Formins
Formins (formin homology proteins) are a group of proteins that are involved in the polymerization of actin and associate with the fast-growing end (barbed end) of actin filaments.[2] Most formins are Rho-GTPase effector proteins. Formins regulate the actin and microtubule cytoskeleton [3][4] and are involved in various cellular functions such as cell polarity, cytokinesis, cell migration and SRF transcriptional activity.[5] Formins are multidomain proteins that interact with diverse signalling molecules and cytoskeletal proteins, although some formins have been assigned functions within the nucleus.
| formin 1 | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | FMN1 | ||||||
| Alt. symbols | LD, FMN | ||||||
| NCBI gene | 342184 | ||||||
| HGNC | 3768 | ||||||
| OMIM | 136535 | ||||||
| RefSeq | NM_001103184 | ||||||
| UniProt | Q68DA7 | ||||||
| Other data | |||||||
| Locus | Chr. 15 q13-q14 | ||||||
| |||||||
| Formin Homology Region 1 | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Drf_FH1 | ||||||||
| Pfam | PF06346 | ||||||||
| InterPro | IPR009408 | ||||||||
| |||||||||
| Formin Homology 2 Domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
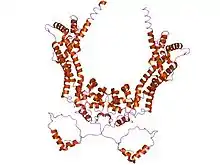 crystal structures of a formin homology-2 domain reveal a tethered-dimer architecture | |||||||||
| Identifiers | |||||||||
| Symbol | FH2 | ||||||||
| Pfam | PF02181 | ||||||||
| InterPro | IPR015425 | ||||||||
| SMART | FH2 | ||||||||
| SCOP2 | 1ux5 / SCOPe / SUPFAM | ||||||||
| |||||||||
| Diaphanous FH3 Domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of mdia1 gbd-fh3 in complex with rhoc-gmppnp | |||||||||
| Identifiers | |||||||||
| Symbol | Drf_FH3 | ||||||||
| Pfam | PF06367 | ||||||||
| Pfam clan | CL0020 | ||||||||
| InterPro | IPR010472 | ||||||||
| |||||||||
| DRF Autoregulatory Domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of the n-terminal mdia1 armadillo repeat region and dimerisation domain in complex with the mdia1 autoregulatory domain (dad) | |||||||||
| Identifiers | |||||||||
| Symbol | Drf_DAD | ||||||||
| Pfam | PF06345 | ||||||||
| InterPro | IPR010465 | ||||||||
| |||||||||
| Diaphanous GTPase-binding Domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of mdia1 gbd-fh3 in complex with rhoc-gmppnp | |||||||||
| Identifiers | |||||||||
| Symbol | Drf_GBD | ||||||||
| Pfam | PF06371 | ||||||||
| Pfam clan | CL0020 | ||||||||
| InterPro | IPR010473 | ||||||||
| |||||||||
Diversity
Formins have been found in all eukaryotes studied.[1] In humans, 15 different formin proteins are present that have been classified in 7 subgroups.[6] By contrast, yeasts contain only 2-3 formins.[7]
Structure and interactions
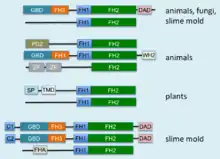
Formins are characterized by the presence of three formin homology (FH) domains (FH1, FH2 and FH3), although members of the formin family do not necessarily contain all three domains.[8][9] In addition, other domains are usually present, such as PDZ, DAD, WH2, or FHA domains.
The proline-rich FH1 domain mediates interactions with a variety of proteins, including the actin-binding protein profilin,[10] SH3 (Src homology 3) domain proteins,[11] and WW domain proteins. The actin nucleation-promoting activity of S. cerevisiae formins has been localized to the FH2 domain.[4] The FH2 domain is required for the self-association of formin proteins through the ability of FH2 domains to directly bind each other, and may also act to inhibit actin polymerization.[12][13] The FH3 domain is less well conserved and is required for directing formins to the correct intracellular location, such the mitotic spindle, or the projection tip during conjugation.[14][15] In addition, some formins can contain a GTPase-binding domain (GBD) required for binding to Rho small GTPases, and a C-terminal conserved Dia-autoregulatory domain (DAD). The GBD is a bifunctional autoinhibitory domain that interacts with and is regulated by activated Rho family members. Mammalian Drf3 contains a CRIB-like motif within its GBD for binding to Cdc42, which is required for Cdc42 to activate and guide Drf3 towards the cell cortex where it remodels the actin skeleton.[16] The DAD binds the N-terminal GBD; this link is broken when GTP-bound Rho binds to the GBD and activates the protein. The addition of the DAD to mammalian cells induces actin filament formation, stabilizes microtubules, and activates SRF mediated transcription.[16] Another commonly found domain is an armadillo repeat region (ARR) located in the FH3 domain.
The FH2 domain, has been shown by X-ray crystallography to have an elongated, crescent shape containing three helical subdomains.[17][18]
Formins also directly bind to microtubules via their FH2 domain. This interaction is important in promoting the capture and stabilization of a subset of microtubules oriented towards the leading edge of migrating cells. Formins also promote the capture of microtubules by the kinetochore during mitosis and for aligning microtubules along actin filaments.[19][20]
See also
References
- Chalkia D, Nikolaidis N, Makalowski W, Klein J, Nei M (December 2008). "Origins and evolution of the formin multigene family that is involved in the formation of actin filaments". Molecular Biology and Evolution. 25 (12): 2717–33. doi:10.1093/molbev/msn215. PMC 2721555. PMID 18840602.
- Evangelista M, Zigmond S, Boone C (July 2003). "Formins: signaling effectors for assembly and polarization of actin filaments". Journal of Cell Science. 116 (Pt 13): 2603–11. doi:10.1242/jcs.00611. PMID 12775772.
- Gunning PW, Ghoshdastider U, Whitaker S, Popp D, Robinson RC (June 2015). "The evolution of compositionally and functionally distinct actin filaments". Journal of Cell Science. 128 (11): 2009–19. doi:10.1242/jcs.165563. PMID 25788699.
- Goode BL, Eck MJ (2007). "Mechanism and function of formins in the control of actin assembly". Annual Review of Biochemistry. 76: 593–627. doi:10.1146/annurev.biochem.75.103004.142647. PMID 17373907.
- Faix J, Grosse R (June 2006). "Staying in shape with formins". Developmental Cell. 10 (6): 693–706. doi:10.1016/j.devcel.2006.05.001. PMID 16740473.
- Higgs HN, Peterson KJ (January 2005). "Phylogenetic analysis of the formin homology 2 domain". Molecular Biology of the Cell. 16 (1): 1–13. doi:10.1091/mbc.E04-07-0565. PMC 539145. PMID 15509653.
- Baarlink C, Brandt D, Grosse R (July 2010). "SnapShot: Formins". Cell. 142 (1): 172–172.e1. doi:10.1016/j.cell.2010.06.030. PMID 20603022. S2CID 2914004.
- Kitayama C, Uyeda TQ (February 2003). "ForC, a novel type of formin family protein lacking an FH1 domain, is involved in multicellular development in Dictyostelium discoideum". Journal of Cell Science. 116 (Pt 4): 711–23. doi:10.1242/jcs.00265. PMID 12538772.
- Wallar BJ, Alberts AS (August 2003). "The formins: active scaffolds that remodel the cytoskeleton". Trends in Cell Biology. 13 (8): 435–46. doi:10.1016/S0962-8924(03)00153-3. PMID 12888296.
- Uetz, Peter (1997). Biochemische Studien am limb deformity-Protein der Vertebraten: Inaugural-Dissertation zur Erlangung der Doktorwürde der Naturwissenschaftlich-Mathematischen Gesamtfakultät der Ruprecht-Karls-Universität Heidelberg. Developmental Biology, European Molecular Biology Laboratory. Heidelberg: European Molecular Biology Laboratory.
- Uetz P, Fumagalli S, James D, Zeller R (December 1996). "Molecular interaction between limb deformity proteins (formins) and Src family kinases". The Journal of Biological Chemistry. 271 (52): 33525–30. doi:10.1074/jbc.271.52.33525. PMID 8969217.
- Takeya R, Sumimoto H (November 2003). "Fhos, a mammalian formin, directly binds to F-actin via a region N-terminal to the FH1 domain and forms a homotypic complex via the FH2 domain to promote actin fiber formation". Journal of Cell Science. 116 (Pt 22): 4567–75. doi:10.1242/jcs.00769. PMID 14576350.
- Shimada A, Nyitrai M, Vetter IR, Kühlmann D, Bugyi B, Narumiya S, Geeves MA, Wittinghofer A (February 2004). "The core FH2 domain of diaphanous-related formins is an elongated actin binding protein that inhibits polymerization". Molecular Cell. 13 (4): 511–22. doi:10.1016/S1097-2765(04)00059-0. PMID 14992721.
- Kato T, Watanabe N, Morishima Y, Fujita A, Ishizaki T, Narumiya S (February 2001). "Localization of a mammalian homolog of diaphanous, mDia1, to the mitotic spindle in HeLa cells". Journal of Cell Science. 114 (Pt 4): 775–84. doi:10.1242/jcs.114.4.775. hdl:2433/150544. PMID 11171383.
- Petersen J, Nielsen O, Egel R, Hagan IM (June 1998). "FH3, a domain found in formins, targets the fission yeast formin Fus1 to the projection tip during conjugation". The Journal of Cell Biology. 141 (5): 1217–28. doi:10.1083/jcb.141.5.1217. PMC 2137179. PMID 9606213.
- Peng J, Wallar BJ, Flanders A, Swiatek PJ, Alberts AS (April 2003). "Disruption of the Diaphanous-related formin Drf1 gene encoding mDia1 reveals a role for Drf3 as an effector for Cdc42". Current Biology. 13 (7): 534–45. doi:10.1016/S0960-9822(03)00170-2. PMID 12676083. S2CID 13902104.
- Xu Y, Moseley JB, Sagot I, Poy F, Pellman D, Goode BL, Eck MJ (March 2004). "Crystal structures of a Formin Homology-2 domain reveal a tethered dimer architecture". Cell. 116 (5): 711–23. doi:10.1016/S0092-8674(04)00210-7. PMID 15006353. S2CID 15855545.
- Thompson ME, Heimsath EG, Gauvin TJ, Higgs HN, Kull FJ (January 2013). "FMNL3 FH2-actin structure gives insight into formin-mediated actin nucleation and elongation". Nature Structural & Molecular Biology. 20 (1): 111–8. doi:10.1038/nsmb.2462. PMC 3876896. PMID 23222643.
- Palazzo AF, Cook TA, Alberts AS, Gundersen GG (August 2001). "mDia mediates Rho-regulated formation and orientation of stable microtubules". Nature Cell Biology. 3 (8): 723–9. doi:10.1038/35087035. PMID 11483957. S2CID 7374170.
- Bartolini F, Gundersen GG (February 2010). "Formins and microtubules". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1803 (2): 164–73. doi:10.1016/j.bbamcr.2009.07.006. PMC 2856479. PMID 19631698.