraints on methane fluxes and rates of anaerobic methane oxidation in a Gulf of Mexico brine pool via in situ mass spectrometry. Deep Sea Research Part II: Topical Studies in Oceanography, 57(21-23), 2022–2029. doi:10.1016/j.dsr2.2010.05.009
