Inositol-3-phosphate synthase
In enzymology, an inositol-3-phosphate synthase (EC 5.5.1.4) is an enzyme that catalyzes the chemical reaction
- D-glucose 6-phosphate 1D-myo-inositol 3-phosphate
| inositol-3-phosphate synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
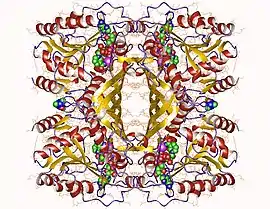 Inositol-3-phosphate synthase homotetramer, Mycobacterium tuberculosis | |||||||||
| Identifiers | |||||||||
| EC no. | 5.5.1.4 | ||||||||
| CAS no. | 9032-95-5 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Myo-inositol-1-phosphate synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
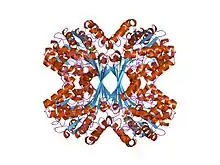 myo-inositol phosphate synthase mips from a. fulgidus | |||||||||
| Identifiers | |||||||||
| Symbol | Inos-1-P_synth | ||||||||
| Pfam | PF01658 | ||||||||
| InterPro | IPR013021 | ||||||||
| SCOP2 | 1gr0 / SCOPe / SUPFAM | ||||||||
| |||||||||
Hence, this enzyme has one substrate, D-glucose 6-phosphate, and one product, 1D-myo-inositol 3-phosphate.
This enzyme belongs to the family of isomerases, specifically the class of intramolecular lyases. The systematic name of this enzyme class is 1D-myo-inositol-3-phosphate lyase (isomerizing). Other names in common use include myo-inositol-1-phosphate synthase, D-glucose 6-phosphate cycloaldolase, inositol 1-phosphate synthatase, glucose 6-phosphate cyclase, inositol 1-phosphate synthetase, glucose-6-phosphate inositol monophosphate cycloaldolase, glucocycloaldolase, and 1L-myo-inositol-1-phosphate lyase (isomerizing).
This enzyme participates in streptomycin biosynthesis and inositol phosphate metabolism. It employs one cofactor, NAD+. The reaction this enzyme catalyses represents the first committed step in the production of all inositol-containing compounds, including phospholipids, either directly or by salvage. The enzyme exists in a cytoplasmic form in a wide range of plants, animals, and fungi. It has also been detected in several bacteria and a chloroplast form is observed in alga and higher plants. Inositol phosphates play an important role in signal transduction.
In Saccharomyces cerevisiae (Baker's yeast), the transcriptional regulation of the INO1 gene encoding inositol-3-phosphate synthase has been studied in detail and its expression is sensitive to the availability of phospholipid precursors as well as growth phase.[1] The regulation of the structural gene encoding 1L-myo-inositol-1-phosphate synthase has also been analyzed at the transcriptional level in the aquatic angiosperm, Spirodela polyrrhiza (Giant duckweed) and the halophyte, Mesembryanthemum crystallinum (Common ice plant).[2]
In prokaryotes, myo-D-inositol phosphate synthase was discovered by Bachhawat and Mande in 1999 (reported in Journal of Molecular Biology). The existence of inositol in prokaryotes is not extensive, but the discovery of this enzyme first in Mycobacterium tuberculosis, nucleated activity towards finding its inhibitors.
Structural studies
As of late 2007, 12 structures have been solved for this class of enzymes, with PDB accession codes 1GR0, 1JKF, 1JKI, 1LA2, 1P1F, 1P1H, 1P1I, 1P1J, 1P1K, 1RM0, 1VJP, and 1VKO.
References
- Klig LS, Zobel PA, Devry CG, Losberger C (June 1994). "Comparison of INO1 gene sequences and products in Candida albicans and Saccharomyces cerevisiae". Yeast. 10 (6): 789–800. doi:10.1002/yea.320100609. PMID 7975896. S2CID 21530024.
- Majumder AL, Johnson MD, Henry SA (September 1997). "1L-myo-inositol-1-phosphate synthase". Biochim. Biophys. Acta. 1348 (1–2): 245–56. doi:10.1016/s0005-2760(97)00122-7. PMID 9370339.
Further reading
- Eisenberg F Jr (1967). "D-myoinositol 1-phosphate as product of cyclization of glucose 6-phosphate and substrate for a specific phosphatase in rat testis". J. Biol. Chem. 242 (7): 1375–82. PMID 4290245.
- Sherman WR, Stewart MA, Zinbo M (1969). "Mass spectrometric study on the mechanism of D-glucose 6-phosphate-L-myo-inositol 1-phosphate cyclase". J. Biol. Chem. 244 (20): 5703–8. PMID 4310603.
- Barnett JE, Corina DL (1968). "The mechanism of glucose 6-phosphate-D-myo-inositol 1-phosphate cyclase of rat testis. The involvement of hydrogen atoms". Biochem. J. 108 (1): 125–9. doi:10.1042/bj1080125. PMC 1198777. PMID 4297937.
- Barnett JE, Rasheed A, Corina DL (1973). "Partial reactions of D-glucose 6-phosphate-1L-myoinositiol 1-phosphate cyclase". Biochem. J. 131 (1): 21–30. doi:10.1042/bj1310021. PMC 1177435. PMID 4352864.
Bachhawat N and Mande SC (1999) J. Mol. Biol. Identification of the INO1 gene of Mycobacterium tuberculosis H37Rv reveals a novel class of inositol-1-phosphate synthase enzyme. 291, 531–536.