LTR retrotransposon
LTR retrotransposons are class I transposable element characterized by the presence of long terminal repeats (LTRs) directly flanking an internal coding region. As retrotransposons, they mobilize through reverse transcription of their mRNA and integration of the newly created cDNA into another location. Their mechanism of retrotransposition is shared with retroviruses, with the difference that most LTR-retrotransposons do not form infectious particles that leave the cells and therefore only replicate inside their genome of origin. Those that do (occasionally) form virus-like particles are classified under Ortervirales.
Their size ranges from a few hundred base pairs to 25kb, for example the Ogre retrotransposon in the pea genome.
In plant genomes, LTR retrotransposons are the major repetitive sequence class, for example, constituting more than 75% of the maize genome.[1] LTR retrotransposons make up about 8% of the human genome and approximately 10% of the mouse genome.[2]
Structure and propagation
LTR retrotransposons have direct long terminal repeats that range from ~100 bp to over 5 kb in size. LTR retrotransposons are further sub-classified into the Ty1-copia-like (Pseudoviridae), Ty3-like (Metaviridae, formally referred to as Gypsy-like, a name that is being considered for retirement[3]), and BEL-Pao-like (Belpaoviridae) groups based on both their degree of sequence similarity and the order of encoded gene products. Ty1-copia and Ty3-Metaviridae groups of retrotransposons are commonly found in high copy number (up to a few million copies per haploid nucleus) in animals, fungi, protista, and plants genomes. BEL-Pao like elements have so far only been found in animals.[4][5]
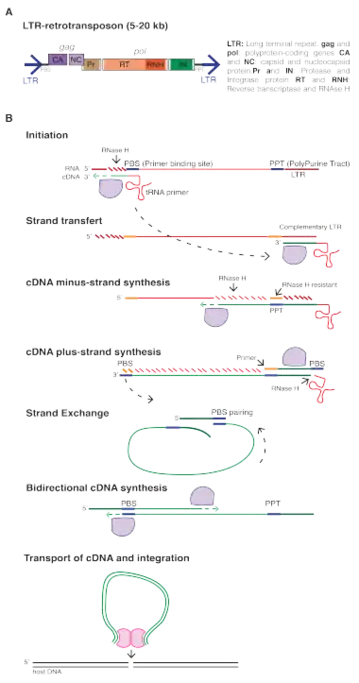
All functional LTR-retrotransposons encode a minimum of two genes, gag and pol, that are sufficient for their replication. Gag encodes a polyprotein with a capsid and a nucleocapsid domain.[6] Gag proteins form virus-like particles in the cytoplasm inside which reverse-transcription occurs. The Pol gene produces three proteins: a protease (PR), a reverse transcriptase endowed with an RT (reverse-transcriptase) and an RNAse H domains, and an integrase (IN).[7]
Typically, LTR-retrotransposon mRNAs are produced by the host RNA pol II acting on a promoter located in their 5’ LTR. The Gag and Pol genes are encoded in the same mRNA. Depending on the host species, two different strategies can be used to express the two polyproteins: a fusion into a single open reading frame (ORF) that is then cleaved or the introduction of a frameshift between the two ORFs.[8] Occasional ribosomal frameshifting allows the production of both proteins, while ensuring that much more Gag protein is produced to form virus-like particles.
Reverse transcription usually initiates at a short sequence located immediately downstream of the 5’-LTR and termed the primer binding site (PBS). Specific host tRNAs bind to the PBS and act as primers for reverse-transcription, which occurs in a complex and multi-step process, ultimately producing a double- stranded cDNA molecule. The cDNA is finally integrated into a new location, creating short TSDs (Target Site Duplications) [9] and adding a new copy in the host genome
Types
Ty1-copia retrotransposons
Ty1-copia retrotransposons are abundant in species ranging from single-cell algae to bryophytes, gymnosperms, and angiosperms. They encode four protein domains in the following order: protease, integrase, reverse transcriptase, and ribonuclease H.
At least two classification systems exist for the subdivision of Ty1-copia retrotransposons into five lineages:[10][11] Sireviruses/Maximus, Oryco/Ivana, Retrofit/Ale, TORK (subdivided in Angela/Sto, TAR/Fourf, GMR/Tork), and Bianca.
Sireviruses/Maximus retrotransposons contain an additional putative envelope gene. This lineage is named for the founder element SIRE1 in the Glycine max genome,[12] and was later described in many species such as Zea mays,[13] Arabidopsis thaliana,[14] Beta vulgaris,[15] and Pinus pinaster.[16] Plant Sireviruses of many sequenced plant genomes are summarized at the MASIVEdb Sirevirus database.[17]
Ty3-retrotransposons (formally gypsy)
Ty3-retrotransposons are widely distributed in the plant kingdom, including both gymnosperms and angiosperms. They encode at least four protein domains in the order: protease, reverse transcriptase, ribonuclease H, and integrase. Based on structure, presence/absence of specific protein domains, and conserved protein sequence motifs, they can be subdivided into several lineages:
Errantiviruses contain an additional defective envelope ORF with similarities to the retroviral envelope gene. First described as Athila-elements in Arabidopsis thaliana,[18][19] they have been later identified in many species, such as Glycine max[20] and Beta vulgaris.[21]
Chromoviruses contain an additional chromodomain (chromatin organization modifier domain) at the C-terminus of their integrase protein.[22][23] They are widespread in plants and fungi, probably retaining protein domains during evolution of these two kingdoms.[24] It is thought that the chromodomain directs retrotransposon integration to specific target sites.[25] According to sequence and structure of the chromodomain, chromoviruses are subdivided into the four clades CRM, Tekay, Reina and Galadriel. Chromoviruses from each clade show distinctive integration patterns, e.g. into centromeres or into the rRNA genes.[26][27]
Ogre-elements are gigantic Ty3-retrotransposons reaching lengths up to 25 kb.[28] Ogre elements have been first described in Pisum sativum.[29]
Metaviruses describe conventional Ty3-gypsy retrotransposons that do not contain additional domains or ORFs.
The Sushi family of Ty3 long terminal repeat retrotransposons were first identified in teleost fish and Sushi-like neogenes were subsequently identified in mammals.[30] Mammalian retrotransposon-derived transcripts (MARTs) cannot transpose but have retained open reading frames, demonstrate high levels of evolutionary conservation and are subject to selective pressures, which suggests some have become neofunctionalized genes with new cellular functions.[31] Retrotransposon gag-like-3 (RTL3/ZCCHC5/MART3) is one of eleven Sushi-like neogenes identified in the human genome.[32]
BEL/pao family
The BEL/pao family is found in animals.[33]
Endogenous retroviruses (ERV)
Although retroviruses are often classified separately, they share many features with LTR retrotransposons. A major difference with Ty1-copia and Ty3-gypsy retrotransposons is that retroviruses have an envelope protein (ENV). A retrovirus can be transformed into an LTR retrotransposon through inactivation or deletion of the domains that enable extracellular mobility. If such a retrovirus infects and subsequently inserts itself in the genome in germ line cells, it may become transmitted vertically and become an Endogenous Retrovirus.[5]
Terminal repeat retrotransposons in miniature (TRIMs)
Some LTR retrotransposons lack all of their coding domains. Due to their short size, they are referred to as terminal repeat retrotransposons in miniature (TRIMs).[34][35] Nevertheless, TRIMs can be able to retrotranspose, as they may rely on the coding domains of autonomous Ty1-copia or Ty3-gypsy retrotransposons. Among the TRIMs, the Cassandra family plays an exceptional role, as the family is unusually wide-spread among higher plants.[36] In contrast to all other characterized TRIMs, Cassandra elements harbor a 5S rRNA promoter in their LTR sequence.[37] Due to their short overall length and the relatively high contribution of the flanking LTRs, TRIMs are prone to re-arrangements by recombination.[38]
References
- Baucom, RS; Estill, JC; Chaparro, C; Upshaw, N; Jogi, A; Deragon, JM; Westerman, RP; Sanmiguel, PJ; Bennetzen, JL (November 2009). "Exceptional diversity, non-random distribution, and rapid evolution of retroelements in the B73 maize genome". PLOS Genetics. 5 (11): e1000732. doi:10.1371/journal.pgen.1000732. PMC 2774510. PMID 19936065.
- McCarthy EM, McDonald JF (2004). "Long terminal repeat retrotransposons of Mus musculus". Genome Biol. 5 (3): R14. doi:10.1186/gb-2004-5-3-r14. PMC 395764. PMID 15003117.
- osf.io https://osf.io/fma57/. Retrieved 2022-11-04.
{{cite web}}: Missing or empty|title=(help) - Copeland CS, Mann VH, Morales ME, Kalinna BH, Brindley PJ (2005). "The Sinbad retrotransposon from the genome of the human blood fluke, Schistosoma mansoni, and the distribution of related Pao-like elements". BMC Evol. Biol. 5 (1): 20. doi:10.1186/1471-2148-5-20. PMC 554778. PMID 15725362.
- Wicker T, Sabot F, Hua-Van A, et al. (December 2007). "A unified classification system for eukaryotic transposable elements". Nat. Rev. Genet. 8 (12): 973–82. doi:10.1038/nrg2165. PMID 17984973.
- Sandmeyer, Suzanne B; Clemens, Kristina A (2010). "Function of a retrotransposon nucleocapsid protein". RNA Biology. 7 (6): 642–654. doi:10.4161/rna.7.6.14117. ISSN 1547-6286. PMC 3073325. PMID 21189452.
- Wicker, Thomas; Sabot, François; Hua-Van, Aurélie; Bennetzen, Jeffrey L.; Capy, Pierre; Chalhoub, Boulos; Flavell, Andrew; Leroy, Philippe; Morgante, Michele (December 2007). "A unified classification system for eukaryotic transposable elements". Nature Reviews. Genetics. 8 (12): 973–982. doi:10.1038/nrg2165. ISSN 1471-0064. PMID 17984973.
- GAO, XIANG; HAVECKER, ERICKA R.; BARANOV, PAVEL V.; ATKINS, JOHN F.; VOYTAS, DANIEL F. (December 2003). "Translational recoding signals between gag and pol in diverse LTR retrotransposons". RNA. 9 (12): 1422–1430. doi:10.1261/rna.5105503. ISSN 1355-8382. PMC 1370496. PMID 14623998.
- "What is Transposon's Target Site Duplication (TSD)?". 22 May 2019.
- Wicker, T; Keller, B (July 2007). "Genome-wide comparative analysis of copia retrotransposons in Triticeae, rice, and Arabidopsis reveals conserved ancient evolutionary lineages and distinct dynamics of individual copia families". Genome Research. 17 (7): 1072–81. doi:10.1101/gr.6214107. PMC 1899118. PMID 17556529.
- Llorens, C; Muñoz-Pomer, A; Bernad, L; Botella, H; Moya, A (2 November 2009). "Network dynamics of eukaryotic LTR retroelements beyond phylogenetic trees". Biology Direct. 4: 41. doi:10.1186/1745-6150-4-41. PMC 2774666. PMID 19883502.
- Laten, HM; Majumdar, A; Gaucher, EA (9 June 1998). "SIRE-1, a copia/Ty1-like retroelement from soybean, encodes a retroviral envelope-like protein". Proceedings of the National Academy of Sciences of the United States of America. 95 (12): 6897–902. Bibcode:1998PNAS...95.6897L. doi:10.1073/pnas.95.12.6897. PMC 22677. PMID 9618510.
- Bousios, A; Kourmpetis, YA; Pavlidis, P; Minga, E; Tsaftaris, A; Darzentas, N (February 2012). "The turbulent life of Sirevirus retrotransposons and the evolution of the maize genome: more than ten thousand elements tell the story". The Plant Journal. 69 (3): 475–88. doi:10.1111/j.1365-313x.2011.04806.x. PMID 21967390.
- Kapitonov, VV; Jurka, J (1999). "Molecular paleontology of transposable elements from Arabidopsis thaliana". Genetica. 107 (1–3): 27–37. doi:10.1023/a:1004030922447. PMID 10952195. S2CID 27249664.
- Weber, B; Wenke, T; Frömmel, U; Schmidt, T; Heitkam, T (February 2010). "The Ty1-copia families SALIRE and Cotzilla populating the Beta vulgaris genome show remarkable differences in abundance, chromosomal distribution, and age". Chromosome Research. 18 (2): 247–63. doi:10.1007/s10577-009-9104-4. PMID 20039119. S2CID 24883110.
- Miguel, C; Simões, M; Oliveira, MM; Rocheta, M (November 2008). "Envelope-like retrotransposons in the plant kingdom: evidence of their presence in gymnosperms (Pinus pinaster)". Journal of Molecular Evolution. 67 (5): 517–25. Bibcode:2008JMolE..67..517M. doi:10.1007/s00239-008-9168-3. PMID 18925379. S2CID 28629941.
- Bousios, A; Minga, E; Kalitsou, N; Pantermali, M; Tsaballa, A; Darzentas, N (30 April 2012). "MASiVEdb: the Sirevirus Plant Retrotransposon Database". BMC Genomics. 13: 158. doi:10.1186/1471-2164-13-158. PMC 3414828. PMID 22545773.
- Pélissier, T; Tutois, S; Deragon, JM; Tourmente, S; Genestier, S; Picard, G (November 1995). "Athila, a new retroelement from Arabidopsis thaliana". Plant Molecular Biology. 29 (3): 441–52. doi:10.1007/bf00020976. PMID 8534844. S2CID 8296300.
- Wright, DA; Voytas, DF (June 1998). "Potential retroviruses in plants: Tat1 is related to a group of Arabidopsis thaliana Ty3/gypsy retrotransposons that encode envelope-like proteins". Genetics. 149 (2): 703–15. doi:10.1093/genetics/149.2.703. PMC 1460185. PMID 9611185.
- Wright, DA; Voytas, DF (January 2002). "Athila4 of Arabidopsis and Calypso of soybean define a lineage of endogenous plant retroviruses". Genome Research. 12 (1): 122–31. doi:10.1101/gr.196001. PMC 155253. PMID 11779837.
- Wollrab, C; Heitkam, T; Holtgräwe, D; Weisshaar, B; Minoche, AE; Dohm, JC; Himmelbauer, H; Schmidt, T (November 2012). "Evolutionary reshuffling in the Errantivirus lineage Elbe within the Beta vulgaris genome". The Plant Journal. 72 (4): 636–51. doi:10.1111/j.1365-313x.2012.05107.x. PMID 22804913.
- Marín, I; Lloréns, C (July 2000). "Ty3/Gypsy retrotransposons: description of new Arabidopsis thaliana elements and evolutionary perspectives derived from comparative genomic data". Molecular Biology and Evolution. 17 (7): 1040–9. doi:10.1093/oxfordjournals.molbev.a026385. PMID 10889217.
- Gorinsek, B; Gubensek, F; Kordis, D (May 2004). "Evolutionary genomics of chromoviruses in eukaryotes". Molecular Biology and Evolution. 21 (5): 781–98. doi:10.1093/molbev/msh057. PMID 14739248.
- Novikova, O; Smyshlyaev, G; Blinov, A (8 April 2010). "Evolutionary genomics revealed interkingdom distribution of Tcn1-like chromodomain-containing Gypsy LTR retrotransposons among fungi and plants". BMC Genomics. 11: 231. doi:10.1186/1471-2164-11-231. PMC 2864245. PMID 20377908.
- Gao, X; Hou, Y; Ebina, H; Levin, HL; Voytas, DF (March 2008). "Chromodomains direct integration of retrotransposons to heterochromatin". Genome Research. 18 (3): 359–69. doi:10.1101/gr.7146408. PMC 2259100. PMID 18256242.
- Neumann, P; Navrátilová, A; Koblížková, A; Kejnovský, E; Hřibová, E; Hobza, R; Widmer, A; Doležel, J; Macas, J (3 March 2011). "Plant centromeric retrotransposons: a structural and cytogenetic perspective". Mobile DNA. 2 (1): 4. doi:10.1186/1759-8753-2-4. PMC 3059260. PMID 21371312.
- Weber, B; Heitkam, T; Holtgräwe, D; Weisshaar, B; Minoche, AE; Dohm, JC; Himmelbauer, H; Schmidt, T (1 March 2013). "Highly diverse chromoviruses of Beta vulgaris are classified by chromodomains and chromosomal integration". Mobile DNA. 4 (1): 8. doi:10.1186/1759-8753-4-8. PMC 3605345. PMID 23448600.
- Macas, J; Neumann, P (1 April 2007). "Ogre elements--a distinct group of plant Ty3/gypsy-like retrotransposons". Gene. 390 (1–2): 108–16. doi:10.1016/j.gene.2006.08.007. PMID 17052864.
- Neumann, P; Pozárková, D; Macas, J (October 2003). "Highly abundant pea LTR retrotransposon Ogre is constitutively transcribed and partially spliced". Plant Molecular Biology. 53 (3): 399–410. CiteSeerX 10.1.1.551.7542. doi:10.1023/b:plan.0000006945.77043.ce. PMID 14750527. S2CID 13412101.
- Ball, Hope C.; Ansari, Mohammad Y.; Ahmad, Nashrah; Novak, Kimberly; Haqqi, Tariq M. (November 2021). "A retrotransposon gag-like-3 gene RTL3 and SOX-9 co-regulate the expression of COL2A1 in chondrocytes". Connective Tissue Research. 62 (6): 615–628. doi:10.1080/03008207.2020.1828380. ISSN 1607-8438. PMC 8404968. PMID 33043724.
- Ball, Hope C.; Ansari, Mohammad Y.; Ahmad, Nashrah; Novak, Kimberly; Haqqi, Tariq M. (November 2021). "A retrotransposon gag-like-3 gene RTL3 and SOX-9 co-regulate the expression of COL2A1 in chondrocytes". Connective Tissue Research. 62 (6): 615–628. doi:10.1080/03008207.2020.1828380. ISSN 1607-8438. PMC 8404968. PMID 33043724.
- Ball, Hope C.; Ansari, Mohammad Y.; Ahmad, Nashrah; Novak, Kimberly; Haqqi, Tariq M. (November 2021). "A retrotransposon gag-like-3 gene RTL3 and SOX-9 co-regulate the expression of COL2A1 in chondrocytes". Connective Tissue Research. 62 (6): 615–628. doi:10.1080/03008207.2020.1828380. ISSN 1607-8438. PMC 8404968. PMID 33043724.
- Llorens, Carlos; Soriano, Beatriz; Navarrete-Muñoz, Maria A.; Hafez, Ahmed; Arnau, Vicente; Benito, Jose Miguel; Gabaldon, Toni; Rallon, Norma; Pérez-Sánchez, Jaume; Krupovic, Mart (2021). "Reverse-Transcribing Viruses (Belpaoviridae, Metaviridae, and Pseudoviridae)". Encyclopedia of virology (Fourth ed.). Amsterdam. pp. 653–666. doi:10.1016/B978-0-12-809633-8.21514-8. ISBN 978-0-12-814516-6. S2CID 243337524.
{{cite book}}: CS1 maint: location missing publisher (link) - Witte, Claus-Peter Le, Quang Hien Bureau, Thomas Kumar, Amar (2001). "Terminal-repeat retrotransposons in miniature (TRIM) are involved in restructuring plant genomes". Proceedings of the National Academy of Sciences of the United States of America. The National Academy of Sciences. 98 (24): 13778–13783. Bibcode:2001PNAS...9813778W. doi:10.1073/pnas.241341898. OCLC 678730241. PMC 61118. PMID 11717436.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Antonius-Klemola, Kristiina; Kalendar, Ruslan; Schulman, Alan H. (2006-01-11). "TRIM retrotransposons occur in apple and are polymorphic between varieties but not sports". Theoretical and Applied Genetics. 112 (6): 999–1008. doi:10.1007/s00122-005-0203-0. ISSN 0040-5752. PMID 16404583. S2CID 7035882.
- Gao, Dongying; Li, Yupeng; Abernathy, Brian; Jackson, Scott (2014-10-29). "Landscape and evolutionary dynamics of terminal-repeat retrotransposons in miniature (TRIMs) in 48 whole plant genomes". dx.doi.org. doi:10.1101/010850. S2CID 10238180. Retrieved 2020-10-03.
- Kalendar, R.; Tanskanen, J.; Chang, W.; Antonius, K.; Sela, H.; Peleg, O.; Schulman, A. H. (2008-04-11). "Cassandra retrotransposons carry independently transcribed 5S RNA". Proceedings of the National Academy of Sciences. 105 (15): 5833–5838. Bibcode:2008PNAS..105.5833K. doi:10.1073/pnas.0709698105. ISSN 0027-8424. PMC 2311327. PMID 18408163.
- Maiwald, Sophie; Weber, Beatrice; Seibt, Kathrin M; Schmidt, Thomas; Heitkam, Tony (2020-10-03). "The Cassandra retrotransposon landscape in sugar beet (Beta vulgaris) and related Amaranthaceae: Recombination and re-shuffling lead to a high structural variability". Annals of Botany. 127 (1): 91–109. doi:10.1093/aob/mcaa176. ISSN 0305-7364. PMC 7750724. PMID 33009553.