Combinatorial chemistry
Combinatorial chemistry comprises chemical synthetic methods that make it possible to prepare a large number (tens to thousands or even millions) of compounds in a single process. These compound libraries can be made as mixtures, sets of individual compounds or chemical structures generated by computer software.[1] Combinatorial chemistry can be used for the synthesis of small molecules and for peptides.
Strategies that allow identification of useful components of the libraries are also part of combinatorial chemistry. The methods used in combinatorial chemistry are applied outside chemistry, too.
History
Combinatorial chemistry had been invented by Furka Á (Eötvös Loránd University Budapest Hungary) who described the principle of it, the combinatorial synthesis and a deconvolution procedure in a document that was notarized in 1982.[2] The principle of the combinatorial method is: synthesize a multi-component compound mixture (combinatorial library) in a single stepwise procedure and screen it to find drug candidates or other kinds of useful compounds also in a single process. The most important innovation of the combinatorial method is to use mixtures in the synthesis and screening that ensures the high productivity of the process. Motivations that led to the invention had been published in 2002.[3]
Introduction
Synthesis of molecules in a combinatorial fashion can quickly lead to large numbers of molecules. For example, a molecule with three points of diversity (R1, R2, and R3) can generate possible structures, where , , and are the numbers of different substituents utilized.[2]
The basic principle of combinatorial chemistry is to prepare libraries of a very large number of compounds then identify the useful components of the libraries.
Although combinatorial chemistry has only really been taken up by industry since the 1990s,[4] its roots can be seen as far back as the 1960s when a researcher at Rockefeller University, Bruce Merrifield, started investigating the solid-phase synthesis of peptides.
In its modern form, combinatorial chemistry has probably had its biggest impact in the pharmaceutical industry.[5] Researchers attempting to optimize the activity profile of a compound create a 'library' of many different but related compounds.[6][7] Advances in robotics have led to an industrial approach to combinatorial synthesis, enabling companies to routinely produce over 100,000 new and unique compounds per year.[8]
In order to handle the vast number of structural possibilities, researchers often create a 'virtual library', a computational enumeration of all possible structures of a given pharmacophore with all available reactants.[9] Such a library can consist of thousands to millions of 'virtual' compounds. The researcher will select a subset of the 'virtual library' for actual synthesis, based upon various calculations and criteria (see ADME, computational chemistry, and QSAR).
Polymers (peptides and oligonucleotides)
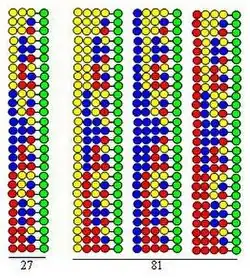
Combinatorial split-mix (split and pool) synthesis
Combinatorial split-mix (split and pool) synthesis [10][11] is based on the solid-phase synthesis developed by Merrifield.[12] If a combinatorial peptide library is synthesized using 20 amino acids (or other kinds of building blocks) the bead form solid support is divided into 20 equal portions. This is followed by coupling a different amino acid to each portion. The third step is the mixing of all portions. These three steps comprise a cycle. Elongation of the peptide chains can be realized by simply repeating the steps of the cycle.
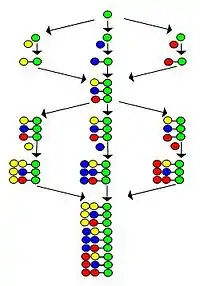
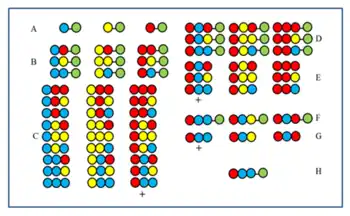
The procedure is illustrated by the synthesis of a dipeptide library using the same three amino acids as building blocks in both cycles. Each component of this library contains two amino acids arranged in different orders. The amino acids used in couplings are represented by yellow, blue and red circles in the figure. Divergent arrows show dividing solid support resin (green circles) into equal portions, vertical arrows mean coupling and convergent arrows represent mixing and homogenizing the portions of the support.
The figure shows that in the two synthetic cycles 9 dipeptides are formed. In the third and fourth cycles, 27 tripeptides and 81 tetrapeptides would form, respectively.
The "split-mix synthesis" has several outstanding features:
- It is highly efficient. As the figure demonstrates the number of peptides formed in the synthetic process (3, 9, 27, 81) increases exponentially with the number of executed cycles. Using 20 amino acids in each synthetic cycle the number of formed peptides are: 400, 8,000, 160,000 and 3,200,000, respectively. This means that the number of peptides increases exponentially with the number of the executed cycles.
- All peptide sequences are formed in the process that can be deduced by a combination of the amino acids used in the cycles.
- Portioning of the support into equal samples assures formation of the components of the library in nearly equal molar quantities.
- Only a single peptide forms on each bead of the support. This is the consequence of using only one amino acid in the coupling steps. It is completely unknown, however, which is the peptide that occupies a selected bead.
- The split-mix method can be used for the synthesis of organic or any other kind of library that can be prepared from its building blocks in a stepwise process.
In 1990 three groups described methods for preparing peptide libraries by biological methods[13][14][15] and one year later Fodor et al. published a remarkable method for synthesis of peptide arrays on small glass slides.[16]
A "parallel synthesis" method was developed by Mario Geysen and his colleagues for preparation of peptide arrays.[17] They synthesized 96 peptides on plastic rods (pins) coated at their ends with the solid support. The pins were immersed into the solution of reagents placed in the wells of a microtiter plate. The method is widely applied particularly by using automatic parallel synthesizers. Although the parallel method is much slower than the real combinatorial one, its advantage is that it is exactly known which peptide or other compound forms on each pin.
Further procedures were developed to combine the advantages of both split-mix and parallel synthesis. In the method described by two groups[18][19] the solid support was enclosed into permeable plastic capsules together with a radiofrequency tag that carried the code of the compound to be formed in the capsule. The procedure was carried out similar to the split-mix method. In the split step, however, the capsules were distributed among the reaction vessels according to the codes read from the radiofrequency tags of the capsules.
A different method for the same purpose was developed by Furka et al.[20] is named "string synthesis". In this method, the capsules carried no code. They are strung like the pearls in a necklace and placed into the reaction vessels in stringed form. The identity of the capsules, as well as their contents, are stored by their position occupied on the strings. After each coupling step, the capsules are redistributed among new strings according to definite rules.
Small molecules
In the drug discovery process, the synthesis and biological evaluation of small molecules of interest have typically been a long and laborious process. Combinatorial chemistry has emerged in recent decades as an approach to quickly and efficiently synthesize large numbers of potential small molecule drug candidates. In a typical synthesis, only a single target molecule is produced at the end of a synthetic scheme, with each step in a synthesis producing only a single product. In a combinatorial synthesis, when using only single starting material, it is possible to synthesize a large library of molecules using identical reaction conditions that can then be screened for their biological activity. This pool of products is then split into three equal portions containing each of the three products, and then each of the three individual pools is then reacted with another unit of reagent B, C, or D, producing 9 unique compounds from the previous 3. This process is then repeated until the desired number of building blocks is added, generating many compounds. When synthesizing a library of compounds by a multi-step synthesis, efficient reaction methods must be employed, and if traditional purification methods are used after each reaction step, yields and efficiency will suffer.
Solid-phase synthesis offers potential solutions to obviate the need for typical quenching and purification steps often used in synthetic chemistry. In general, a starting molecule is adhered to a solid support (typically an insoluble polymer), then additional reactions are performed, and the final product is purified and then cleaved from the solid support. Since the molecules of interest are attached to a solid support, it is possible to reduce the purification after each reaction to a single filtration/wash step, eliminating the need for tedious liquid-liquid extraction and solvent evaporation steps that most synthetic chemistry involves. Furthermore, by using heterogeneous reactants, excess reagents can be used to drive sluggish reactions to completion, which can further improve yields. Excess reagents can simply be washed away without the need for additional purification steps such as chromatography.

Over the years, a variety of methods have been developed to refine the use of solid-phase organic synthesis in combinatorial chemistry, including efforts to increase the ease of synthesis and purification, as well as non-traditional methods to characterize intermediate products. Although the majority of the examples described here will employ heterogeneous reaction media in every reaction step, Booth and Hodges provide an early example of using solid-supported reagents only during the purification step of traditional solution-phase syntheses.[21] In their view, solution-phase chemistry offers the advantages of avoiding attachment and cleavage reactions necessary to anchor and remove molecules to resins as well as eliminating the need to recreate solid-phase analogues of established solution-phase reactions.
The single purification step at the end of a synthesis allows one or more impurities to be removed, assuming the chemical structure of the offending impurity is known. While the use of solid-supported reagents greatly simplifies the synthesis of compounds, many combinatorial syntheses require multiple steps, each of which still requires some form of purification. Armstrong, et al. describe a one-pot method for generating combinatorial libraries, called multiple-component condensations (MCCs).[22] In this scheme, three or more reagents react such that each reagent is incorporated into the final product in a single step, eliminating the need for a multi-step synthesis that involves many purification steps. In MCCs, there is no deconvolution required to determine which compounds are biologically active because each synthesis in an array has only a single product, thus the identity of the compound should be unequivocally known.
In another array synthesis, Still generated a large library of oligopeptides by split synthesis.[23] The drawback to making many thousands of compounds is that it is difficult to determine the structure of the formed compounds. Their solution is to use molecular tags, where a tiny amount (1 pmol/bead) of a dye is attached to the beads, and the identity of a certain bead can be determined by analyzing which tags are present on the bead. Despite how easy attaching tags makes identification of receptors, it would be quite impossible to individually screen each compound for its receptor binding ability, so a dye was attached to each receptor, such that only those receptors that bind to their substrate produce a color change.
When many reactions need to be run in an array (such as the 96 reactions described in one of Armstrong's MCC arrays), some of the more tedious aspects of synthesis can be automated to improve efficiency. DeWitt and Czarnik detail a method called the "DIVERSOMER method," in which many miniaturized versions of chemical reactions are all run simultaneously.[24] This method uses a device that automates the resin loading and wash cycles, as well as the reaction cycle monitoring and purification, and demonstrate the feasibility of their method and apparatus by using it to synthesize a variety of molecule classes, such as hydantoins and benzodiazepines, running 40 individual reactions in most cases.
Oftentimes, it is not possible to use expensive equipment, and Schwabacher, et al. describe a simple method of combining parallel synthesis of library members and evaluation of entire libraries of compounds.[25] In their method, a thread that is partitioned into different regions is wrapped around a cylinder, where a different reagent is then coupled to each region which bears only a single species. The thread is then re-divided and wrapped around a cylinder of a different size, and this process is then repeated. The beauty of this method is that the identity of each product can be known simply by its location along the thread, and the corresponding biological activity is identified by Fourier transformation of fluorescence signals.
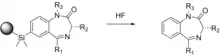
In most of the syntheses described here, it is necessary to attach and remove the starting reagent to/from a solid support. This can lead to the generation of a hydroxyl group, which can potentially affect the biological activity of a target compound. Ellman uses solid phase supports in a multi-step synthesis scheme to obtain 192 individual 1,4-benzodiazepine derivatives, which are well-known therapeutic agents.[26] To eliminate the possibility of potential hydroxyl group interference, a novel method using silyl-aryl chemistry is used to link the molecules to the solid support which cleaves from the support and leaves no trace of the linker.
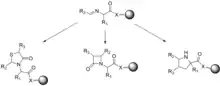
When anchoring a molecule to a solid support, intermediates cannot be isolated from one another without cleaving the molecule from the resin. Since many of the traditional characterization techniques used to track reaction progress and confirm product structure are solution-based, different techniques must be used. Gel-phase 13 C NMR spectroscopy, MALDI mass spectrometry, and IR spectroscopy have been used to confirm structure and monitor the progress of solid-phase reactions.[27] Gordon et al., describe several case studies that utilize imines and peptidyl phosphonates to generate combinatorial libraries of small molecules.[27] To generate the imine library, an amino acid tethered to a resin is reacted in the presence of an aldehyde. The authors demonstrate the use of fast 13 C gel phase NMR spectroscopy and magic angle spinning 1 H NMR spectroscopy to monitor the progress of reactions and showed that most imines could be formed in as little as 10 minutes at room temperature when trimethyl orthoformate was used as the solvent. The formed imines were then derivatized to generate 4-thiazolidinones, B-lactams, and pyrrolidines.
The use of solid-phase supports greatly simplifies the synthesis of large combinatorial libraries of compounds. This is done by anchoring a starting material to a solid support and then running subsequent reactions until a sufficiently large library is built, after which the products are cleaved from the support. The use of solid-phase purification has also been demonstrated for use in solution-phase synthesis schemes in conjunction with standard liquid-liquid extraction purification techniques.
Deconvolution and screening
Combinatorial libraries
Combinatorial libraries are special multi-component mixtures of small-molecule chemical compounds that are synthesized in a single stepwise process. They differ from collection of individual compounds as well as from series of compounds prepared by parallel synthesis. It is an important feature that mixtures are used in their synthesis. The use of mixtures ensures the very high efficiency of the process. Both reactants can be mixtures and in this case the procedure would be even more efficient. For practical reasons however, it is advisable to use the split-mix method in which one of two mixtures is replaced by single building blocks (BBs). The mixtures are so important that there are no combinatorial libraries without using mixture in the synthesis, and if a mixture is used in a process inevitably combinatorial library forms. The split-mix synthesis is usually realized using solid support but it is possible to apply it in solution, too. Since he structures the components are unknown deconvolution methods need to be used in screening. One of the most important features of combinatorial libraries is that the whole mixture can be screened in a single process. This makes these libraries very useful in pharmaceutical research. Partial libraries of full combinatorial libraries can also be synthesized. Some of them can be used in deconvolution[28]
Deconvolution of libraries cleaved from the solid support
If the synthesized molecules of a combinatorial library are cleaved from the solid support a soluble mixture forms. In such solution, millions of different compounds may be found. When this synthetic method was developed, it first seemed impossible to identify the molecules, and to find molecules with useful properties. Strategies for identification of the useful components had been developed, however, to solve the problem. All these strategies are based on synthesis and testing of partial libraries. The earliest iterative strategy is described in the above-mentioned document of Furka notarized in 1982 and.[2] The method was later independently published by Erb et al. under the name "Recursive deconvolution"[29]
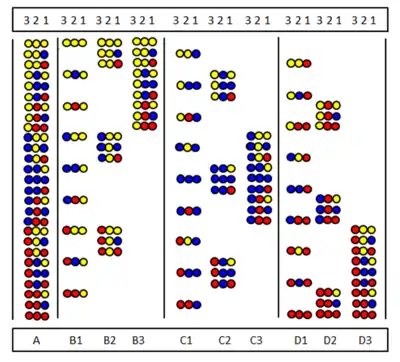
Recursive deconvolution
The method is made understandable by the figure. A 27-member peptide library is synthesized from three amino acids. After the first (A) and second (B) cycles samples were set aside before mixing them. The products of the third cycle (C) are cleaved down before mixing then are tested for activity. Suppose the group labeled by + sign is active. All members have the red amino acid at the last coupling position (CP). Consequently, the active member also has the red amino acid at the last CP. Then the red amino acid is coupled to the three samples set aside after the second cycle (B) to get samples D. After cleaving, the three E samples are formed. If after testing the sample marked by + is the active one it shows that the blue amino acid occupies the second CP in the active component. Then to the three A samples first the blue then the red amino acid is coupled (F) then tested again after cleaving (G). If the + component proves to be active, the sequence of the active component is determined and shown in H.
Positional scanning
Positional scanning was introduced independently by Furka et al.[30] and Pinilla et al.[31] The method is based on the synthesis and testing of series of sublibraries. in which a certain sequence position is occupied by the same amino acid. The figure shows the nine sublibraries (B1-D3) of a full peptide trimer library (A) made from three amino acids. In sublibraries there is a position which is occupied by the same amino acid in all components. In the synthesis of a sublibrary the support is not divided and only one amino acid is coupled to the whole sample. As a result, one position is really occupied by the same amino acid in all components. For example, in the B2 sublibrary position 2 is occupied by the "yellow" amino acid in all the nine components. If in a screening test this sublibrary gives positive answer it means that position 2 in the active peptide is also occupied by the "yellow" amino acid. The amino acid sequence can be determined by testing all the nine (or sometime less) sublibraries.
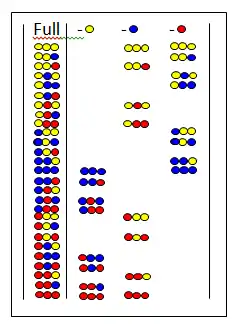
Omission libraries
In omission libraries[32][33] a certain amino acid is missing from all peptides of the mixture. The figure shows the full library and the three omission libraries. At the top the omitted amino acids are shown. If the omission library gives a negative test the omitted amino acid is present in the active component.
Deconvolution of tethered combinatorial libraries
If the peptides are not cleaved from the solid support we deal with a mixture of beads, each bead containing a single peptide. Smith and his colleagues[34] showed earlier that peptides could be tested in tethered form, too. This approach was also used in screening peptide libraries. The tethered peptide library was tested with a dissolved target protein. The beads to which the protein was attached were picked out, removed the protein from the bead then the tethered peptide was identified by sequencing. A somewhat different approach was followed by Taylor and Morken.[35] They used infrared thermography to identify catalysts in non-peptide tethered libraries. The method is based on the heat that is evolved in the beads that contain a catalyst when the tethered library immersed into a solution of a substrate. When the beads are examined through an infrared microscope the catalyst containing beads appear as bright spots and can be picked out.
Encoded combinatorial libraries
If we deal with a non-peptide organic libraries library it is not as simple to determine the identity of the content of a bead as in the case of a peptide one. In order to circumvent this difficulty methods had been developed to attach to the beads, in parallel with the synthesis of the library, molecules that encode the structure of the compound formed in the bead. Ohlmeyer and his colleagues published a binary encoding method[36] They used mixtures of 18 tagging molecules that after cleaving them from the beads could be identified by Electron Capture Gas Chromatography. Sarkar et al. described chiral oligomers of pentenoic amides (COPAs) that can be used to construct mass encoded OBOC libraries.[37] Kerr et al. introduced an innovative encoding method[38] An orthogonally protected removable bifunctional linker was attached to the beads. One end of the linker was used to attach the non-natural building blocks of the library while to the other end encoding amino acid triplets were linked. The building blocks were non-natural amino acids and the series of their encoding amino acid triplets could be determined by Edman degradation. The important aspect of this kind of encoding was the possibility to cleave down from the beads the library members together with their attached encoding tags forming a soluble library. The same approach was used by Nikolajev et al. for encoding with peptides.[39] In 1992 by Brenner and Lerner introduced DNA sequences to encode the beads of the solid support that proved to be the most successful encoding method.[40] Nielsen, Brenner and Janda also used the Kerr approach for implementing the DNA encoding[41] In the latest period of time there were important advancements in DNA sequencing. The next generation techniques make it possible to sequence large number of samples in parallel that is very important in screening of DNA encoded libraries. There was another innovation that contributed to the success of DNA encoding. In 2000 Halpin and Harbury omitted the solid support in the split-mix synthesis of the DNA encoded combinatorial libraries and replaced it by the encoding DNA oligomers. In solid phase split and pool synthesis the number of components of libraries can't exceed the number of the beads of the support. By the novel approach of the authors, this restraint was eliminated and made it possible to prepare new compounds in practically unlimited number.[42] The Danish company Nuevolution for example synthesized a DNA encoded library containing 40 trillion! components[43] The DNA encoded libraries are soluble that makes possible to apply the efficient affinity binding in screening. Some authors apply the DEL for acromim of DNA encoded combinatorial libraries others are using DECL. The latter seems better since in this name the combinatorial nature of these libraries is clearly expressed. Several types of DNA encoded combinatorial libraries had been introduced and described in the first decade of the present millennium. These libraries are very successfully applied in drug research.
- DNA templated synthesis of combinatorial libraries described in 2001 by Gartner et al.[44]
- Dual pharmacophore DNA encoded combinatorial libraries invented in 2004 by Mlecco et al.[45]
- Sequence encoded routing published by Harbury Halpin and Harbury in 2004.[46]
- Single pharmacophore DNA encoded combinatorial libraries introduced in 2008 by Manocci et al.[47]
- DNA encoded combinatorial libraries formed by using yoctoliter-scale reactor published by Hansen et al. in 2009[48]
Details are found about their synthesis and application in the page DNA-encoded chemical library. The DNA encoded soluble combinatorial libraries have drawbacks, too. First of all the advantage coming from the use of solid support is completely lost. In addition, the polyionic character of DNA encoding chains limits the utility of non-aqueous solvents in the synthesis. For this reason many laboratories choose to develop DNA compatible reactions for use in the synthesis of DECLs. Quite a few of available ones are already described[49][50][51]
Materials science
Materials science has applied the techniques of combinatorial chemistry to the discovery of new materials. This work was pioneered by P.G. Schultz et al. in the mid-nineties [52] in the context of luminescent materials obtained by co-deposition of elements on a silicon substrate. His work was preceded by J. J. Hanak in 1970[53] but the computer and robotics tools were not available for the method to spread at the time. Work has been continued by several academic groups[54][55][56][57] as well as companies with large research and development programs (Symyx Technologies, GE, Dow Chemical etc.). The technique has been used extensively for catalysis,[58] coatings,[59] electronics,[60] and many other fields.[61] The application of appropriate informatics tools is critical to handle, administer, and store the vast volumes of data produced.[62] New types of design of experiments methods have also been developed to efficiently address the large experimental spaces that can be tackled using combinatorial methods.[63]
Diversity-oriented libraries
Even though combinatorial chemistry has been an essential part of early drug discovery for more than two decades, so far only one de novo combinatorial chemistry-synthesized chemical has been approved for clinical use by FDA (sorafenib, a multikinase inhibitor indicated for advanced renal cancer).[64] The analysis of the poor success rate of the approach has been suggested to connect with the rather limited chemical space covered by products of combinatorial chemistry.[65] When comparing the properties of compounds in combinatorial chemistry libraries to those of approved drugs and natural products, Feher and Schmidt[65] noted that combinatorial chemistry libraries suffer particularly from the lack of chirality, as well as structure rigidity, both of which are widely regarded as drug-like properties. Even though natural product drug discovery has not probably been the most fashionable trend in the pharmaceutical industry in recent times, a large proportion of new chemical entities still are nature-derived compounds,[66][67][68][69][70][71] and thus, it has been suggested that effectiveness of combinatorial chemistry could be improved by enhancing the chemical diversity of screening libraries.[72] As chirality and rigidity are the two most important features distinguishing approved drugs and natural products from compounds in combinatorial chemistry libraries, these are the two issues emphasized in so-called diversity oriented libraries, i.e. compound collections that aim at coverage of the chemical space, instead of just huge numbers of compounds.[73][74][75][76][77][78]
Patent classification subclass
In the 8th edition of the International Patent Classification (IPC), which entered into force on January 1, 2006, a special subclass has been created for patent applications and patents related to inventions in the domain of combinatorial chemistry: "C40B".
See also
References
- Pottel, J.; Moitessier, N. (2017). "Customizable Generation of Synthetically Accessible, Local Chemical Subspaces". J. Chem. Inf. Model. 57 (3): 454–467. doi:10.1021/acs.jcim.6b00648. PMID 28234470.
- Furka Á. Tanulmány, gyógyászatilag hasznosítható peptidek szisztematikus felkutatásának lehetőségéről (and Study on the possibility of systematic searching for pharmaceutically useful peptides https://mersz.hu/mod/object.php?objazonosito=matud202006_f42772_i2
- Furka Á (2002). Combinatorial Chemistry 20 years on… Drug DiscovToday 7; 1-4. https://doi.org/10.1016/S1359-6446(02)00001-6
- "COMBINATORIAL CHEMISTRY: A REVIEW". International Journal of Pharmaceutical Sciences and Research. 4 (7). July 2013. doi:10.13040/IJPSR.0975-8232.4(7).2502-16. Retrieved June 21, 2020.
- Lesney, Mark S. (2002). "Concocting combinatorials: Chemistry in drug development". Retrieved October 19, 2018.
- Dolle, Roland (2011). Historical overview of chemical library design. in Zhou, J.Z. (ed.), Chemical Library Design, Methods in Molecular Biology, Springer, 2011, Chapter 1, pp. 3-25.
- Podlewska, S.; Czarnecki, W.M.; Kafel, R.; Bojarski, A.J. (2017). "Creating the New from the Old: Combinatorial Libraries Generation with Machine-Learning-Based Compound Structure Optimization". J. Chem. Inf. Modeling. 57 (2): 133–147. doi:10.1021/acs.jcim.6b00426. PMID 28158942.
- Jeffrey W. Noonan et al. "Advancing Parallel Solution Phase Library Synthesis through Efficient Purification, Quantitation, and Characterization Techniques" Journal of Laboratory Automation, 48 (1992) 3789.
- E. V.Gordeeva et al. "COMPASS program - an original semi-empirical approach to computer-assisted synthesis" Tetrahedron, 48 (1992) 3789.
- Furka Á, Sebestyén F, Asgedom M, Dibó G. Cornucopia of peptides by synthesis. In Highlights of Modern Biochemistry, Proceedings of the 14th International Congress of Biochemistry. VSP.Utrecht.1988; 5; p. 47.
- Á. Furka, F. Sebestyen, M. Asgedom, G. Dibo, General method for rapid synthesis of multicomponent peptide mixtures. Int. J. Peptide Protein Res., 1991, 37, 487-493.
- Merrifield RB, 1963 J. Am. Chem. Soc. 85, 2149.
- Scott, J.; Smith, G. (1990-07-27). "Searching for peptide ligands with an epitope library". Science. American Association for the Advancement of Science (AAAS). 249 (4967): 386–390. Bibcode:1990Sci...249..386S. doi:10.1126/science.1696028. ISSN 0036-8075. PMID 1696028.
- Cwirla, S. E.; Peters, E. A.; Barrett, R. W.; Dower, W. J. (1990-08-01). "Peptides on phage: a vast library of peptides for identifying ligands". Proceedings of the National Academy of Sciences. 87 (16): 6378–6382. Bibcode:1990PNAS...87.6378C. doi:10.1073/pnas.87.16.6378. ISSN 0027-8424. PMC 54537. PMID 2201029.
- J. J. Devlin, L. C. Panganiban and P. E. Devlin Science 1990, 249, 404.
- Fodor SP, Read JL, Pirrung MC, Stryer L, Lu AT, Solas D, 1991. Light-directed, spatially addressable parallel chemical synthesis. Science 251, 767-73.
- H. M. Geysen, R. H. Meloen, S. J. Barteling Proc. Natl. Acad. Sci. USA 1984, 81, 3998.
- E. J. Moran, S. Sarshar, J. F. Cargill, M. Shahbaz, A Lio, A. M. M. Mjalli, R. W. Armstrong J. Am. Chem. Soc. 1995, 117, 10787.
- K. C. Nicolaou, X –Y. Xiao, Z. Parandoosh, A. Senyei, M. P. Nova Angew. Chem. Int. Ed. Engl. 1995, 36, 2289.
- Á. Furka, J. W. Christensen, E. Healy, H. R. Tanner, H. Saneii J. Comb. Chem. 2000, 2, 220.
- Booth, R. John; Hodges, John C. (1999–2001). "Solid-Supported Reagent Strategies for Rapid Purification of Combinatorial Synthesis Products". Accounts of Chemical Research. 32 (1): 18–26. doi:10.1021/ar970311n. ISSN 0001-4842.
- Armstrong, Robert W.; Combs, Andrew P.; Tempest, Paul A.; Brown, S. David; Keating, Thomas A. (1996–2001). "Multiple-Component Condensation Strategies for Combinatorial Library Synthesis". Accounts of Chemical Research. 29 (3): 123–131. doi:10.1021/ar9502083. ISSN 0001-4842. S2CID 95815562.
- Still, W. Clark (1996–2001). "Discovery of Sequence-Selective Peptide Binding by Synthetic Receptors Using Encoded Combinatorial Libraries". Accounts of Chemical Research. 29 (3): 155–163. doi:10.1021/ar950166i. ISSN 0001-4842.
- DeWitt, Sheila Hobbs; Czarnik, Anthony W. (1996–2001). "Combinatorial Organic Synthesis Using Parke-Davis's DIVERSOMER Method". Accounts of Chemical Research. 29 (3): 114–122. doi:10.1021/ar950209v. ISSN 0001-4842.
- Schwabacher, Alan W.; Shen, Yixing; Johnson, Christopher W. (1999–2009). "Fourier Transform Combinatorial Chemistry". Journal of the American Chemical Society. 121 (37): 8669–8670. doi:10.1021/ja991452i. ISSN 0002-7863.
- Ellman, Jonathan A. (1996–2001). "Design, Synthesis, and Evaluation of Small-Molecule Libraries". Accounts of Chemical Research. 29 (3): 132–143. doi:10.1021/ar950190w. ISSN 0001-4842.
- Gordon, E. M.; Gallop, M. A.; Patel, D. V. (1996–2001). "Strategy and Tactics in Combinatorial Organic Synthesis. Applications to Drug Discovery". Accounts of Chemical Research. 29 (3): 144–154. doi:10.1021/ar950170u. ISSN 0001-4842.
- A. Furka Sub-Library Composition of Peptide Libraries. Potential Application in Screening. Drug Development Research 33, 90-97 (1994).
- Erb E, Janda KD, Brenner S (1994) Recursive deconvolution of combinatorial chemical libraries Proc. Natl Acad Sci.USA 91; 11422-11426.
- Furka Á, Sebestyén F, WC 93/24517, 1993.
- Pinilla C, Appel JR, Blanc P, Houghten RA (1993) Rapid identification of high affinity peptide ligands using positional scanning synthetic peptide combinatorial libraries. BioTechniques 13(6); 901-5.
- Carell TE, Winter A, Rebek J Jr. (1994) A Novel Procedure for the Synthesis of Libraries Containing Small Organic Molecules, Angew Chem Int Ed Engl 33; 2059-2061.
- Câmpian E, Peterson M, Saneii HH, Furka Á, (1998) Deconvolution by omission libraries, Bioorg &[ Med Chem Letters 8; 2357-2362.
- J. A. Smith J. G. R. Hurrel, S. J. Leach A novel method for delineating antigenic determinants: peptide synthesis and radioimmunoassay using the same solid support. Immunochemistry 1977, 14, 565.
- S. J. Taylor, J. P. Morken Thermographic Selection of Effective Catalysts from an Encoded Polymer-Bound Library Science 1998, 280, 267.
- Ohlmeyer MHJ, Swanson RN, Dillard LW, Reader JC, Asouline G, Kobayashi R, Wigler M, Still WC (1993) Complex synthetic chemical libraries indexed with molecular tags, Proc Natl Acad Sci USA 90; 10922-10926.
- Sarkar M, Pascal BD, Steckler C, Aquino C., Micalizio GC, Kodadek T, Chalmers MJ (1993) Decoding Split and Pool Combinatorial Libraries with Electron Transfer Dissociation Tandem Mass Spectrometry, J Am Soc Mass Spectrom 24(7): 1026-36.
- Kerr JM, Banville SC, Zuckermann RN (1993) Encoded Combinatorial Peptide Libraries Containing Non-Natural Amino Acids, J Am Chem. Soc 115; 2529-2531.
- Nikolaiev V, Stierandová A, Krchnák V, Seligmann B, Lam KS, Salmon SE, Lebl M, (1993) Peptide-encoding for structure determination of nonsequenceable polymers within libraries synthesized and tested on solid-phase supports, Pept Res. 6(3):161-70.
- Brenner S, Lerner RA. (1992) Encoded combinatorial chemistry. Proc Natl Acad Sci USA 89; 5381–5383.
- Nielsen J, Brenner S, Janda KD. (1993) Synthetic methods for the implementation of encoded combinatorial chemistry. Journal of the American Chemical Society, 115 (21); 9812–9813.
- Harbury DR, Halpin DR (2000) WO 00/23458.
- B. Halford How DNA-encoded libraries are revolutionizing drug discovery. C&EN 2017, 95, Issue 25.
- Gartner ZJ, Tse BN, Grubina RB, Doyon JB, Snyder TM, Liu DR (2004) DNA-Templated Organic Synthesis and Selection of a Library of Macrocycles, Science 305; 1601-1605.
- Melkko S, Scheuermann J, Dumelin CE, Neri D (2004) Encoded self-assembling chemical libraries Nat Biotechnol 22; 568-574.
- Halpin DR, Harbury PB (2004) DNA Display I. Sequence-Encoded Routing of DNA Populations, PLoS Biology 2; 1015-102.
- Mannocci L, Zhang Y, Scheuermann J, Leimbacher M, De Bellis G, Rizzi E, Dumelin C, Melkko S, and Neri N (2008) High-throughput sequencing allows the identification of binding molecules isolated from DNA-encoded chemical libraries, Proc Natl Acad Sci USA 105;17670–17675.
- Hansen MH, Blakskjær P, Petersen LK, Hansen TH, Højfeldt JW, Gothelf KV, HansenNJV (2009) A Yoctoliter-Scale DNA Reactor for Small-Molecule Evolution (2009) J Am Chem Soc 131; 1322-1327.
- Luk KC, Satz AL (2014) DNA‐Compatible Chemistry in: Goodnow Jr. RA Editor A Handbook for DNA‐Encoded Chemistry: Theory and Applications for Exploring Chemical Space and Drug Discovery, Wiley, pp 67-98.
- Satz AL, Cai J, Chen Y,§, Goodnow R, Felix Gruber F, Kowalczyk A, Petersen A, Naderi-Oboodi G, Orzechowski L, Strebel Q (2015) DNA Compatible Multistep Synthesis and Applications to DNA Encoded Libraries Bioconjugate Chem 26; 1623−1632.
- Li Y, Gabriele E, Samain F, Favalli N, Sladojevich F, Scheuermann J, Neri D (2016) Optimized reaction conditions for amide bond formation in DNA-encoded combinatorial libraries, ACS Comb Sci 18(8); 438–443.
- X. -D. Xiang et al. "A Combinatorial Approach to Materials Discovery" Science 268 (1995) 1738
- J.J. Hanak, J. Mater. Sci, 1970, 5, 964-971
- Combinatorial methods for development of sensing materials, Springer, 2009. ISBN 978-0-387-73712-6
- V. M. Mirsky, V. Kulikov, Q. Hao, O. S. Wolfbeis. Multiparameter High Throughput Characterization of Combinatorial Chemical Microarrays of Chemosensitive Polymers. Macromolec. Rap. Comm., 2004, 25, 253-258
- H. Koinuma et al. "Combinatorial solid state materials science and technology" Sci. Technol. Adv. Mater. 1 (2000) 1 free download
- Andrei Ionut Mardare et al. "Combinatorial solid state materials science and technology" Sci. Technol. Adv. Mater. 9 (2008) 035009 free download
- Applied Catalysis A, Volume 254, Issue 1, Pages 1-170 (10 November 2003)
- J. N. Cawse et al., Progress in Organic Coatings, Volume 47, Issue 2, August 2003, Pages 128-135
- Combinatorial Methods for High-Throughput Materials Science, MRS Proceedings Volume 1024E, Fall 2007
- Combinatorial and Artificial Intelligence Methods in Materials Science II, MRS Proceedings Volume 804, Fall 2004
- QSAR and Combinatorial Science, 24, Number 1 (February 2005)
- J. N. Cawse, Ed., Experimental Design for Combinatorial and High Throughput Materials Development, John Wiley and Sons, 2002.
- D. Newman and G. Cragg "Natural Products as Sources of New Drugs over the Last 25 Years" J Nat Prod 70 (2007) 461
- M. Feher and J. M. Schmidt "Property Distributions: Differences between Drugs, Natural Products, and Molecules from Combinatorial Chemistry" J. Chem. Inf. Comput. Sci., 43 (2003) 218
- Cordier, C.; Morton, D.; Murrison, S.; Nelson, A.; O'Leary-Steele, C. (2008). "Natural products as an inspiration in the diversity-oriented synthesis of bioactive compound libraries". Nat. Prod. Rep. 25 (4): 719–737. doi:10.1039/B706296F. PMC 2496956. PMID 18663392. S2CID 15697405.
- de Sousa Luis, J.A.; Costa Barros, R.P.; de Sousa, N.F.; Muratov, E.; Scotti, L.; Scotti, M.T. (2021). "Virtual screening of natural products database". Mini-Rev. Med. Chem. 21 (18): 2657–2730. doi:10.2174/1389557520666200730161549. PMID 32744975. S2CID 220965407.
- Harvey, A.L. (2008). "Natural products in drug discovery". Drug Discov. Today. 13 (19–20): 894–901. doi:10.1016/j.drudis.2008.07.004. PMID 18691670.
- Kohli, S. (2018). Integrated approach to nature as source of new drug lead, in Molecular Insight of Drug Design, Ed. Aditya, A. (PDF). IntechOpen. pp. 29–44. doi:10.5772/intechopen.74961. S2CID 64364163. Archived (PDF) from the original on 2022-05-27.
- Kaiser, M.; Wetzel, S.; Kumar, K.; Waldmann, H. (2008). "Biology-inspired synthesis of compound libraries". Cell. Mol. Life Sci. 65 (7–8): 1186–1201. doi:10.1007/s00018-007-7492-1. PMID 18193390. S2CID 22601514.
- Teague, S.J.; Davis, A.M.; Leeson, P.D.; Oprea, T. (1999). "The design of leadlike combinatorial libraries". Angew. Chem. Int. Ed. 38 (24): 3743–3748. doi:10.1002/(SICI)1521-3773(19991216)38:24<3743::AID-ANIE3743>3.0.CO;2-U. PMID 10649345.
- Su QB, Beeler AB, Lobkovsky E, Porco JA, Panek JS "Stereochemical diversity through cyclodimerization: Synthesis of polyketide-like macrodiolides." Org Lett 2003, 5:2149-2152.
- Medina-Franco, J.L.; Martínez-Mayorga, K.; Giulianotti, M.A.; Houghten, R.A.; Pinilla, C. (2008). "Visualization of the chemical space in drug discovery". Curr Comput-Aided Drug Des. 4 (4): 322–333. doi:10.2174/157340908786786010.
- Rosén, J.; Gottfries, J.; Muresan, S.; Backlund, A.; Oprea, T.I. (2009). "Novel Chemical Space Exploration via Natural Products". J. Med. Chem. 52 (7): 1953–1962. doi:10.1021/jm801514w. PMC 2696019. PMID 19265440.
- Lachance, H.; Wetzel, S.; Kumar, K.; Waldmann, H. (2012). "Charting, Navigating, and Populating Natural Product Chemical Space for Drug Discovery". J. Med. Chem. 55 (13): 5989–6001. doi:10.1021/jm300288g. PMID 22537178.
- Reymond, J.-L.; Awale, M. (2012). "Exploring Chemical Space for Drug Discovery Using the Chemical Universe Database". ACS Chem. Neurosci. 3 (9): 649–657. doi:10.1021/cn3000422. PMC 3447393. PMID 23019491.
- Medina-Franco, J.L. (2013). Chemoinformatic characterization of the chemical space and molecular diversity of compound libraries, in Diversity-Oriented Synthesis: Basics and Applications in Organic Synthesis, Drug Discovery, and Chemical Biology, Trabocchi, A.; Ed., Chapter 10, 2013, 325-352. John Wiley & Sons, Inc.
- Klein, R.; Lindell, S.D. (2014). Combinatorial Chemistry Library Design, in, Plant Chemical Biology, Audenaert, D.; Overvoorde, P.; Eds. New Jersey: John Wiley & Sons, Inc. pp. 40–63.
External links
- English version of the 1982 document
- "The concealed side of the history of combinatorial chemistry"
- IUPAC's "Glossary of Terms Used in Combinatorial Chemistry"
- ACS Combinatorial Science (formerly Journal of Combinatorial Chemistry)
- Combinatorial Chemistry Review
- Molecular Diversity
- Combinatorial Chemistry and High Throughput Screening
- Combinatorial Chemistry: an Online Journal
- SmiLib - A free open-source software for combinatorial library enumeration
- GLARE - A free open-source software for combinatorial library design