Computer-aided diagnosis
Computer-aided detection (CADe), also called computer-aided diagnosis (CADx), are systems that assist doctors in the interpretation of medical images. Imaging techniques in X-ray, MRI, Endoscopy, and ultrasound diagnostics yield a great deal of information that the radiologist or other medical professional has to analyze and evaluate comprehensively in a short time. CAD systems process digital images or videos for typical appearances and to highlight conspicuous sections, such as possible diseases, in order to offer input to support a decision taken by the professional.
| Computer-aided diagnosis | |
|---|---|
 | |
| Purpose | computer assistance diagnosis of images |
CAD also has potential future applications in digital pathology with the advent of whole-slide imaging and machine learning algorithms. So far its application has been limited to quantifying immunostaining but is also being investigated for the standard H&E stain.[1]
CAD is an interdisciplinary technology combining elements of artificial intelligence and computer vision with radiological and pathology image processing. A typical application is the detection of a tumor. For instance, some hospitals use CAD to support preventive medical check-ups in mammography (diagnosis of breast cancer), the detection of polyps in Colonoscopy, and lung cancer.
Computer-aided detection (CADe) systems are usually confined to marking conspicuous structures and sections. Computer-aided diagnosis (CADx) systems evaluate the conspicuous structures. For example, in mammography CAD highlights microcalcification clusters and hyperdense structures in the soft tissue. This allows the radiologist to draw conclusions about the condition of the pathology. Another application is CADq, which quantifies, e.g., the size of a tumor or the tumor's behavior in contrast medium uptake. Computer-aided simple triage (CAST) is another type of CAD, which performs a fully automatic initial interpretation and triage of studies into some meaningful categories (e.g. negative and positive). CAST is particularly applicable in emergency diagnostic imaging, where a prompt diagnosis of critical, life-threatening condition is required.
Although CAD has been used in clinical environments for over 40 years, CAD usually does not substitute the doctor or other professional, but rather plays a supporting role. The professional (generally a radiologist) is generally responsible for the final interpretation of a medical image.[2] However, the goal of some CAD systems is to detect earliest signs of abnormality in patients that human professionals cannot, as in diabetic retinopathy, architectural distortion in mammograms,[3][4] ground-glass nodules in thoracic CT,[5][6] and non-polypoid (“flat”) lesions in CT colonography.[7]
Topics
A Brief History
In the late 1950s, with the dawn of modern computers researchers in various fields started exploring the possibility of building computer-aided medical diagnostic (CAD) systems.[8] These first CAD systems used flow-charts, statistical pattern-matching, probability theory or knowledge bases to drive their decision-making process.[9]
Since the early 1970s, some of the very early CAD systems in medicine, which were often referred as “expert systems” in medicine, were developed and used mainly for educational purposes. The MYCIN expert system,[10] the Internist-I expert system[11] and the CADUCEUS (expert system)[12] are some of such examples.
During the beginning of the early developments, the researchers were aiming at building entirely automated CAD / expert systems. The expectation of what computers can do was unrealistically optimistic among these scientists. However, after the breakthrough paper, “Reducibility among Combinatorial Problems” by Richard M. Karp,[13] it became clear that there were limitations but also potential opportunities when one develops algorithms to solve groups of important computational problems.[9]
As result of the new understanding of the various algorithmic limitations that Karp discovered in the early 1970s, researchers started realizing the serious limitations that CAD and expert systems in medicine have.[9] The recognition of these limitations brought the investigators to develop new kinds of CAD systems by using advanced approaches. Thus, by the late 1980s and early 1990s the focus sifted in the use of data mining approaches for the purpose of using more advanced and flexible CAD systems.
In 1998, the first commercial CAD system for mammography, the ImageChecker system, was approved by the US Food and Drug Administration (FDA). In the following years several commercial CAD systems for analyzing mammography, breast MRI, medical imagining of lung, colon, and heart also received FDA approvals. Currently, CAD systems are used as a diagnostic aid to provide physicians for better medical decision-making.[14]
Methodology
CAD is fundamentally based on highly complex pattern recognition. X-ray or other types of images are scanned for suspicious structures. Normally a few thousand images are required to optimize the algorithm. Digital image data are copied to a CAD server in a DICOM-format and are prepared and analyzed in several steps.
1. Preprocessing for
- Reduction of artifacts (bugs in images)
- Image noise reduction
- Leveling (harmonization) of image quality (increased contrast) for clearing the image's different basic conditions e.g. different exposure parameter.
- Filtering
2. Segmentation for
- Differentiation of different structures in the image, e.g. heart, lung, ribcage, blood vessels, possible round lesions
- Matching with anatomic databank
- Sample gray-values in volume of interest[15]
3. Structure/ROI (Region of Interest) Analyze Every detected region is analyzed individually for special characteristics:
- Compactness
- Form, size and location
- Reference to close by structures / ROIs
- Average grey level value analyze within a ROI
- Proportion of grey levels to border of the structure inside the ROI
4. Evaluation / classification After the structure is analyzed, every ROI is evaluated individually (scoring) for the probability of a TP. The following procedures are examples of classification algorithms.
- Nearest-Neighbor Rule (e.g. k-nearest neighbors)[16]
- Minimum distance classifier
- Cascade classifier
- Naive Bayes classifier
- Artificial neural network[17][18][19][20][21]
- Radial basis function network (RBF)
- Support vector machine (SVM)[22][23]
- Principal component analysis (PCA)
If the detected structures have reached a certain threshold level, they are highlighted in the image for the radiologist. Depending on the CAD system these markings can be permanently or temporary saved. The latter's advantage is that only the markings which are approved by the radiologist are saved. False hits should not be saved, because an examination at a later date becomes more difficult then.
Sensitivity and specificity
CAD systems seek to highlight suspicious structures. Today's CAD systems cannot detect 100% of pathological changes. The hit rate (sensitivity) can be up to 90% depending on system and application.[24] A correct hit is termed a True Positive (TP), while the incorrect marking of healthy sections constitutes a False Positive (FP). The less FPs indicated, the higher the specificity is. A low specificity reduces the acceptance of the CAD system because the user has to identify all of these wrong hits. The FP-rate in lung overview examinations (CAD Chest) could be reduced to 2 per examination. In other segments (e.g. CT lung examinations) the FP-rate could be 25 or more. In CAST systems the FP rate must be extremely low (less than 1 per examination) to allow a meaningful study triage.
Absolute detection rate
The absolute detection rate of the radiologist is an alternative metric to sensitivity and specificity. Overall, results of clinical trials about sensitivity, specificity, and the absolute detection rate can vary markedly. Each study result depends on its basic conditions and has to be evaluated on those terms. The following facts have a strong influence:
- Retrospective or prospective design
- Quality of the used images
- Condition of the x-ray examination
- Radiologist's experience and education
- Type of lesion
- Size of the considered lesion
Challenges that CAD in Medicine Faces Today
Despite the many developments that CAD has achieved since the dawn of computers, there are still certain challenges that CAD systems face today.[25]
Some challenges are related to various algorithmic limitations in the procedures of a CAD system including input data collection, preprocessing, processing and system assessments. Algorithms are generally designed to select a single likely diagnosis, thus providing suboptimal results for patients with multiple, concurrent disorders.[26] Today input data for CAD mostly come from electronic health records (EHR). Effective designing, implementing and analyzing for EHR is a major necessity on any CAD systems.[25]
Due to the massive availability of data and the need to analyze such data, big data is also one of the biggest challenges that CAD systems face today. The increasingly vast amount of patient data is a serious problem. Often the patient data are complex and can be semi-structured or unstructured data. It requires highly developed approaches to store, retrieve and analyze them in reasonable time.[25]
During the preprocessing stage, input data requires to be normalized. The normalization of input data includes noise reduction, and filtering. Processing may contain a few sub-steps depending on applications. Basic three sub-steps on medical imaging are segmentation, feature extraction / selection and classification. These sub-steps require advanced techniques to analyze input data with less computational time. Although much effort has been devoted on creating innovative techniques for these procedures of CAD systems, there is still not the single best algorithm for each step. Ongoing studies in building innovative algorithms for all the aspects of CAD systems is essential.[25]
There is also a lack of standardized assessment measures for CAD Systems.[25] This fact may cause the difficulty for obtaining FDA approval for commercial use. Moreover, while many positive developments of CAD systems have been proven, studies for validating their algorithms for clinical practice has hardly been confirmed.[27]
Other challenges are related to the problem for healthcare providers to adopt new CAD systems in clinical practice. Some negative studies may discourage the use of CAD. In addition, the lack of training of health professionals on the use of CAD sometimes brings the incorrect interpretation of the system outcomes. These challenges are described in more detail in.[25]
Applications
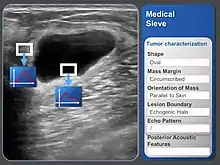
CAD is used in the diagnosis of breast cancer, lung cancer, colon cancer, prostate cancer, bone metastases, coronary artery disease, congenital heart defect, pathological brain detection, fracture detection, Alzheimer's disease, and diabetic retinopathy.
Breast cancer
CAD is used in screening mammography (X-ray examination of the female breast). Screening mammography is used for the early detection of breast cancer. CAD systems are often utilized to help classify a tumor as malignant or benign. CAD is especially established in US and the Netherlands and is used in addition to human evaluation, usually by a radiologist. The first CAD system for mammography was developed in a research project at the University of Chicago. Today it is commercially offered by iCAD and Hologic. However, while achieving high sensitivities, CAD systems tend to have very low specificity and the benefits of using CAD remain uncertain. A 2008 systematic review on computer-aided detection in screening mammography concluded that CAD does not have a significant effect on cancer detection rate, but does undesirably increase recall rate (i.e. the rate of false positives). However, it noted considerable heterogeneity in the impact on recall rate across studies.[28]
Recent advances in machine learning, deep-learning and artificial intelligence technology have enabled the development of CAD systems that are clinically proven to assist radiologists in addressing the challenges of reading mammographic images by improving cancer detection rates and reducing false positives and unnecessary patient recalls, while significantly decreasing reading times.[29]
Procedures to evaluate mammography based on magnetic resonance imaging exist too.
Lung cancer (bronchial carcinoma)
In the diagnosis of lung cancer, computed tomography with special three-dimensional CAD systems are established and considered as appropriate second opinions.[30] At this a volumetric dataset with up to 3,000 single images is prepared and analyzed. Round lesions (lung cancer, metastases and benign changes) from 1 mm are detectable. Today all well-known vendors of medical systems offer corresponding solutions.
Early detection of lung cancer is valuable. However, the random detection of lung cancer in the early stage (stage 1) in the X-ray image is difficult. Round lesions that vary from 5–10 mm are easily overlooked.[31] The routine application of CAD Chest Systems may help to detect small changes without initial suspicion. A number of researchers developed CAD systems for detection of lung nodules (round lesions less than 30 mm) in chest radiography[32][33][34] and CT,[35][36] and CAD systems for diagnosis (e.g., distinction between malignant and benign) of lung nodules in CT. Virtual dual-energy imaging[37][38][39][40] improved the performance of CAD systems in chest radiography.[41]
Colon cancer
CAD is available for detection of colorectal polyps in the colon in CT colonography.[42][43] Polyps are small growths that arise from the inner lining of the colon. CAD detects the polyps by identifying their characteristic "bump-like" shape. To avoid excessive false positives, CAD ignores the normal colon wall, including the haustral folds.
Cardiovascular disease
State-of-the-art methods in cardiovascular computing, cardiovascular informatics, and mathematical and computational modeling can provide valuable tools in clinical decision-making.[44] CAD systems with novel image-analysis-based markers as input can aid vascular physicians to decide with higher confidence on best suitable treatment for cardiovascular disease patients.
Reliable early-detection and risk-stratification of carotid atherosclerosis is of outmost importance for predicting strokes in asymptomatic patients.[45] To this end, various noninvasive and low-cost markers have been proposed, using ultrasound-image-based features.[46] These combine echogenicity, texture, and motion[47][48][49][50] characteristics to assist clinical decision towards improved prediction, assessment and management of cardiovascular risk.[51]
CAD is available for the automatic detection of significant (causing more than 50% stenosis) coronary artery disease in coronary CT angiography (CCTA) studies.[52]
Congenital heart defect
Early detection of pathology can be the difference between life and death. CADe can be done by auscultation with a digital stethoscope and specialized software, also known as Computer-aided auscultation. Murmurs, irregular heart sounds, caused by blood flowing through a defective heart, can be detected with high sensitivity and specificity. Computer-aided auscultation is sensitive to external noise and bodily sounds and requires an almost silent environment to function accurately.
Pathological brain detection (PBD)
Chaplot et al. was the first to use Discrete Wavelet Transform (DWT) coefficients to detect pathological brains.[53] Maitra and Chatterjee employed the Slantlet transform, which is an improved version of DWT. Their feature vector of each image is created by considering the magnitudes of Slantlet transform outputs corresponding to six spatial positions chosen according to a specific logic.[54]
In 2010, Wang and Wu presented a forward neural network (FNN) based method to classify a given MR brain image as normal or abnormal. The parameters of FNN were optimized via adaptive chaotic particle swarm optimization (ACPSO). Results over 160 images showed that the classification accuracy was 98.75%.[55]
In 2011, Wu and Wang proposed using DWT for feature extraction, PCA for feature reduction, and FNN with scaled chaotic artificial bee colony (SCABC) as classifier.[56]
In 2013, Saritha et al. were the first to apply wavelet entropy (WE) to detect pathological brains. Saritha also suggested to use spider-web plots.[57] Later, Zhang et al. proved removing spider-web plots did not influence the performance.[58] Genetic pattern search method was applied to identify abnormal brain from normal controls. Its classification accuracy was reported as 95.188%.[59] Das et al. proposed to use Ripplet transform.[60] Zhang et al. proposed to use particle swarm optimization (PSO).[61] Kalbkhani et al. suggested to use GARCH model.[62]
In 2014, El-Dahshan et al. suggested to use pulse coupled neural network.[63]
In 2015, Zhou et al. suggested to apply naive Bayes classifier to detect pathological brains.[64]
Alzheimer's disease
CADs can be used to identify subjects with Alzheimer's and mild cognitive impairment from normal elder controls.
In 2014, Padma et al. used combined wavelet statistical texture features to segment and classify AD benign and malignant tumor slices.[57] Zhang et al. found kernel support vector machine decision tree had 80% classification accuracy, with an average computation time of 0.022s for each image classification.[65]
In 2019, Signaevsky et al. have first reported a trained Fully Convolutional Network (FCN) for detection and quantification of neurofibrillary tangles (NFT) in Alzheimer's disease and an array of other tauopathies. The trained FCN achieved high precision and recall in naive digital whole slide image (WSI) semantic segmentation, correctly identifying NFT objects using a SegNet model trained for 200 epochs. The FCN reached near-practical efficiency with average processing time of 45 min per WSI per Graphic Processing Unit (GPU), enabling reliable and reproducible large-scale detection of NFTs. The measured performance on test data of eight naive WSI across various tauopathies resulted in the recall, precision, and an F1 score of 0.92, 0.72, and 0.81, respectively.[66]
Eigenbrain is a novel brain feature that can help to detect AD, based on Principal Component Analysis[67] or Independent Component Analysis decomposition.[68] Polynomial kernel SVM has been shown to achieve good accuracy. The polynomial KSVM performs better than linear SVM and RBF kernel SVM.[69] Other approaches with decent results involve the use of texture analysis,[70] morphological features,[71] or high-order statistical features[72]
Nuclear medicine
CADx is available for nuclear medicine images. Commercial CADx systems for the diagnosis of bone metastases in whole-body bone scans and coronary artery disease in myocardial perfusion images exist.[73]
With a high sensitivity and an acceptable false lesions detection rate, computer-aided automatic lesion detection system is demonstrated as useful and will probably in the future be able to help nuclear medicine physicians to identify possible bone lesions.[74]
Diabetic retinopathy
Diabetic retinopathy is a disease of the retina that is diagnosed predominantly by fundoscopic images. Diabetic patients in industrialised countries generally undergo regular screening for the condition. Imaging is used to recognize early signs of abnormal retinal blood vessels. Manual analysis of these images can be time-consuming and unreliable.[75][76] CAD has been employed to enhance the accuracy, sensitivity, and specificity of automated detection method. The use of some CAD systems to replace human graders can be safe and cost effective.[76]
Image pre-processing, and feature extraction and classification are two main stages of these CAD algorithms.[77]
Pre-processing methods
Image normalization is minimizing the variation across the entire image. Intensity variations in areas between periphery and central macular region of the eye have been reported to cause inaccuracy of vessel segmentation.[78] Based on the 2014 review, this technique was the most frequently used and appeared in 11 out of 40 recently (since 2011) published primary research.[77]
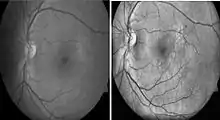
Histogram equalization is useful in enhancing contrast within an image. This technique is used to increase local contrast. At the end of the processing, areas that were dark in the input image would be brightened, greatly enhancing the contrast among the features present in the area. On the other hand, brighter areas in the input image would remain bright or be reduced in brightness to equalize with the other areas in the image. Besides vessel segmentation, other features related to diabetic retinopathy can be further separated by using this pre-processing technique. Microaneurysm and hemorrhages are red lesions, whereas exudates are yellow spots. Increasing contrast between these two groups allow better visualization of lesions on images. With this technique, 2014 review found that 10 out of the 14 recently (since 2011) published primary research.[77]
Green channel filtering is another technique that is useful in differentiating lesions rather than vessels. This method is important because it provides the maximal contrast between diabetic retinopathy-related lesions.[80] Microaneurysms and hemorrhages are red lesions that appear dark after application of green channel filtering. In contrast, exudates, which appear yellow in normal image, are transformed into bright white spots after green filtering. This technique is mostly used according to the 2014 review, with appearance in 27 out of 40 published articles in the past three years.[77] In addition, green channel filtering can be used to detect center of optic disc in conjunction with double-windowing system.
Non-uniform illumination correction is a technique that adjusts for non-uniform illumination in fundoscopic image. Non-uniform illumination can be a potential error in automated detection of diabetic retinopathy because of changes in statistical characteristics of image.[77] These changes can affect latter processing such as feature extraction and are not observable by humans. Correction of non-uniform illumination (f') can be achieved by modifying the pixel intensity using known original pixel intensity (f), and average intensities of local (λ) and desired pixels (μ) (see formula below).[81] Walter-Klein transformation is then applied to achieve the uniform illumination.[81] This technique is the least used pre-processing method in the review from 2014.
Morphological operations is the second least used pre-processing method in 2014 review.[77] The main objective of this method is to provide contrast enhancement, especially darker regions compared to background.
Feature extractions and classifications
After pre-processing of funduscopic image, the image will be further analyzed using different computational methods. However, the current literature agreed that some methods are used more often than others during vessel segmentation analyses. These methods are SVM, multi-scale, vessel-tracking, region growing approach, and model-based approaches.
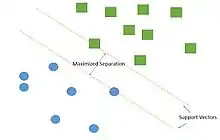
Support vector machine is by far the most frequently used classifier in vessel segmentation, up to 90% of cases. SVM is a supervised learning model that belongs to the broader category of pattern recognition technique. The algorithm works by creating a largest gap between distinct samples in the data. The goal is to create the largest gap between these components that minimize the potential error in classification.[82] In order to successfully segregate blood vessel information from the rest of the eye image, SVM algorithm creates support vectors that separate the blood vessel pixel from the rest of the image through a supervised environment. Detecting blood vessel from new images can be done through similar manner using support vectors. Combination with other pre-processing technique, such as green channel filtering, greatly improves the accuracy of detection of blood vessel abnormalities.[77] Some beneficial properties of SVM include[82]
- Flexibility – Highly flexible in terms of function
- Simplicity – Simple, especially with large datasets (only support vectors are needed to create separation between data)
Multi-scale approach is a multiple resolution approach in vessel segmentation. At low resolution, large-diameter vessels can first be extracted. By increasing resolution, smaller branches from the large vessels can be easily recognized. Therefore, one advantage of using this technique is the increased analytical speed.[75] Additionally, this approach can be used with 3D images. The surface representation is a surface normal to the curvature of the vessels, allowing the detection of abnormalities on vessel surface.
Vessel tracking is the ability of the algorithm to detect "centerline" of vessels. These centerlines are maximal peak of vessel curvature. Centers of vessels can be found using directional information that is provided by Gaussian filter. Similar approaches that utilize the concept of centerline are the skeleton-based and differential geometry-based.[75]
Region growing approach is a method of detecting neighboring pixels with similarities. A seed point is required for such method to start. Two elements are needed for this technique to work: similarity and spatial proximity. A neighboring pixel to the seed pixel with similar intensity is likely to be the same type and will be added to the growing region. One disadvantage of this technique is that it requires manual selection of seed point, which introduces bias and inconsistency in the algorithm.[75] This technique is also being used in optic disc identification.
Model-based approaches employ representation to extract vessels from images. Three broad categories of model-based are known: deformable, parametric, and template matching.[75] Deformable methods uses objects that will be deformed to fit the contours of the objects on the image. Parametric uses geometric parameters such as tubular, cylinder, or ellipsoid representation of blood vessels. Classical snake contour in combination with blood vessel topological information can also be used as a model-based approach.[83] Lastly, template matching is the usage of a template, fitted by stochastic deformation process using Hidden Markov Mode 1.
Effects on employment
Automation of medical diagnosis labor (for example, quantifying red blood cells) has some historical precedent.[84] The deep learning revolution of the 2010s has already produced AIs that are more accurate in many areas of visual diagnosis than radiologists and dermatologists, and this gap is expected to grow. Some experts, including many doctors, are dismissive of the effects that AI will have on medical specialties. In contrast, many economists and artificial intelligence experts believe that fields such as radiology will be massively disrupted, with unemployment or downward pressure on the wages of radiologists; hospitals will need fewer radiologists overall, and many of the radiologists who still exist will require substantial retraining. Geoffrey Hinton, the "Godfather of deep learning", argues that (in view of the likely advances expected in the next five or ten years) hospitals should immediately stop training radiologists, as their time-consuming and expensive training on visual diagnosis will soon be mostly obsolete, leading to a glut of traditional radiologists.[85][86] An op-ed in JAMA argues that pathologists and radiologists should merge into a single "information specialist" role, and state that "To avoid being replaced by computers, radiologists must allow themselves to be displaced by computers." Information specialists would be trained in "Bayesian logic, statistics, data science", and some genomics and biometrics; manual visual pattern recognition would be greatly de-emphasized compared with current onerous radiology training.[87]
References
- "Computer-aided Diagnosis: The Tipping Point for Digital Pathology". Digital Pathology Association. 27 April 2017.
- Oakden-Rayner, Luke (May 2019). "The Rebirth of CAD: How Is Modern AI Different from the CAD We Know?". Radiology: Artificial Intelligence. 1 (3): e180089. doi:10.1148/ryai.2019180089. ISSN 2638-6100. PMC 8017402. PMID 33937793.
- Bird, R. E.; Wallace, T. W.; Yankaskas, B. C. (1992). "Analysis of cancers missed at screening mammography". Radiology. 184 (3): 613–617. doi:10.1148/radiology.184.3.1509041. PMID 1509041.
- Baker, J. A.; Rosen, E. L.; Lo, J. Y.; et al. (2003). "Computer-Aided Detection (CAD) in Screening Mammography: Sensitivity of Commercial CAD Systems for Detecting Architectural Distortion". American Journal of Roentgenology. 181 (4): 1083–1088. doi:10.2214/ajr.181.4.1811083. PMID 14500236.
- Jang, H. J.; Lee, K. S.; Kwon, O. J.; et al. (1996). "Bronchioloalveolar carcinoma: focal area of ground-glass attenuation at thin-section CT as an early sign". Radiology. 199 (2): 485–488. doi:10.1148/radiology.199.2.8668800. PMID 8668800.
- Suzuki, K.; Li, F.; Sone, S.; Doi, K. (2005). "Computer-aided diagnostic scheme for distinction between benign and malignant nodules in thoracic low-dose CT by use of massive training artificial neural network". IEEE Transactions on Medical Imaging. 24 (9): 1138–1150. doi:10.1109/tmi.2005.852048. PMID 16156352. S2CID 2690415.
- Lostumbo, A.; Suzuki, K.; Dachman, A. H. (2010). "Flat lesions in CT colonography". Abdom Imaging. 35 (5): 578–583. doi:10.1007/s00261-009-9562-3. PMID 19633882. S2CID 13487349.
- Lea, Andrew S. (2023). Digitizing Diagnosis: Medicine, Minds, and Machines in Twentieth-Century America. Johns Hopkins University Press. pp. 1–256. ISBN 978-1421446813.
- Yanase J, Triantaphyllou E (2019). "A Systematic Survey of Computer-Aided Diagnosis in Medicine: Past and Present Developments". Expert Systems with Applications. 138: 112821. doi:10.1016/j.eswa.2019.112821. S2CID 199019309.
- Shortliffe EH, and Buchanan BG (1975). "A model of inexact reasoning in medicine". Mathematical Biosciences. 23 (3–4): 351–379. doi:10.1016/0025-5564(75)90047-4. S2CID 118063112.
- Miller RA, Pople Jr HE, and Myers JD (1982). "Internist-I, an experimental computer-based diagnostic consultant for general internal medicine". New England Journal of Medicine. 307 (8): 468–476. doi:10.1056/NEJM198208193070803. PMID 7048091.
- Feigenbaum, Edward; McCorduck, Pamela (1984). The fifth generation. Addison-Wesley. pp. 1–275. ISBN 978-0451152640.
- Richard M. Karp (1972). "Reducibility Among Combinatorial Problems" (PDF). In R. E. Miller; J. W. Thatcher (eds.). Complexity of Computer Computations. New York: Plenum. pp. 85–103. Archived from the original (PDF) on 2011-06-29. Retrieved 2019-08-14.
- Doi K (2007). "Computer-aided diagnosis in medical imaging: historical review, current status and future potential". Computerized Medical Imaging and Graphics. 31 (4): 198–211. doi:10.1016/j.compmedimag.2007.02.002. PMC 1955762. PMID 17349778.
- Echegaray, Sebastian; Gevaert, Olivier; Shah, Rajesh; Kamaya, Aya; Louie, John; Kothary, Nishita; Napel, Sandy (18 November 2015). "Core samples for radiomics features that are insensitive to tumor segmentation: method and pilot study using CT images of hepatocellular carcinoma". Journal of Medical Imaging. 2 (4): 041011. doi:10.1117/1.JMI.2.4.041011. PMC 4650964. PMID 26587549.
- Murphy, K.; van Ginneken, B.; Schilham, A. M.; et al. (2009). "A large-scale evaluation of automatic pulmonary nodule detection in chest CT using local image features and k-nearest-neighbour classification". Medical Image Analysis. 13 (5): 757–770. doi:10.1016/j.media.2009.07.001. hdl:2066/81262. PMID 19646913.
- Suzuki, K.; Armato, 3rd, S. G.; Li, F.; Sone, S.; Doi, K. (2003). "Massive training artificial neural network (MTANN) for reduction of false positives in computerized detection of lung nodules in low-dose computed tomography". Medical Physics. 30 (7): 1602–1617. Bibcode:2003MedPh..30.1602S. doi:10.1118/1.1580485. PMID 12906178.
- Chan, H. P.; Lo, S. C.; Sahiner, B.; et al. (1995). "Computer-aided detection of mammographic microcalcifications: pattern recognition with an artificial neural network". Medical Physics. 22 (10): 1555–1567. Bibcode:1995MedPh..22.1555C. doi:10.1118/1.597428. hdl:2027.42/134770. PMID 8551980.
- Gletsos, Miltiades; Mougiakakou, Stavroula; Matsopoulos, George; Nikita, Konstantina; Nikita, Alexandra; Kelekis, Dimitrios (2003). "A computer-aided diagnostic system to characterize CT focal liver lesions: design and optimization of a neural network classifier". IEEE Transactions on Information Technology in Biomedicine. 7 (3): 153–162. doi:10.1109/TITB.2003.813793. PMID 14518728. S2CID 18918667.
- Mougiakakou, Stavroula; Golemati, Spyretta; Gousias, Ioannis; Nicolaides, Andrew; Nikita, Konstantina (2007). "Computer-aided diagnosis of carotid atherosclerosis based on ultrasound image statistics, laws' texture and neural networks". Ultrasound in Medicine and Biology. 33 (1): 26–36. doi:10.1016/j.ultrasmedbio.2006.07.032. PMID 17189044.
- Stoitsis, John; Valavanis, Ioannis; Mougiakakou, Stavroula; Golemati, Spyretta; Nikita, Alexandra; Nikita, Konstantina (2006). "Computer aided diagnosis based on medical image processing and artificial intelligence methods". Nuclear Instruments and Methods in Physics Research Section A: Accelerators, Spectrometers, Detectors and Associated Equipment. 569 (2): 591–595. Bibcode:2006NIMPA.569..591S. doi:10.1016/j.nima.2006.08.134.
- Chen, S.; Suzuki, K.; MacMahon, H. (2011). "Development and evaluation of a computer-aided diagnostic scheme for lung nodule detection in chest radiographs by means of two-stage nodule enhancement with support vector classification". Medical Physics. 38 (4): 1844–1858. Bibcode:2011MedPh..38.1844C. doi:10.1118/1.3561504. PMC 3069992. PMID 21626918.
- Papadopoulos, A.; Fotiadis, D. I.; Likas, A. (2005). "Characterization of clustered microcalcifications in digitized mammograms using neural networks and support vector machines". Artif Intell Med. 34 (2): 141–150. doi:10.1016/j.artmed.2004.10.001. PMID 15894178.
- Wollenweber T.; Janke B.; Teichmann A.; Freund M. (2007). "Korrelation zwischen histologischem Befund und einem Computer-assistierten Detektionssystem (CAD) für die Mammografie". Geburtsh Frauenheilk. 67 (2): 135–141. doi:10.1055/s-2006-955983. S2CID 73122975.
- Yanase J, Triantaphyllou E (2019). "The Seven Key Challenges for the Future of Computer-Aided Diagnosis in Medicine". International Journal of Medical Informatics. 129: 413–422. doi:10.1016/j.ijmedinf.2019.06.017. PMID 31445285. S2CID 198287435.
- Wadhwa, R. R.; Park, D. Y.; Natowicz, M. R. (2018). "The accuracy of computer‐based diagnostic tools for the identification of concurrent genetic disorders". American Journal of Medical Genetics Part A. 176 (12): 2704–2709. doi:10.1002/ajmg.a.40651. PMID 30475443. S2CID 53758271.
- Bron EE, Smits M, Van Der Flier WM, Vrenken H, Barkhof F, Scheltens P, Papma JM, Steketee RM, Orellana CM, Meijboom R, and Pinto M (2015). "Standardized evaluation of algorithms for computer-aided diagnosis of dementia based on structural MRI: the CAD Dementia challenge". NeuroImage. 111: 562–579. doi:10.1016/j.neuroimage.2015.01.048. PMC 4943029. PMID 25652394.
- Taylor P, Potts HW (2008). "Computer aids and human second reading as interventions in screening mammography: Two systematic reviews to compare effects on cancer detection and recall rate" (PDF). European Journal of Cancer. 44 (6): 798–807. doi:10.1016/j.ejca.2008.02.016. PMID 18353630.
- Benjamens, Stan; Dhunnoo, Pranavsingh; Mesko, Bertalan (2020). "The state of artificial intelligence-based FDA-approved medical devices and algorithms: an online database". npj Digital Medicine. 3: 118. doi:10.1038/s41746-020-00324-0. PMC 7486909. PMID 32984550.
- Abe, Yoshiyuki; Hanai, Kouzo; Nakano, Makiko; Ohkubo, Yasuyuki; Hasizume, Toshinori; Kakizaki, Toru; Nakamura, Masato; Niki, Noboru; Eguchi, Kenji (2005-01-01). "A Computer-aided Diagnosis (CAD) System in Lung Cancer Screening with Computed Tomography". Anticancer Research. 25 (1B): 483–488. ISSN 0250-7005. PMID 15816616.
- Wu N, Gamsu G, Czum J, Held B, Thakur R, Nicola G (Mar 2006). "Detection of small pulmonary nodules using direct digital radiography and picture archiving and communication systems". J Thorac Imaging. 21 (1): 27–31. doi:10.1097/01.rti.0000203638.28511.9b. PMID 16538152. S2CID 31230950.
- Giger, Maryellen Lissak; Doi, Kunio; MacMahon, Heber (1988-03-01). "Image feature analysis and computer-aided diagnosis in digital radiography. 3. Automated detection of nodules in peripheral lung fields". Medical Physics. 15 (2): 158–166. Bibcode:1988MedPh..15..158G. doi:10.1118/1.596247. ISSN 2473-4209. PMID 3386584.
- Ginneken, B. Van; Romeny, B. M. Ter Haar; Viergever, M. A. (2001-12-01). "Computer-aided diagnosis in chest radiography: a survey". IEEE Transactions on Medical Imaging. 20 (12): 1228–1241. doi:10.1109/42.974918. ISSN 0278-0062. PMID 11811823. S2CID 6280485.
- Coppini, G.; Diciotti, S.; Falchini, M.; Villari, N.; Valli, G. (2003-12-01). "Neural networks for computer-aided diagnosis: detection of lung nodules in chest radiograms". IEEE Transactions on Information Technology in Biomedicine. 7 (4): 344–357. doi:10.1109/TITB.2003.821313. ISSN 1089-7771. PMID 15000360. S2CID 15121082.
- Giger, M. L.; Bae, K. T.; MacMahon, H. (1994-04-01). "Computerized detection of pulmonary nodules in computed tomography images". Investigative Radiology. 29 (4): 459–465. doi:10.1097/00004424-199404000-00013. ISSN 0020-9996. PMID 8034453. S2CID 9800069.
- Kanazawa, K.; Kawata, Y.; Niki, N.; Satoh, H.; Ohmatsu, H.; Kakinuma, R.; Kaneko, M.; Moriyama, N.; Eguchi, K. (1998-03-01). "Computer-aided diagnosis for pulmonary nodules based on helical CT images". Computerized Medical Imaging and Graphics. 22 (2): 157–167. doi:10.1016/S0895-6111(98)00017-2. ISSN 0895-6111. PMID 9719856.
- Chen, Sheng; Zhong, Sikai; Yao, Liping; Shang, Yanfeng; Suzuki, Kenji (2016). "Enhancement of chest radiographs obtained in the intensive care unit through bone suppression and consistent processing". Physics in Medicine and Biology. 61 (6): 2283–2301. Bibcode:2016PMB....61.2283C. doi:10.1088/0031-9155/61/6/2283. PMID 26930386. S2CID 206020910.
- Chen, S.; Suzuki, K. (2014-02-01). "Separation of Bones From Chest Radiographs by Means of Anatomically Specific Multiple Massive-Training ANNs Combined With Total Variation Minimization Smoothing". IEEE Transactions on Medical Imaging. 33 (2): 246–257. doi:10.1109/TMI.2013.2284016. ISSN 0278-0062. PMID 24132005. S2CID 922550.
- Suzuki, K.; Abe, H.; MacMahon, H.; Doi, K. (2006-04-01). "Image-processing technique for suppressing ribs in chest radiographs by means of massive training artificial neural network (MTANN)". IEEE Transactions on Medical Imaging. 25 (4): 406–416. CiteSeerX 10.1.1.589.8748. doi:10.1109/TMI.2006.871549. ISSN 0278-0062. PMID 16608057. S2CID 17961280.
- LOOG, M; VANGINNEKEN, B; SCHILHAM, A (2006-12-01). "Filter learning: Application to suppression of bony structures from chest radiographs". Medical Image Analysis. 10 (6): 826–840. doi:10.1016/j.media.2006.06.002. ISSN 1361-8415. PMID 16859953.
- Chen, S.; Suzuki, K. (2013-02-01). "Computerized Detection of Lung Nodules by Means of #x201C;Virtual Dual-Energy #x201D; Radiography". IEEE Transactions on Biomedical Engineering. 60 (2): 369–378. doi:10.1109/TBME.2012.2226583. ISSN 0018-9294. PMC 4283823. PMID 23193306.
- Bell, L.T.O.; Gandhi, S. (2018). "A comparison of computer-assisted detection (CAD) programs for the identification of colorectal polyps: Performance and sensitivity analysis, current limitations and practical tips for radiologists". Clinical Radiology. 73 (6): 593.e11–593.e18. doi:10.1016/j.crad.2018.02.009. PMID 29602538. S2CID 4500060.
- Suzuki, Kenji; Yoshida, Hiroyuki; Näppi, Janne; Dachman, Abraham H. (2006-10-01). "Massive-training artificial neural network (MTANN) for reduction of false positives in computer-aided detection of polyps: Suppression of rectal tubes". Medical Physics. 33 (10): 3814–3824. Bibcode:2006MedPh..33.3814S. doi:10.1118/1.2349839. ISSN 2473-4209. PMID 17089846.
- Golemati, Spyretta; Nikita, Konstantina (2019). Cardiovascular Computing-Methodologies and Clinical Applications. Springer.
- Gastounioti, Aimilia; Makrodimitris, Stavros; Golemati, Spyretta; Kadoglou, Nikolaos; Liapis, Christos; Nikita, Konstantina (2014). "A novel computerized tool to stratify risk in carotid atherosclerosis using kinematic features of the arterial wall". IEEE Journal of Biomedical and Health Informatics. 19 (3): 1137–1145. doi:10.1109/JBHI.2014.2329604. PMID 24951709. S2CID 5924749.
- Golemati, Spyretta; Gastounioti, Aimilia; Nikita, Konstantina (2013). "Toward novel noninvasive and low-cost markers for predicting strokes in asymptomatic carotid atherosclerosis: the role of ultrasound image analysis". IEEE Transactions on Biomedical Engineering. 60 (3): 652–658. doi:10.1109/TBME.2013.2244601. PMID 23380846. S2CID 5653986.
- Golemati, Spyretta; Patelaki, Eleni; Gastounioti, Aimilia; Andreadis, Ioannis; Liapis, Christos; Nikita, Konstantina (2020). "Motion synchronization patterns of the carotid atheromatous plaque from B-mode ultrasound". Scientific Reports. 10 (1): 11221. Bibcode:2020NatSR..1011221G. doi:10.1038/s41598-020-65340-2. PMC 7343786. PMID 32641773.
- Rizi, Fereshteh; Au, Jason; Yli-Ollila, Heikki; Golemati, Spyretta; Makunaite, Monika; Orkisz, Maciej; Navab, Nassir; MacDonald, Maureen; Laitinen, Tiina Marja; Behnam, Hamid; Gao, Zhifan; Gastounioti, Aimilia; Jurkonis, Rytis; Vray, Didier; Laitinen, Tomi; Serusclat, Andre; Nikita, Konstantina; Zahnd, Guillaume (2020). "Carotid Wall Longitudinal Motion in Ultrasound Imaging: An Expert Consensus Review". Ultrasound in Medicine and Biology. 46 (10): 2605–2624. doi:10.1016/j.ultrasmedbio.2020.06.006. PMID 32709520. S2CID 225545904.
- Golemati, Spyretta; Gastounioti, Aimilia; Nikita, Konstantina (2016). "Ultrasound-Image-Based Cardiovascular Tissue Motion Estimation". IEEE Reviews on Biomedical Engineering. 9: 208–218. doi:10.1109/RBME.2016.2558147. S2CID 23333131.
- Gastounioti, Aimilia; Golemati, Spyretta; Stoitsis, John; Nikita, Konstantina (2013). "Carotid artery wall motion analysis from B-mode ultrasound using adaptive block matching: in silico evaluation and in vivo application". Physics in Medicine and Biology. 58 (24): 8647–8661. Bibcode:2013PMB....58.8647G. doi:10.1088/0031-9155/58/24/8647. PMID 24256708. S2CID 11571104.
- Gastounioti, Aimilia; Kolias, Vasileios; Golemati, Spyretta; Tsiaparas, Nikolaos; Matsakou, Aikaterini; Stoitsis, John; Kadoglou, Nikolaos; Gkekas, Christos; Kakisis, John; Liapis, Christos; Karakitsos, Petros; Sarafis, Ioannis; Angelidis, Padelis; Nikita, Konstantina (2014). "CAROTID - a web-based platform for optimal personalized management of atherosclerotic patients". Computer Methods and Programs in Biomedicine. 114 (2): 183–193. doi:10.1016/j.cmpb.2014.02.006. PMID 24636805.
- Goldenberg, Roman; Eilot, Dov; Begelman, Grigory; Walach, Eugene; Ben-Ishai, Eyal; Peled, Nathan (2012). "Computer-aided simple triage (CAST) for coronary CT angiography (CCTA)". International Journal of Computer Assisted Radiology and Surgery. 7 (6): 819–827. doi:10.1007/s11548-012-0684-7. ISSN 1861-6410. PMID 22484719. S2CID 5627031.
- Chaplot, Sandeep; Patnaik, L.M.; Jagannathan, N.R. (2006). "Classification of magnetic resonance brain images using wavelets as input to support vector machine and neural network". Biomedical Signal Processing and Control. 1: 86–92. doi:10.1016/j.bspc.2006.05.002.
- Maitra, Madhubanti; Chatterjee, Amitava (2006). "A Slantlet transform based intelligent system for magnetic resonance brain image classification". Biomedical Signal Processing and Control. 1 (4): 299–306. doi:10.1016/j.bspc.2006.12.001.
- Wang, S.; Wu, W. (2010). "A Novel Method for Magnetic Resonance Brain Image Classification based on Adaptive Chaotic PSO". Progress in Electromagnetics Research. 109: 325–343. doi:10.2528/PIER10090105.
- Zhang, Yudong; Wu, L. (2011). "Magnetic Resonance Brain Image Classification by an Improved Artificial Bee Colony Algorithm". Progress in Electromagnetics Research. 2011: 65–79. doi:10.2528/PIER11031709.
- Padma, A.; Sukanesh, R. (2014). "Segmentation and Classification of Brain CT Images Using Combined Wavelet Statistical Texture Features". Arabian Journal for Science and Engineering. 39 (2): 767–776. doi:10.1007/s13369-013-0649-3. S2CID 62615810.
- Zhang, Yudong; Dong, Zhengchao; Ji, Genlin (2015). "Effect of spider-web-plot in MR brain image classification". Pattern Recognition Letters. 62: 14–16. Bibcode:2015PaReL..62...14Z. doi:10.1016/j.patrec.2015.04.016.
- Zhang, Y.; Wang, S.; Ji, G.; Dong, Z. (2013). "Genetic Pattern Search and its Application to Brain Image Classification". Mathematical Problems in Engineering. 2013: 1–8. doi:10.1155/2013/580876.
- Das S.; Chowdhury M.; Kundu M.K. (2013). "Brain MR Image Classification Using Multiscale Geometric Analysis of Ripplet". Progress in Electromagnetics Research. 137: 1–17. doi:10.2528/pier13010105.
- Zhang, Y.; Wang, S. (2013). "An MR Brain Images Classifier System via Particle Swarm Optimization and Kernel Support Vector Machine". The Scientific World Journal. 2013: 130134. doi:10.1155/2013/130134. PMC 3791634. PMID 24163610.
- Kalbkhani H.; Shayesteh M.G.; Zali-Vargahan B. (2013). "Robust algorithm for brain magnetic resonance image (MRI) classification based on GARCH variances series". Biomedical Signal Processing and Control. 8 (6): 909–919. doi:10.1016/j.bspc.2013.09.001.
- El-Dahshan E.S.A.; Mohsen H.M.; Revett K.; et al. (2014). "Computer-aided diagnosis of human brain tumor through MRI: A survey and a new algorithm". Expert Systems with Applications. 41 (11): 5526–5545. doi:10.1016/j.eswa.2014.01.021.
- Zhou, Xing-Xing (2015). "Detection of Pathological Brain in MRI Scanning Based on Wavelet-Entropy and Naive Bayes Classifier". Bioinformatics and Biomedical Engineering. Lecture Notes in Computer Science. Vol. 9043. pp. 201–209. doi:10.1007/978-3-319-16483-0_20. ISBN 978-3-319-16482-3.
- Zhang, Yudong; Wang, Shuihua; Dong, Zhengchao (2014). "Classification of Alzheimer Disease Based on Structural Magnetic Resonance Imaging by Kernel Support Vector Machine Decision Tree". Progress in Electromagnetics Research. 144: 185–191. doi:10.2528/PIER13121310.
- Signaevsky, Maxim; Prastawa, Marcel; Farrell, Kurt; Tabish, Nabil; Baldwin, Elena; Han, Natalia; Iida, Megan; Koll, John; Bryce, Clare; Purohit, Dushyant; Haroutunian, Vahram; McKee, Ann; Stein, Thor; White-III, Charles; Walker, Jamie; Richardson, Timothy; Hanson, Russell; Cordon-Cardo, Carlos; Donovan, Michael; Zeineh, Jack; Fernandez, Gerardo; Crary, John (2019). "Artificial intelligence in neuropathology: deep learning-based assessment of tauopathy". Laboratory Investigation. 99 (7): 1019–1029. doi:10.1038/s41374-019-0202-4. PMC 7684013. PMID 30770886.
- Friston, K.; Poline, J-P.; Holmes, C.J.; Frith, C.D.; Frackowiak, R.S. (1996). "A multivariate Analysis of PET activation studies". Hum. Brain Mapp. 4 (2): 140–151. doi:10.1002/(SICI)1097-0193(1996)4:2<140::AID-HBM5>3.0.CO;2-3. PMID 20408193. S2CID 9074888.
- Martínez-Murcia, F.J.; Górriz, J.M.; Ramírez, J.; Puntonet, C.G.; Illán, I.A. (2013). "Functional activity maps based on significance measures and Independent Component Analysis". Computer Methods and Programs in Biomedicine. 111 (1): 255–268. doi:10.1016/j.cmpb.2013.03.015. PMC 6701938. PMID 23660005.
- Dong, Z.C. (2015). "Detection of subjects and brain regions related to Alzheimer's disease using 3D MRI scans based on eigenbrain and machine learning". Frontiers in Computational Neuroscience. 66 (9): 66. doi:10.3389/fncom.2015.00066. PMC 4451357. PMID 26082713.
- Zhang, J.; Yu, C.; Jiang, G.; Liu, W.; Tong, L. (2012). "3d texture analysis on mri images of alzheimer's disease". Brain Imaging and Behavior. 6 (1): 61–69. doi:10.1007/s11682-011-9142-3. PMID 22101754. S2CID 10069584.
- Chupin, Marie; Gérardin, Emilie; Cuingnet, Rémi; Boutet, Claire; Lemieux, Louis; Lehéricy, Stéphane; Benali, Habib; Garnero, Line; Colliot, Olivier (2009). "Fully automatic hippocampus segmentation and classification in Alzheimer's disease and mild cognitive impairment applied on data from ADNI". Hippocampus. 19 (6): 579–587. doi:10.1002/hipo.20626. PMC 2837195. PMID 19437497.
- Martinez-Murcia, F.J.; Gorriz, J.M.; Ramirez, J.; Ortiz, A. (2016). "A Spherical Brain Mapping of MR Images for the Detection of Alzheimer's Disease". Current Alzheimer Research. 13 (5): 575–588. doi:10.2174/1567205013666160314145158. hdl:10481/42543. PMID 26971941.
- "EXINI Diagnostics".
- Huang, Kao and Chen (18 June 2007). "A Set of Image Processing Algorithms for Computer-Aided Diagnosis in Nuclear Medicine Whole Body Bone Scan Images". IEEE Transactions on Nuclear Science. 54 (3): 514–522. Bibcode:2007ITNS...54..514H. doi:10.1109/TNS.2007.897830. S2CID 20730927.
- Kaur, M; Talwar, R (2014). "Review on: blood vessel extraction and eye retinopathy detection". International Journal of Computer Science and Information Technologies. 5 (6): 7513–7516. S2CID 17460643.
- Tufail, A; Rudisill, C; Egan, C; Kapetanakis, VV; Salas-Vega, S; Owen, CG; Lee, A; Louw, V; Anderson, J; Liew, G; Bolter, L; Srinivas, S; Nittala, M; Sadda, S; Taylor, P; Rudnicka, AR (n.d.). "Automated diabetic retinopathy image assessment software: diagnostic accuracy and cost-effectiveness compared to human graders". Ophthalmology. 124 (3): 343–351. doi:10.1016/j.ophtha.2016.11.014. PMID 28024825.
- Ahmad, A.; Mansoor, A. B.; Mumtaz, R.; Khan, M.; Mirza, S. H. (2014-12-01). "Image processing and classification in diabetic retinopathy: A review". 2014 5th European Workshop on Visual Information Processing (EUVIP). pp. 1–6. doi:10.1109/EUVIP.2014.7018362. ISBN 978-1-4799-4572-6. S2CID 16465894.
- Fraz, M. M.; Barman, S. A.; Remagnino, P.; Hoppe, A.; Basit, A.; Uyyanonvara, B.; Rudnicka, A. R.; Owen, C. G. (2012-11-01). "An Approach to Localize the Retinal Blood Vessels Using Bit Planes and Centerline Detection". Comput. Methods Prog. Biomed. 108 (2): 600–616. doi:10.1016/j.cmpb.2011.08.009. ISSN 0169-2607. PMID 21963241.
- Priya, R; Aruna, P (2011). "Review of automated diagnosis of diabetic retinopathy using the support vector machine". International Journal of Applied Engineering Research, Dindigul. 1 (4): 844–862.
- Saleh, Marwan D.; Eswaran, C. (2012-10-01). "An Automated Decision-support System for Non-proliferative Diabetic Retinopathy Disease Based on MAs and HAs Detection". Comput. Methods Prog. Biomed. 108 (1): 186–196. doi:10.1016/j.cmpb.2012.03.004. ISSN 0169-2607. PMID 22551841.
- Antal, B.; Hajdu, A. (2012-06-01). "An Ensemble-Based System for Microaneurysm Detection and Diabetic Retinopathy Grading". IEEE Transactions on Biomedical Engineering. 59 (6): 1720–1726. arXiv:1410.8577. doi:10.1109/TBME.2012.2193126. ISSN 0018-9294. PMID 22481810. S2CID 16382245.
- Administrator (2015-05-20). "Review on: Detection of Diabetic Retinopathy using SVM and MDA". International Journal of Computer Applications. 117 (20): 1–3. Bibcode:2015IJCA..117t...1S. doi:10.5120/20667-2485.
- Espona, L.; Carreira, M. J.; Ortega, M.; Penedo, M. G. (2007-06-06). Martí, Joan; Benedí, José Miguel; Mendonça, Ana Maria; Serrat, Joan (eds.). Pattern Recognition and Image Analysis. Lecture Notes in Computer Science. Springer Berlin Heidelberg. pp. 178–185. doi:10.1007/978-3-540-72849-8_23. ISBN 9783540728481.
- Paiva, Omir Antunes; Prevedello, Luciano M. (October 2017). "The potential impact of artificial intelligence in radiology". Radiologia Brasileira. 50 (5): V–VI. doi:10.1590/0100-3984.2017.50.5e1. PMC 5656066. PMID 29085178.
- Mukherjee, Siddhartha (27 March 2017). "A.I. Versus M.D." The New Yorker. Retrieved 3 February 2018.
- "Why scan-reading artificial intelligence is bad news for radiologists". The Economist. 29 November 2017. Retrieved 3 February 2018.
- Jha, Saurabh; Topol, Eric J. (13 December 2016). "Adapting to Artificial Intelligence". JAMA. 316 (22): 2353–2354. doi:10.1001/jama.2016.17438. PMID 27898975. S2CID 3662362.
External links
- Digital Retinal Images for Vessel Extraction (DRIVE)
- STructured Analysis of the REtina (STARE)
- High-Resolution Fundus (HRF) Image Database