Dispersin B
Dispersin B (also known as DspB or DispersinB) is a 40 kDa glycoside hydrolase produced by the periodontal pathogen, Aggregatibacter actinomycetemcomitans.[1] The bacteria secrete Dispersin B to release adherent cells from a mature biofilm colony by disrupting biofilm formation. The enzyme catalyzes the hydrolysis of linear polymers of N-acetyl-D-glucosamines found in the biofilm matrices. Poly-acetyl glucosamines are integral to the structural integrity of the biofilms of various Gram-positive bacteria and Gram-negative bacteria and are referred to as PIA (PNAG,PS/A) in Staphylococcus species and PGA in Escherichia coli.[2][3] By degrading the biofilm matrix, Dispersin B allows for the release of bacterial cells that can adhere to new surfaces close by and extend the biofilm or start new colonies. Currently there is interest in Dispersin B as a commercial anti-biofilm agent that could be combined with antibiotics for the treatment of bacterial infections.
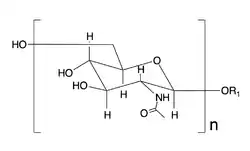
| Dispersin B | |||||||||
|---|---|---|---|---|---|---|---|---|---|
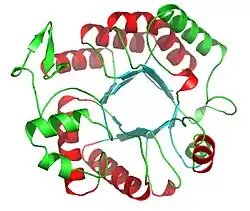 Ribbon diagram of Dispersin B | |||||||||
| Identifiers | |||||||||
| EC no. | 3.2.1.52 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
Biological function
Dispersin B is produced by Aggregatibacter actinomycetemcomitans, a Gram-negative oral bacterium, when it needs to detach and disperse adherent bacterial cells.[4] A. actinomycetemcomitans forms asymmetric biofilm lobed colonies that release single cells or small clusters of bacterial cells, which can attach to nearby surfaces, form new colonies, and enable the biofilm to spread.[2][5] A biofilm is a matrix of extracellular polymeric substances that are synthesized by the bacteria. The biofilm's structural integrity is dependent on poly-N-acetylglucosamine (PGA), extracellular DNA, and proteinaceous adhesins. It allows bacteria to adhere to host surfaces, protects the bacterial cells from host defenses, results in increased resistance to antibiotics, and provides a protected environment with microchannels for the flow of water and other essential nutrients.[2] By hydrolyzing PGA, Dispersin B disrupts the formation of the biofilm matrix and allows adherent cells to be released.[6] Dispersin B has also been shown to cause the detachment of biofilm cells that have adhered to abiotic surfaces as well as cause the disaggregaton of highly auto-aggregated clumps of bacterial cells.[2]
Enzyme structure
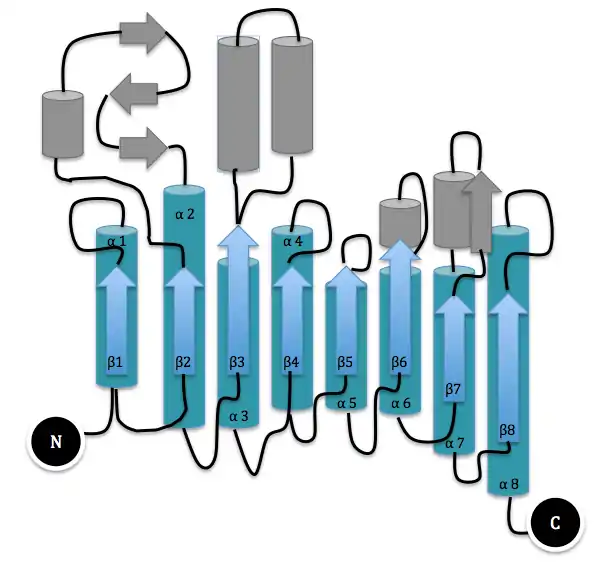
The three-dimensional structure of dispersin B was determined by expressing the protein in E. coli, purifying it from cultures, and crystallizing it in the orthoorhombic space group C2221 with one molecule in the asymmetric unit.[2] It is a 361 amino acid protein consisting of a single domain. The major substructure consists of residues 1-60 and residues 91-358, which fold into a β/α structure of a typical TIM barrel. The loop β4 contains two highly conserved residues Asp183 and Glu184 that are part of the enzyme's active site, a large cavity in the center of the bowl-shaped Dispersin B molecule. The oligomer substrate binding and cleaving at the active site of Dispersin B was studied to learn more about the enzymatic degradation.[7] Reverse phase HPLC was used to analyze the products of the hydrolysis of substrates with differing degrees of polymerization. Chain lengthening of the substrate was shown to increase the catalytic efficiency of Dispersin B. A substrate with a degree of polymerization of five was not enough for total subsite occupancy.[7] Carbamate substrates were also shown to improve the activity of the enzyme relative to the benchmark substrate 4-nitrophenyl N-acetyl-β-D-glucosamine.[8]
Enzyme mechanism
Dispersin B is a β-hexosaminidase that specifically hydrolyzes β-1,6-glycosidic linkages of acetylglucosamine polymers found in biofilm matrices.[9] As a member of family 20 β-hexosaminidases, it cleaves terminal monosaccharide residues from the non-reducing end of the polymers. The active site of Dispersin B contains three highly conserved acidic residues: an aspartic acid at residue 183 (D183), a glutamic acid at residue 184 (E184), and a glutamic acid at residue 332 (E332).[10] In the proposed mechanism, E184 serves as the acid/base and donates a proton to the -OR on C1. Crystallographic evidence suggests that substrate-assisted catalysis occurs and D183 is believed to assist in activating the N-acetyl group, which acts as a nucleophilic in an attack of C1.[11] Water then attacks the anomeric centre to release the cleaved polymer from the active site with retention of anomeric configuration. (The mechanism shown below is thus incorrect).[10][12] The use of the neighboring C2-acetamido group of the substrate differs from the majority of β-glycosidases, which use a carboxyl group as a nucleophile in their mechanisms.[13][14] Although E332 is located farther away from the anomeric carbon, the lower activity of Dispersin B with the mutation E332Q suggests that it is important for catalysis. It may be critical in stabilizing the transition state of cleaving the terminal monosaccharide.[15]
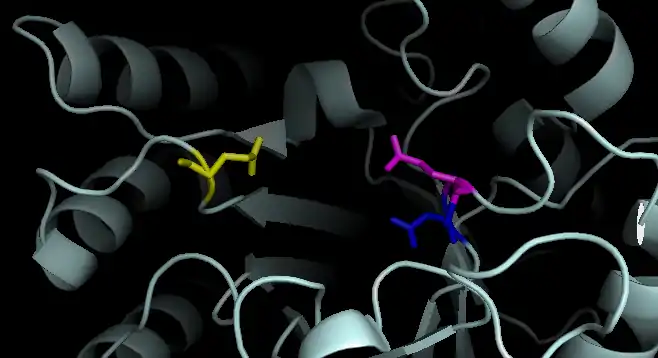 Active Site of Dispersin B: E184 shown in blue. D183 shown in pink. E332 shown in yellow.
Active Site of Dispersin B: E184 shown in blue. D183 shown in pink. E332 shown in yellow. Proposed Mechanism of Dispersin B: E184 shown in blue. D183 shown in pink. E332 not shown.
Proposed Mechanism of Dispersin B: E184 shown in blue. D183 shown in pink. E332 not shown.
Applications
Dispersin B may be useful for treating and preventing biofilm-associated infections caused by poly-N-acetylglucosamine-producing bacteria. Dispersin B prevented the formation of S. epidermidis biofilm, which suggests that biofilm-releasing enzymes can exhibit broad-spectrum activity and could be potential antibiofilm agents.[16]
Commercial development
Dispersin B is being commercially developed as a wound care gel and medical device coating by Kane Biotech, Inc., a Canadian biotech company. External and internal catheters that were coated with a combination of Dispersin B and triclosan were shown to be as effective at preventing bacterial infection as already commercially available chlorhexidine-sulfadiazine-coated catheters. These findings support that Dispersin B exhibits synergistic activity when combined with antibiotics.[7] Carbamate substrates were also shown to improve the activity of the enzyme relative to the benchmark substrate 4-nitrophenyl N-acetyl-β-D-glucosamine.[17] Pharmaceutical-grade Dispersin B has been successfully manufactured by BioVectra, Inc., Charlottetown, P.E.I., Canada, and is currently undergoing biocompatibility testing. Clinical trials to test the effectiveness of dispersin B for the treatment and prevention of diabetic foot ulcers and pressure sores should begin in 2010.
References
- Kaplan JB, Ragunath C, Ramasubbu N, Fine DH (August 2003). "Detachment of Actinobacillus actinomycetemcomitans biofilm cells by an endogenous beta-hexosaminidase activity". J. Bacteriol. 185 (16): 4693–8. doi:10.1128/JB.185.16.4693-4698.2003. PMC 166467. PMID 12896987.
- PDB: 1YHT; Ramasubbu N, Thomas LM, Ragunath C, Kaplan JB (June 2005). "Structural analysis of dispersin B, a biofilm-releasing glycoside hydrolase from the periodontopathogen Actinobacillus actinomycetemcomitans". J. Mol. Biol. 349 (3): 475–86. doi:10.1016/j.jmb.2005.03.082. PMID 15878175.
- Mack D, Fischer W, Krokotsch A, Leopold K, Hartmann R, Egge H, Laufs R (1996). "The intercellular adhesin involved in biofilm accumulation of Staphylococcus epidermidis is a linear beta-1,6-linked glucosaminoglycan: purification and structural analysis". J. Bacteriol. 178 (1): 175–183. PMC 177636. PMID 8550413.
- Blackledge MS, Worthington RJ, Melander C (2013). "Biologically inspired strategies for combating bacterial biofilms". Curr Opin Pharmacol. 13 (5): 699–706. doi:10.1016/j.coph.2013.07.004. PMC 3795836. PMID 23871261.
- Kaplan JB, Meyenhofer MF, Fine DH (2003). "Biofilm growth and detachment of Actinobacillus actinomycetemcomitans". J. Bacteriol. 185 (4): 1399–1404. doi:10.1128/JB.185.4.1399-1404.2003. PMC 142852. PMID 12562811.
- Izano EA, Sadovskaya I, Vinogradov E, Mulks MH, Velliyagounder K, Ragunath C, Kher WB, Ramasubbu N, Jabbouri S, Perry MB, Kaplan JB (2007). "Poly-N-acetylglucosamine mediates biofilm formation and antibiotic resistance in Actinobacillus pleuropneumoniae". Microb Pathog. 43 (1): 1–9. doi:10.1016/j.micpath.2007.02.004. PMC 1950449. PMID 17412552.
- Fazekas E, Kandra L, Gyémánt G (2012). "Model for β-1,6-N-acetylglucosamine oligomer hydrolysis catalysed by DispersinB, a biofilm degrading enzyme". Carbohydrate Res. 363: 7–13. doi:10.1016/j.carres.2012.09.016. PMID 23103508.
- Chibba A, Dasgupta S, Yakandawala N, Madhyastha S, Nitz M (2011). "Chromogenic Carbamate and Acetal Substrates for Glycosaminidases". J. Carbohydr. Chem. 30: 549–558. doi:10.1080/07328303.2011.610543. ISSN 0732-8303.
- Itoh Y, Wang X, Hinnebusch BJ, Preston JF, Romeo T (2005). "Depolymerization of beta-1,6-N-acetyl-D-glucosamine disrupts the integrity of diverse bacterial biofilms". J. Bacteriol. 187 (1): 382–386. doi:10.1128/JB.187.1.382-387.2005. PMC 538831. PMID 15601723.
- Manuel SG, Ragunath C, Sait HB, Izano EA, Kaplan JB, Ramasubbu N (2007). "Role of active-site residues of dispersin B, a biofilm-releasing beta-hexosaminidase from a periodontal pathogen, in substrate hydrolysis". FEBS J. 274 (22): 5987–99. doi:10.1111/j.1742-4658.2007.06121.x. PMID 17949435.
- Mark BL, Vocadlo DJ, Knapp S, Triggs-Raine BL, Withers SG, James MN (2001). "Crystallographic evidence for substrate-assisted catalysis in a bacterial beta-hexosaminidase". J Biol Chem. 276 (13): 10330–10337. doi:10.1074/jbc.M011067200. PMID 11124970.
- Hou Y, Vocadlo DJ, Leung A, Withers SG, Mahuran D (2001). "Characterization of the Glu and Asp residues in the active site of human beta-hexosaminidase B". Biochemistry. 40 (7): 2201–2209. doi:10.1021/bi002018s. PMC 2910085. PMID 11329289.
- Maier T, Strater N, Schuette CG, Klingenstein R, Sandhoff K, Saenger W (2003). "The X-ray crystal structure of human beta-hexosaminidase B provides new insights into Sandhoff disease". J Mol Biol. 328 (3): 669–681. doi:10.1016/s0022-2836(03)00311-5. PMID 12706724.
- Drouillard S, Armand S, Davies GJ, Vorgias CE, Henrissat B (1997). "hem J. 1997 Dec 15;328 ( Pt 3):945-9. Serratia marcescens chitobiase is a retaining glycosidase utilizing substrate acetamido group participation". Biochem. J. 328 (3): 945–949. doi:10.1042/bj3280945. PMC 1219008. PMID 9396742.
- Williams SJ, Mark BL, Vocadlo DJ, James MN, Withers SG (2002). "Aspartate 313 in the Streptomyces plicatus hexosaminidase plays a critical role in substrate-assisted catalysis by orienting the 2-acetamido group and stabilizing the transition state". J Biol Chem. 277 (42): 40055–40065. doi:10.1074/jbc.M206481200. PMID 12171933.
- Kaplan JB, Ragunath C, Velliyagounder K, Fine DH, Ramasubbu N (2004). "Enzymatic detachment of Staphylococcus epidermidis biofilms". Antimicrob. Agents Chemother. 48 (7): 2633–2636. doi:10.1128/AAC.48.7.2633-2636.2004. PMC 434209. PMID 15215120.
- Darouiche RO, Mansouri MD, Gawande PV, Madhyastha S (2009). "Antimicrobial and antibiofilm efficacy of triclosan and DispersinB combination". Journal of Antimicrobial Chemotherapy. 64 (1): 88–93. doi:10.1093/jac/dkp158. PMID 19447791.