Inbreeding depression
Inbreeding depression is the reduced biological fitness which has the potential to result from inbreeding (the breeding of related individuals). Biological fitness refers to an organism's ability to survive and perpetuate its genetic material. Inbreeding depression is often the result of a population bottleneck. In general, the higher the genetic variation or gene pool within a breeding population, the less likely it is to suffer from inbreeding depression, though inbreeding and outbreeding depression can simultaneously occur.
Inbreeding depression seems to be present in most groups of organisms, but varies across mating systems. Hermaphroditic species often exhibit lower degrees of inbreeding depression than outcrossing species, as repeated generations of selfing is thought to purge deleterious alleles from populations. For example, the outcrossing nematode (roundworm) Caenorhabditis remanei has been demonstrated to suffer severely from inbreeding depression, unlike its hermaphroditic relative C. elegans, which experiences outbreeding depression.[2]
Mechanisms

Inbreeding (i.e., breeding between closely related individuals) results in more recessive traits manifesting themselves, as the genomes of pair-mates are more similar. Recessive traits can only occur in an offspring if present in both parents' genomes. The more genetically similar the parents are, the more often recessive traits appear in their offspring. This normally has a positive effect, as most genes are undergoing purifying selection (the homozygous state is favored). However, for very closely related individuals, there is an increased likelihood of homozygous deleterious genes in the offspring which can result in unfit individuals.[3] For the alleles that confer an advantage in the heterozygous and/or homozygous-dominant state, the fitness of the homozygous-recessive state may even be zero (meaning sterile or unviable offspring).
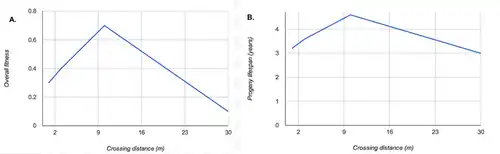
An example of inbreeding depression is shown to the right. In this case, a is the recessive allele which has negative effects. In order for the a phenotype to become active, the gene must end up as homozygous aa because in the geneotype Aa, the A takes dominance over the a and the a does not have any effect. Some recessive genes result in detrimental phenotypes by causing the organism to be less fit to its natural environment.
Another mechanism responsible for inbreeding depression is the fitness advantage of heterozygosity, which is known as overdominance. This can lead to reduced fitness of a population with many homozygous genotypes, even if they are not deleterious or recessive. Here, even the dominant alleles result in reduced fitness if present homozygously (see also hybrid vigour).
Overdominance is rare in nature.[3] For practical applications, e.g. in livestock breeding, the former is thought to be more significant – it may yield completely unviable offspring (meaning outright failure of a pedigree), while the latter can only result in relatively reduced fitness.
Natural selection
Natural selection cannot effectively remove all deleterious recessive genes from a population for several reasons. First, deleterious genes arise constantly through de novo mutation within a population. Second, most offspring will have some deleterious traits, so few will be more fit for survival than the others. Different deleterious traits are extremely unlikely to equally affect reproduction – an especially disadvantageous recessive trait expressed in a homozygous recessive individual is likely to eliminate itself, naturally limiting the expression of its phenotype. Third, recessive deleterious alleles will be "masked" by heterozygosity, and so in a dominant-recessive trait, heterozygotes will not be selected against.
When recessive deleterious alleles occur in the heterozygous state, where their potentially deleterious expression is masked by the corresponding wild-type allele, this masking phenomenon is referred to as complementation (see complementation (genetics)).
In general, sexual reproduction in eukaryotes has two fundamental aspects: genetic recombination during meiosis, and outcrossing. It has been proposed that these two aspects have two natural selective advantages respectively. A proposed adaptive advantage of meiosis is that it facilitates recombinational repair of DNA damages that are otherwise difficult to repair (see DNA repair as the adaptive advantage of meiosis). A proposed adaptive advantage of outcrossing is complementation, which is the masking of deleterious recessive alleles[4][5] (see hybrid vigor or heterosis). The selective advantage of complementation may largely account for avoidance of inbreeding (see kin recognition), though it is unlikely that animals avoid inbreeding.[6]
Management
Hybridization as a conservation effort is appropriate if the population has lost "substantial genetic variation through genetic drift and the detrimental effects of inbreeding depression are apparent" and a similar population should be used.[7][8] Different populations of the same species have different deleterious traits, and therefore their cross breeding is less likely to result in homozygosity at most loci in the offspring. This is known as outbreeding enhancement, which can be performed in extreme cases of severe inbreeding[7] by conservation managers and zoo captive breeders to prevent inbreeding depression.
However, intermixing two different populations can give rise to unfit polygenic traits in outbreeding depression (i.e. yielding offspring which lack the genetic adaptations to specific environmental conditions). These, then, will have a lowered fitness than pure-bred individuals of a particular subspecies that has adapted to its local environment.
In humans
Inbreeding may have both detrimental and beneficial effects.[9] The biological effects of inbreeding depression in humans can on occasion be confounded by socioeconomic and cultural influences on reproductive behavior.[10] Studies in human populations have shown that age at marriage, duration of marriage, illiteracy, contraceptive use, and reproductive compensation are the major determinants of apparent fertility, even amongst populations with a high proportion of consanguinous unions.[11] However, several small effects on increased mortality,[12] longer inter-birth intervals[12] and reduced overall productivity[10] have been noted in certain isolated populations, though other studies show increased fitness of offspring and no effect on lifespan past the 2nd cousin level.[13]
Charles Darwin was one of the first scientists to demonstrate the effects of inbreeding depression, through numerous experiments on plants. Darwin's wife, Emma, was his first cousin, and he was concerned about the impact of inbreeding on his ten children, three of whom died at age ten or younger; three others had childless long-term marriages.[14][15][16]
Humans do not seek to completely minimize inbreeding, but rather to maintain an optimal amount of inbreeding vs. outbreeding. Close inbreeding reduces fitness through inbreeding depression, but some inbreeding brings benefits.[17][18] Indeed, inbreeding "increases the speed of selection of beneficial recessive and co-dominant alleles, e.g. those that protect against diseases."[19]
In wolves
A small isolated highly inbred population of gray wolves in Isle Royale National Park, Michigan, USA, was considered in 2019 to be at imminent risk of extinction.[20] This gray wolf population had been experiencing severe inbreeding depression primarily due to the homozygous expression of strongly deleterious recessive mutations.[20][21] Defects arising from severe inbreeding among the wolves included reduced survival and reproduction, malformed vertebrae, syndactyly, probable cataracts, an unusual “rope tail” and anomalous fur phenotypes.[20] A separate small inbred population of gray wolves in Scandinavia was also found to suffer from inbreeding depression due to the homozygous expression of deleterious recessive mutations.[22]
Factors reducing inbreeding depression
Whilst inbreeding depression has been found to occur in almost all sufficiently studied species, some taxa, most notably some angiosperms, appear to suffer lower fitness costs than others in inbred populations.[23] Three mechanisms appear to be responsible for this: purging, differences in ploidy, and selection for heterozygosity.[23] It must be cautioned that some studies failing to show an absence of inbreeding depression in certain species can arise from small sample sizes or where the supposedly outbred control group is already suffering inbreeding depression, which frequently occurs in populations that have undergone a recent bottleneck, such as those of the naked mole rat.[23][24]
Purging selection
Purging selection occurs where the phenotypes of deleterious recessive alleles are exposed through inbreeding, and thus can be selected against. This can lead to such detrimental mutations being removed from the population, and has been demonstrated to occur rapidly where the recessive alleles have a lethal effect.[23] The efficiency of purging will depend on the relationship between the magnitude of the deleterious effect that is unmasked in the homozygotes and the importance of genetic drift, so that purging is weaker for non-lethal than for recessive lethal alleles.[25] For very small populations, drift has a strong influence, which can cause the fixation of sublethal alleles under weak selection.[23] The fixation of a single allele for a specific gene can also reduce fitness where heterozygote advantage was previously present (i.e., where heterozygous individuals have higher fitness than homozygotes of either allele), although this phenomenon seems to make a usually small contribution to inbreeding depression. Although naturally occurring, purging can be important for population survival, deliberately attempting to purge deleterious mutations from a population is not generally recommended as a technique to improve the fitness of captive bred animals.[26][27][28] In plants, genetic load can be assessed through a test analogous to an inbreeding depression test called an Autogamy depression test.
Polyploidy
Many angiosperms (flowering plants) can self-fertilise for several generations and suffer little from inbreeding depression. This is very useful for species which disperse widely and can therefore find themselves growing in a novel environment with no conspecifics present.[23] Polyploidy (having more than two paired sets of each chromosome), which is prevalent in angiosperms, ferns and a select few animal taxa, accounts for this. By having several copies of a chromosome, as opposed to two, homozygosity is less likely to occur in inbred offspring. This means that recessive deleterious alleles are not expressed as frequently as with many copies of a chromosome; it is more likely that at least one will contain a functional allele.[23]
Selection for heterozygosity
Selection for heterozygosity is rare, as lost loci undergo purifying selection for homozygous loci.[3] Inbreeding depression has also been found to occur more gradually than predicted in some wild populations, such as in the highly inbred population of Scandinavian wolves. This appears to be due to a selection pressure for more heterozygous individuals, which generally are in better condition and so are more likely to become one of the few animals to breed and produce offspring.[29]
See also
References
- Begon, Michael, Colin R. Townsend, and John L. Harper. Ecology: from individuals to ecosystems. 4th ed. Malden, MA: Blackwell Pub., 2006. Print.
- Dolgin, Elie S.; Charlesworth, Brian; Baird, Scott; Cutter, Asher D.; et al. (2007). "Inbreeding and Outbreeding Depression in Caenorhabditis Nematodes". Evolution. 61 (6): 1339–1352. doi:10.1111/j.1558-5646.2007.00118.x. PMID 17542844.
- Charlesworth, Deborah; Willis, John H. (November 2009). "The genetics of inbreeding depression". Nature Reviews Genetics. 10 (11): 783–796. doi:10.1038/nrg2664. ISSN 1471-0064. PMID 19834483. S2CID 771357.
- Bernstein, H; Byerly, HC; Hopf, FA; Michod, RE (1985). "Genetic damage, mutation, and the evolution of sex". Science. 229 (4719): 1277–1281. Bibcode:1985Sci...229.1277B. doi:10.1126/science.3898363. PMID 3898363.
- Michod, R.E. Eros and Evolution: A Natural Philosophy of Sex. (1996) Perseus Books ISBN 0201442329 ISBN 978-0201442328
- de Boer, Raïssa A.; Vega-Trejo, Regina; Kotrschal, Alexander; Fitzpatrick, John L. (July 2021). "Meta-analytic evidence that animals rarely avoid inbreeding". Nature Ecology & Evolution. 5 (7): 949–964. doi:10.1038/s41559-021-01453-9. ISSN 2397-334X. PMID 33941905. S2CID 233718913.
- Allendorf, Fred W.; Leary, Robb F.; Spruell, Paul; Wenburg, John K. (November 2001). "The problems with hybrids: setting conservation guidelines". Trends in Ecology & Evolution. 16 (11): 613–622. doi:10.1016/S0169-5347(01)02290-X.
- Edmands, Suzanne (2006-11-15). "Between a rock and a hard place: evaluating the relative risks of inbreeding and outbreeding for conservation and management: RELATIVE RISKS OF INBREEDING AND OUTBREEDING". Molecular Ecology. 16 (3): 463–475. doi:10.1111/j.1365-294X.2006.03148.x. PMID 17257106. S2CID 457825.
- Denic, Srdjan; Nicholls, Michael Gary (2007). "Genetic Benefits of Consanguinity Through Selection of Genotypes Protective Against Malaria". Human Biology. 79 (2): 145–158. doi:10.1353/hub.2007.0030. ISSN 1534-6617. PMID 18027811. S2CID 32818511.
- Robert, Alexandre; Toupance, Bruno; Tremblay, Marc; Heyer, Evelyne (2009). "Impact of inbreeding on fertility in a pre-industrial population". European Journal of Human Genetics. 17 (5): 673–681. doi:10.1038/ejhg.2008.237. PMC 2986271. PMID 19092776.
- Bittles, A. H.; Grant, J. C.; Sullivan, S. G.; Hussain, R. (2002). "Does inbreeding lead to decreased human fertility?". Annals of Human Biology. 29 (2): 111–130. doi:10.1080/03014460110075657. PMID 11874619. S2CID 31317976.
- Ober, C; Hyslop, T; Hauck, WW (January 1999). "Inbreeding effects on fertility in humans: evidence for reproductive compensation". Am. J. Hum. Genet. 64 (1): 225–31. doi:10.1086/302198. PMC 1377721. PMID 9915962.
- "Third Cousins Have Greatest Number Of Offspring, Data From Iceland Shows". ScienceDaily. Retrieved 2022-07-26.
- Berra, Tim M.; Alvarez, Gonzalo; Ceballos, Francisco C.; et al. (2010). "Was the Darwin/Wedgwood Dynasty Adversely Affected by Consanguinity?". BioScience. 60 (5): 376–383. doi:10.1525/bio.2010.60.5.7. S2CID 35915651.
- "Inbreeding May Have Caused Darwin Family Ills, Study Suggests". Science Daily.
- Clark, R.W. (1984) "The Survival of Charles Darwin" Random House [see pgs. 76 and 78]. ISBN 039452134X ISBN 978-0394521343
- Berghe, Pierre L. van den (March 1983). "Human inbreeding avoidance: Culture in nature". Behavioral and Brain Sciences. 6 (1): 91–102. doi:10.1017/S0140525X00014850. ISSN 1469-1825. S2CID 146133244.
- Alvarez, Liliana; Jaffe, Klaus (2004-01-01). "Narcissism Guides Mate Selection: Humans Mate Assortatively, as Revealed by Facial Resemblance, following an Algorithm of "Self Seeking Like"". Evolutionary Psychology. 2 (1): 147470490400200. doi:10.1177/147470490400200123. ISSN 1474-7049. S2CID 16207548.
- Denic, S.; Nagelkerke, N.; Agarwal, M. M. (2011). "On Some Novel Aspects of Consanguineous Marriages". Public Health Genomics. 14 (3): 162–168. doi:10.1159/000321771. ISSN 1662-4246. PMID 21150168. S2CID 38342159.
- Robinson JA, Räikkönen J, Vucetich LM, Vucetich JA, Peterson RO, Lohmueller KE, Wayne RK. Genomic signatures of extensive inbreeding in Isle Royale wolves, a population on the threshold of extinction. Sci Adv. 2019 May 29;5(5):eaau0757. doi: 10.1126/sciadv.aau0757. PMID: 31149628; PMCID: PMC6541468
- Kyriazis CC, Wayne RK, Lohmueller KE. Strongly deleterious mutations are a primary determinant of extinction risk due to inbreeding depression. Evol Lett. 2020 Dec 17;5(1):33-47. doi: 10.1002/evl3.209. PMID: 33552534; PMCID: PMC7857301
- Smeds L, Ellegren H. From high masked to high realized genetic load in inbred Scandinavian wolves. Mol Ecol. 2023 Apr;32(7):1567-1580. doi: 10.1111/mec.16802. Epub 2022 Dec 13. PMID: 36458895
- Frankham, Ballou, Briscoe (2002). Introduction to conservation genetics. Cambridge university press.
{{cite book}}: CS1 maint: multiple names: authors list (link) - Liberg, Paul; Firmin B. (2007). "Role of inbreeding depression and purging in captive breeding and restoration programmes". Molecular Ecology. 17 (1): 334–343. doi:10.1111/j.1365-294X.2007.03433.x. PMID 18173505. S2CID 28421723.
- García-Dorado, A (2012). "Understanding and predicting the fitness decline of shrunk populations: inbreeding, purging, mutation and standard selection". Genetics. 190 (4): 1461–1476. doi:10.1534/genetics.111.135541. PMC 3316656. PMID 22298709.
- Garcia-Dorado, A (2015). "On the consequences of ignoring purging on genetic recommendations of MVP rules". Heredity. 115 (3): 185–187. doi:10.1038/hdy.2015.28. PMC 4814235. PMID 25873145.
- Leberg, P. L.; Firmin, B. D. (2008). "Role of inbreeding depression and purging in captive breeding and restoration programmes". Molecular Ecology. 17 (1): 334–343. doi:10.1111/j.1365-294X.2007.03433.x. PMID 18173505. S2CID 28421723.
- Crnokrak, P; Barrett, SCH (2002). "Purging the genetic load: a review of the experimental evidence". Evolution. 56 (12): 2347–2358. doi:10.1111/j.0014-3820.2002.tb00160.x. PMID 12583575.
- Bensch, Staffan; Andren, Hanson; Pederson, Sand; Sejberg, Wabakken; Akesson, Liberg (2006). "Selection for heterozygosity gives hope to wild wolves". PLOS ONE. 1 (1): e72. doi:10.1371/journal.pone.0000072. PMC 1762340. PMID 17183704.