Junctional adhesion molecule
A junctional adhesion molecule (JAM) is a protein that is a member of the immunoglobulin superfamily,[1][2] and is expressed in a variety of different tissues, such as leukocytes, platelets, and epithelial and endothelial cells.[2] They have been shown to regulate signal complex assembly on both their cytoplasmic and extracellular domains through interaction with scaffolding that contains a PDZ domain and adjacent cell's receptors, respectively.[3] JAMs adhere to adjacent cells through interactions with integrins LFA-1 and Mac-1, which are contained in leukocyte β2 and α4β1, which is contained in β1. JAMs have many influences on leukocyte-endothelial cell interactions, which are primarily moderated by the integrins discussed above.[4] They interact in their cytoplasmic domain with scaffold proteins that contain a PDZ domain, which are common protein interaction modules that target short amino acid sequences at the C-terminus of proteins, to form tight junctions in both epithelial and endothelial cells as polarity is gained in the cell.[3]
| Junctional Adhesion Molecule | |
|---|---|
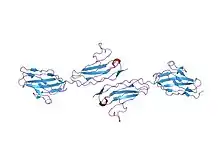 Crystallographic structure of junctional adhesion molecules linked together | |
| Identifiers | |
| Symbol | JAM |
| Membranome | Immunoglobulinset domain V set domain |
Structure
JAMs are usually around 40 kDa in size.[5] Based on crystallographic studies conducted with recombinant extracellular mouse JAMs (rsJAM) and human JAMs (hJAM), it has been shown that JAM consists of immunoglobulin-like V-set domain followed by a second immunoglobulin domain that are linked together by a short linker sequence. The linker makes extensive hydrogen bonds to both domains, and the side chain of one of the main linker residues, Leu128, is commonly embedded in a hydrophobic cleft between each immunoglobulin-like domain.[6] Two JAM molecules contain N-terminal domains that react in a highly complementary fashion due to prolific ionic and hydrophobic interactions. These two molecules form U-shaped dimers and salt bridges are then formed by a R(V,I,L)E motif.[6] This motif has been proven to be important in dimer formation and is common among different types of JAMs. It commonly consists of Arg58-Val59-Glu60 located on the N-terminus and can dissociate into monomers based on the conditions of the solution it is exposed to. This motif has been shown to be present in many common variants of JAMs, including rsJAM, hJAM, JAM-1, JAM-2, and JAM-3.[7]
Types
Three major JAM molecules interact with various molecules and receptors within the body:
JAM-1
JAM-1 was the first of the junctional adhesion molecules to be discovered, and is located in the tight junctions of both epithelial and endothelial cells.[8] JAM-1interacts with cells in a homophilic manner in order to preserve the structure of the junction while moderating its permeability. It can also interact with receptors as a heterophilic structure by acting as a ligand for LFA-1 and facilitating leukocyte transmigration.[8] JAM-1 also plays a significant role in many different cellular functions, including being both a reovirus receptor and a platelet receptor.[8]
JAM-2
Like JAM-1, JAM-2 also is a member of the immunoglobulin superfamily.[9] JAM-2 localization is moderated by serine phosphorylation at tight junctions as the molecule adheres to other tight junction proteins like PAR-3 and ZO-1. JAM-2 has been shown to interact with these proteins, primarily through the PDZ1 domain, and also through the PDZ3 domain.[10] JAM-2 has also shown to act as a ligand for many immune cells, and plays a role in lymphocyte attraction to specific organs.[10]
JAM-3
JAM-3 functions similarly to JAM-2 as it is localized around the tight junctions of epithelial and endothelial cells, but has been shown to be unable to adhere to leukocytes in the manner that other JAMs can.[11] Mutations of JAM-3 introns have been shown to lead to brain hemorrhages and development of cataracts.[11] Like JAM-2, JAM-3 has been shown associate with tight junction proteins like PAR-3 and ZO-1. JAM-3 has also been shown to interact with PARD3 (partitioning defective 3 homolog).
Function
JAMs serve many different functions within the cell:
Cell motility
JAMs play a critical role in the regulation of cell movement in multiple different cell types, such as epithelial, endothelial, leukocyte, and germ cells.[10] JAM-1 regulates motility in epithelial cells by moderating expression of β1 integrin protein downstream of Rap1. JAM-1 has been shown to be able to cause cell adhesion, spreading and movement along β1 ligands, like collagen IV and fibronectin.[3] JAM-1 also acts to moderate migration of vitronectin in endothelial cells. Vitronectin is a ligand for integrins αvβ3 and αvβ5, which exhibit selective cooperativity with bFGF and VEGF in the activation of the MAPK pathway. JAM-1 and JAM-3 allow leukocytes to migrate into connective tissue by freeing polymorphonuclear leukocytes from entrapment in endothelial cells and basement membranes.[3] In the absence of JAM-1, these leukocytes cannot moderate β1 integrin endocytosis, and cannot be effectively expressed on the surface of the cell (which is essential for motility).[11]
Cell polarity
JAM-1 and JAM-3 have significant roles in regulating cell polarity through their interactions with cell polarity proteins.[5] JAM-1, JAM-2, and JAM-3 all interact with PAR-3 to influence cell polarity. PAR-3 is a significant factor in a cell's polarity-regulating complex, and regulates polarity in different cell types in many different organisms.[12] All components of the PAR complex are required for tight junction formation between cells, but premature adherens junctions can form without PAR complex components being present.[3] However, these junctions cannot efficiently develop into mature epithelial cell junctions. JAM-3 has also shown to affect cell polarity in spermatids by regulating the localization of cytosolic polarity.[10]
Cell Proliferation
In order to preserve homeostasis of adult tissue, aged cells must be replaced with new cells at varying frequency, depending on the organ. Some organs that require high rates of cellular turnover are the small intestine and the colon. JAM-1 has been shown to regulate the proliferation of cells in the colon.[8] In JAM-1 deficient mice, it has been found that the amount of proliferating cells in the colon greatly increased due to the increased proliferation of TA cells. JAM-1 acts to suppress cell proliferation, which is performed by restricting Akt activity.[8] Recent studies have also pointed to JAM-1 preserving structural integrity of tissues more so than regulating cell number.
Role in physiological processes
JAMs play a significant role in many diverse physiological processes within the human body, including:
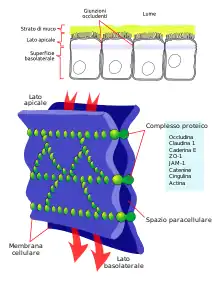
Tight junction formation
Tight junctions serve to provide most of the function for the barrier that is present on epithelial cell surfaces. Tight junctions feature the localization of both JAM-1 and JAM-3, and JAM-3 is localized exclusively at tight junctions.[3] The role of JAM-1 in tight junction biology is to function through mediation partly due to the localization of the Par-αPKC complex at adherens junctions during junction creation.[3] Once the tight junction is formed, many JAM-1 proteins are present, many of which are now phosphorylated at Ser285.[3] JAM-1 also regulates the activity of many different claudins within different epithelial cells.[7]
Angiogenesis
Angiogenesis is the generation of blood vessels from old blood vessels. Studies have shown that proteins found in tight junctions serve as intermediaries that moderate angiogenic signaling pathways. JAM-1 induces proliferation of endothelial cells, which begins the process of angiogenesis.[13] An analysis of JAM-1 showed a correlation between JAM-1 activity and FGF2-induced angiogenesis in both cancerous proliferation or vascular repair.[13]
Male fertility
JAM-3 has been shown to be a primary regulator of the development of spermatids as well as the rest of the male reproductive system. Within the Sertoli cells of the male reproductive system, JAM-3 interacts with JAM-2 to influence the polarity of both round and elongated spermatids.[12] JAM-1 and JAM-2 are also present in and contribute to the polarity of the blood-testis barrier. Studies have also shown that inactivation of JAM-3 has been shown to significantly impede fertility by blocking male germ cell development and proliferation.[3]
References
- Greene C, Campbell M, Janigro D (January 2019). "Chapter 1 - Fundamentals of Brain–Barrier Anatomy and Global Functions". In Lonser RR, Sarntinoranont M, Bankiewicz K (eds.). Nervous System Drug Delivery. Academic Press. pp. 3–20. doi:10.1016/b978-0-12-813997-4.00001-3. ISBN 978-0-12-813997-4. S2CID 198273920.
- Ebnet K, Suzuki A, Ohno S, Vestweber D (January 2004). "Junctional adhesion molecules (JAMs): more molecules with dual functions?". Journal of Cell Science. 117 (Pt 1): 19–29. doi:10.1242/jcs.00930. PMID 14657270.
- Ebnet K (October 2017). "Junctional Adhesion Molecules (JAMs): Cell Adhesion Receptors With Pleiotropic Functions in Cell Physiology and Development". Physiological Reviews. 97 (4): 1529–1554. doi:10.1152/physrev.00004.2017. PMID 28931565. S2CID 10846721.
- Lee HJ, Zheng JJ (May 2010). "PDZ domains and their binding partners: structure, specificity, and modification". Cell Communication and Signaling. 8 (1): 8. doi:10.1186/1478-811X-8-8. PMC 2891790. PMID 20509869.
- Bauer HC, Traweger A, Bauer H (2004-01-01). "Proteins of the Tight Junction in the Blood-Brain Barrier". In HS, Westman J (eds.). 1 - Proteins of the Tight Junction in the Blood-Brain Barrier. pp. 1–10. doi:10.1016/b978-012639011-7/50005-x. ISBN 978-0-12-639011-7.
{{cite book}}:|work=ignored (help) - Kostrewa D, Brockhaus M, D'Arcy A, Dale GE, Nelboeck P, Schmid G, et al. (August 2001). "X-ray structure of junctional adhesion molecule: structural basis for homophilic adhesion via a novel dimerization motif". The EMBO Journal. 20 (16): 4391–8. doi:10.1093/emboj/20.16.4391. PMC 125582. PMID 11500366.
- Prota AE, Campbell JA, Schelling P, Forrest JC, Watson MJ, Peters TR, et al. (April 2003). "Crystal structure of human junctional adhesion molecule 1: implications for reovirus binding". Proceedings of the National Academy of Sciences of the United States of America. 100 (9): 5366–71. Bibcode:2003PNAS..100.5366P. doi:10.1073/pnas.0937718100. PMC 404559. PMID 12697893.
- Naik UP, Eckfeld K (2003). "Junctional adhesion molecule 1 (JAM-1)". Journal of Biological Regulators and Homeostatic Agents. 17 (4): 341–7. PMID 15065765.
- "JAM2 junctional adhesion molecule 2 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2019-11-29.
- Ebnet K, Aurrand-Lions M, Kuhn A, Kiefer F, Butz S, Zander K, et al. (October 2003). "The junctional adhesion molecule (JAM) family members JAM-2 and JAM-3 associate with the cell polarity protein PAR-3: a possible role for JAMs in endothelial cell polarity". Journal of Cell Science. 116 (Pt 19): 3879–91. doi:10.1242/jcs.00704. PMID 12953056.
- "JAM3 junctional adhesion molecule 3 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2019-11-29.
- Rehder D, Iden S, Nasdala I, Wegener J, Brickwedde MK, Vestweber D, Ebnet K (October 2006). "Junctional adhesion molecule-a participates in the formation of apico-basal polarity through different domains". Experimental Cell Research. 312 (17): 3389–403. doi:10.1016/j.yexcr.2006.07.004. PMID 16919624.
- Naik TU, Naik MU, Naik UP (January 2008). "Junctional adhesion molecules in angiogenesis". Frontiers in Bioscience. 13 (13): 258–62. doi:10.2741/2676. PMID 17981544. S2CID 11562413.