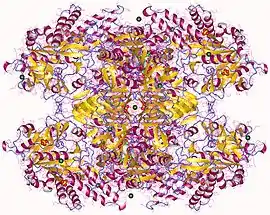
UTP—glucose-1-phosphate uridylyltransferase
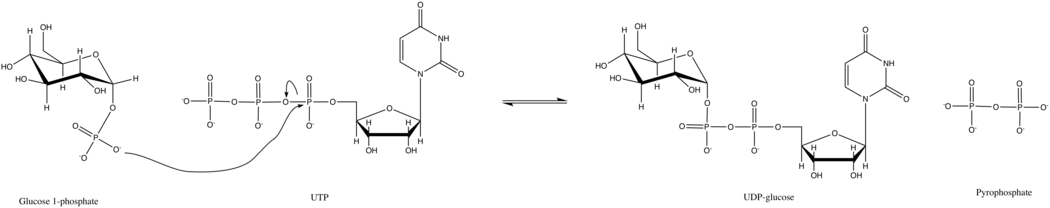
UTP—glucose-1-phosphate uridylyltransferase also known as glucose-1-phosphate uridylyltransferase (or UDP–glucose pyrophosphorylase) is an enzyme involved in carbohydrate metabolism. It synthesizes UDP-glucose from glucose-1-phosphate and UTP; i.e.,
| UTP—glucose-1-phosphate uridylyltransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Human UTP—glucose-1-phosphate uridylyltransferase cartoon created in pymol | |||||||||
| Identifiers | |||||||||
| EC no. | 2.7.7.9 | ||||||||
| CAS no. | 9026-22-6 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
UTP—glucose-1-phosphate uridylyltransferase is an enzyme found in all three domains (bacteria, eukarya, and archaea) as it is a key player in glycogenesis and cell wall synthesis. Its role in sugar metabolism has been studied extensively in plants in order to understand plant growth and increase agricultural production. Recently, human UTP—glucose-1-phosphate uridylyltransferase has been studied and crystallized, revealing a different type of regulation than other organisms previously studied. Its significance is derived from the many uses of UDP-glucose including galactose metabolism, glycogen synthesis, glycoprotein synthesis, and glycolipid synthesis.[1][2]
Structure
The structure of UTP—glucose-1-phosphate uridylyltransferase is significantly different between prokaryotes and eukaryotes, but within eukaryotes, the primary, secondary, and tertiary structures of the enzyme are quite conserved.[3] In many species, UTP—glucose-1-phosphate uridylyltransferase is found as a homopolymer consisting of identical subunits in a symmetrical quaternary structure.[4][5] The number of subunits varies across species: for instance, in Escherichia coli, the enzyme is found as a tetramer, whereas in Burkholderia xenovorans, the enzyme is dimeric.[5][6] In humans and in yeast, the enzyme is active as an octamer consisting of two tetramers stacked onto one another with conserved hydrophobic residues at the interfaces between the subunits.[7][8] In contrast, the enzyme in plants has conserved charged residues forming the interface between subunits.
In humans, each enzyme subunit contains several residues (L113, N251, and N328) that are highly conserved in eukaryotes. A Rossman fold motif participates in binding of the UTP nucleotide and a sugar-binding domain (residues T286–G293) coordinates with the glucose ring.[9] A missense mutation (G115D) in the region of the enzyme containing the active site (which is conserved in eukaryotes) causes a dramatic decrease in enzymatic activity in vitro.[10]
 Crystal structure of UTP—glucose-1-phosphate uridylyltransferase from Burkholderia xenovorans
Crystal structure of UTP—glucose-1-phosphate uridylyltransferase from Burkholderia xenovorans Human UTP—glucose-1-phosphate uridylyltransferase isoform 1 subunit with UDP-glucose bound
Human UTP—glucose-1-phosphate uridylyltransferase isoform 1 subunit with UDP-glucose bound
Examples
Human genes encoding proteins with UTP—glucose-1-phosphate uridylyltransferase activity include two isoforms with molecular weights of 56.9 and 55.7 kDa, respectively.[11]
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
Function
UTP—glucose-1-phosphate uridylyltransferase is ubiquitous in nature due to its important role in the generation of UDP-glucose, a central compound in carbohydrate metabolism. In plant leaves, UTP—glucose-1-phosphate uridylyltransferase is a key part of the sucrose biosynthesis pathway, supplying Uridine diphosphate glucose to Sucrose-phosphate synthase which converts UDP-glucose and D-fructose 6-phosphate into sucrose-6-phosphate.[12] It may also be partially responsible for the breakdown of sucrose in other tissues using UDP-glucose.
In higher animals, the enzyme is highly active in tissues involved in glycogenesis, including the liver and the muscles.[13] An exception is the brain, which has high levels of glycogen but low specific activity of UTP—glucose-1-phosphate uridylyltransferase.[14] In animal cells, UTP—glucose-1-phosphate uridylyltransferase is found predominantly in the cytoplasm.
UTP—glucose-1-phosphate uridylyltransferase is also required for galactose metabolism in animals and microorganisms. In galactose metabolism, the enzyme galactose 1-phosphate uridylyltransferase transfers a phosphate from UDP-glucose to galactose 1-phosphate to produce UDP-galactose, which is then converted to UDP-glucose.[15] Bacteria with defective UTP—glucose-1-phosphate uridylyltransferase are unable to incorporate galactose into their cell walls.[16]
Mechanism
In this enzyme's primary reaction, the phosphate group on glucose-1-phosphate replaces the phosphoanhydride bond on UTP. This reaction is readily reversible and the Gibbs Free Energy is close to zero. However, under typical cellular conditions, inorganic pyrophosphatase quickly hydrolyzes the pyrophosphate product and drives the reaction forward by Le Chatelier's Principle.
UTP—glucose-1-phosphate uridylyltransferase uses an ordered sequential Bi Bi mechanism for both the forward and reverse reactions.[17] In yeast, the enzyme follows simple Michaelis-Menten kinetics and does not exhibit cooperativity between the subunits in the octamer.[8]
Similar to other sugar nucleotidyltransferases, UTP—glucose-1-phosphate uridylyltransferase activity requires two divalent cations to stabilize the binding of negatively charged phosphate groups.[18] Magnesium typically serves in this role, but other ions such as manganese(II), cobalt(II), and nickel(II) can also substitute with a ~75% reduction in the optimal activity.[19] X-ray crystallography experiments have shown that one Mg2+ ion is coordinated by a phosphoryl oxygen on glucose 1-phosphate and by an α-phosphoryl oxygen on UTP.[5] In addition to stabilizing the negatively charged phosphates, Mg2+ is thought to orient the glucose 1-phosphate for nucleophilic attack of the α-phosphorus of UTP.[20]
Regulation
Although functionally similar across species, UDP-glucose pyrophosphorylase has different structures and regulation mechanisms in different organisms.
Microorganisms
In yeast, UTP—glucose-1-phosphate uridylyltransferase is regulated by phosphorylation by PAS kinase.[21] This phosphorylation is reversible and controls the partition of sugar flux towards glycogen and cell wall synthesis.
Plants
UTP—glucose-1-phosphate uridylyltransferase in plants is regulated through oligomerization and possibly phosphorylation.[22] In barley, it has been shown that UDP-glucose pyrophosphorylase is only active in monomeric form but readily forms oligomers, suggesting that oligomerization may be a form of regulation of the enzyme. In rice, cold stress decreases N-glycosylation of the enzyme, which is thought to alter the enzyme's activity in response to cold.[23]
In Arabidopsis, there are two isozymes of UTP—glucose-1-phosphate uridylyltransferase: UGP1 and UGP2.[24] These two isozymes have almost identical activities and differ in only 32 amino acids, all of which are located on the outer surface of the protein away from the active site. These minor differences may allow for differential allosteric regulation of isozyme activity. UGP1 and UGP2 are differentially expressed in different parts of the plant. UGP1 expression is widely expressed in the majority of tissues while UGP2 is expressed primarily in flowers, suggesting that UGP1 is the major form of the enzyme and UGP2 serves an auxiliary function. Indeed, UGP2 expression is increased in response to stressors such as phosphate deficiency, indicating that UGP2 probably functions as a backup to UGP1 when the plant is under environmental stress.
Animals
The control of UTP—glucose-1-phosphate uridylyltransferase activity is primarily achieved by genetic means (i.e. regulation of transcription and translation). Similar to most enzymes, UTP—glucose-1-phosphate uridylyltransferase is inhibited by its product, UDP-glucose. However, the enzyme is not subject to significant allosteric regulation, which is logical given the widespread use of UDP-glucose in a variety of metabolic pathways.
Humans
In humans, UDP-glucose pyrophosphorylase is active as an octamer.[7] The enzyme's activity is also modified by O-glycosylation.[25] Similar to other mamallian species, there two different isoforms in humans that are produced by alternative splicing of the gene.[3][11][26] The isoforms differ by only 11 amino acids at the N-terminus and no significant differences in their functional activity have been identified.
Disease relevance
In humans, galactosemia is a disorder that affects the development of newborns and children as they cannot metabolize the sugar galactose properly. It is speculated that overexpression of UDP-glucose pyrophosphorylase may relieve symptoms in humans with galactosemia.[27]
In cancer cells, which typically have high rates of glycolysis and decreased glycogen content, the activity of UTP—glucose-1-phosphate uridylyltransferase is often downregulated by up to 50-60% compared to normal cells.[28] The abnormally low activity of UTP—glucose-1-phosphate uridylyltransferase is due to decreased levels of the enzyme and the downregulation of other enzymes in the glycogenic pathway including glycogen synthase and phosphoglucomutase.
UTP—glucose-1-phosphate uridylyltransferase has been found to be an important virulence factor in a variety of pathogens including bacteria and protozoa.[29][30] For example, the enzyme has been found to be required for the biosynthesis of capsular polysaccharide, an important virulence factor of streptococcus pneumoniae, a bacterial cause of pneumonia, bronchitis, and other breathing issues.[31] As a result, the enzyme has attracted attention as a potential target for pharmaceuticals. However, in order to achieve specificity, the drugs must be designed to specifically target allosteric sites on the surface of the protein because the active site is highly conserved across species.[3]
UDP-glucose pyrophosphorylase (UGP2) was recently found to be implicated in novel neurodevelopmental disorder in humans, known as [32] also referred to as Barakat-Perenthaler syndrome.[33] This disorder was first described in 22 individuals from 15 families, presenting with a severe epileptic encephalopathy, neurodevelopmental delay with absence of virtually all developmental milestones, intractable seizures, progressive microcephaly, visual disturbance and similar minor dysmorphisms. Barakat and colleagues identified a recurrent homozygous mutation in all affected individuals (chr2:64083454A > G), which mutates the translational start site of the shorter protein isoform of UGP2. Therefore, the shorter protein isoform can no longer be produced in patients harboring the homozygous mutation. Functional studies from the same group showed that the short protein isoform is normally predominantly expressed in human brain. Therefore, the recurrent mutation leads to a tissue-specific absence of UGP2 in brain, which leads to altered glycogen metabolism, upregulated unfolded protein response and premature neuronal differentiation. Other bi-allelic loss-of-function mutations in UGP2 are likely lethal, as human embryonic stem cells depleted of both short and long isoforms of UGP2 fail to differentiate in cardiomyocytes and blood cells. Hence, the identification of this new disease also shows that isoform-specific start-loss mutations causing expression loss of a tissue-relevant isoform of an essential protein can cause a genetic disease, even when an organism-wide protein absence is incompatible with life. A therapy for Barakat-Perenthaler syndrome does currently not exist.
References
- Sandhoff K, van Echten G, Schröder M, Schnabel D, Suzuki K (August 1992). "Metabolism of glycolipids: the role of glycolipid-binding proteins in the function and pathobiochemistry of lysosomes". Biochemical Society Transactions. 20 (3): 695–9. doi:10.1042/bst0200695. PMID 1426613.
- Alonso MD, Lomako J, Lomako WM, Whelan WJ (September 1995). "A new look at the biogenesis of glycogen". FASEB Journal. 9 (12): 1126–37. doi:10.1096/fasebj.9.12.7672505. PMID 7672505. S2CID 40281321.
- Führing JI, Cramer JT, Schneider J, Baruch P, Gerardy-Schahn R, Fedorov R (April 2015). "A quaternary mechanism enables the complex biological functions of octameric human UDP-glucose pyrophosphorylase, a key enzyme in cell metabolism". Scientific Reports. 5 (1): 9618. Bibcode:2015NatSR...5E9618F. doi:10.1038/srep09618. PMC 5381698. PMID 25860585.
- Kim H, Choi J, Kim T, Lokanath NK, Ha SC, Suh SW, Hwang HY, Kim KK (April 2010). "Structural basis for the reaction mechanism of UDP-glucose pyrophosphorylase". Molecules and Cells. 29 (4): 397–405. doi:10.1007/s10059-010-0047-6. PMID 20238176. S2CID 25022544.
- Thoden JB, Holden HM (July 2007). "Active site geometry of glucose-1-phosphate uridylyltransferase". Protein Science. 16 (7): 1379–88. doi:10.1110/ps.072864707. PMC 2206702. PMID 17567737.
- Disease, Seattle Structural Genomics Center for Infectious (2016). "Crystal structure of UDP-glucose pyrophosporylase / UTP-glucose-1-phosphate uridylyltransferase from Burkholderia xenovorans". Worldwide Protein Data Bank. doi:10.2210/pdb5j49/pdb.
- Yu Q, Zheng X (March 2012). "The crystal structure of human UDP-glucose pyrophosphorylase reveals a latch effect that influences enzymatic activity". The Biochemical Journal. 442 (2): 283–91. doi:10.1042/BJ20111598. PMID 22132858.
- Roeben A, Plitzko JM, Körner R, Böttcher UM, Siegers K, Hayer-Hartl M, Bracher A (December 2006). "Structural basis for subunit assembly in UDP-glucose pyrophosphorylase from Saccharomyces cerevisiae". Journal of Molecular Biology. 364 (4): 551–60. doi:10.1016/j.jmb.2006.08.079. PMID 17010990.
- Kleczkowski LA, Geisler M, Fitzek E, Wilczynska M (November 2011). "A common structural blueprint for plant UDP-sugar-producing pyrophosphorylases". The Biochemical Journal. 439 (3): 375–9. doi:10.1042/BJ20110730. PMID 21992098.
- Flores-Díaz M, Alape-Girón A, Persson B, Pollesello P, Moos M, von Eichel-Streiber C, Thelestam M, Florin I (September 1997). "Cellular UDP-glucose deficiency caused by a single point mutation in the UDP-glucose pyrophosphorylase gene". The Journal of Biological Chemistry. 272 (38): 23784–91. doi:10.1074/jbc.272.38.23784. PMID 9295324.
- "UGP2 - UTP--glucose-1-phosphate uridylyltransferase - Homo sapiens (Human) - UGP2 gene & protein". www.uniprot.org. Retrieved 2017-03-06.
- Mendicino J (December 1960). "Sucrose phosphate synthesis in wheat germ and green leaves". The Journal of Biological Chemistry. 235 (12): 3347–52. doi:10.1016/S0021-9258(18)64469-2. PMID 13769376.
- Turnquist, Richard L.; Hansen, R. Gaurth (1973-01-01). "2 Uridine Diphosphoryl Glucose Pyrophosphorylase". In Boyer, Paul D. (ed.). The Enzymes. Group Transfer Part A: Nucleotidyl Transfer Nucleosidyl Transfer Acyl Transfer Phosphoryl Transfer. Vol. 8. Academic Press. pp. 51–71. doi:10.1016/S1874-6047(08)60062-1. ISBN 9780121227081.
- Villar-Palasi C, Larner J (March 1960). "Insulin-mediated effect on the activity of UDPG-glycogen transglucosylase of muscle". Biochimica et Biophysica Acta. 39: 171–3. doi:10.1016/0006-3002(60)90142-6. PMID 13842294.
- Bosch AM (August 2006). "Classical galactosaemia revisited". Journal of Inherited Metabolic Disease. 29 (4): 516–25. doi:10.1007/s10545-006-0382-0. PMID 16838075. S2CID 16382462.
- Sundararajan TA, Rapin AM, Kalckar HM (December 1962). "Biochemical observations on E. coli mutants defective in uridine diphosphoglucose". Proceedings of the National Academy of Sciences of the United States of America. 48 (12): 2187–93. Bibcode:1962PNAS...48.2187S. doi:10.1073/pnas.48.12.2187. PMC 221142. PMID 13979281.
- Tsuboi KK, Fukunaga K, Petricciani JC (February 1969). "Purification and specific kinetic properties of erythrocyte uridine diphosphate glucose pyrophosphorylase". The Journal of Biological Chemistry. 244 (3): 1008–15. doi:10.1016/S0021-9258(18)91886-7. PMID 5782905.
- Zea CJ, Camci-Unal G, Pohl NL (July 2008). "Thermodynamics of binding of divalent magnesium and manganese to uridine phosphates: implications for diabetes-related hypomagnesaemia and carbohydrate biocatalysis". Chemistry Central Journal. 2 (1): 15. doi:10.1186/1752-153x-2-15. PMC 2490692. PMID 18627619.
- Gustafson GL, Gander JE (March 1972). "Uridine diphosphate glucose pyrophosphorylase from Sorghum vulgare. Purification and kinetic properties". The Journal of Biological Chemistry. 247 (5): 1387–97. doi:10.1016/S0021-9258(19)45571-3. PMID 5012314.
- Sivaraman J, Sauvé V, Matte A, Cygler M (November 2002). "Crystal structure of Escherichia coli glucose-1-phosphate thymidylyltransferase (RffH) complexed with dTTP and Mg2+". The Journal of Biological Chemistry. 277 (46): 44214–9. doi:10.1074/jbc.M206932200. PMID 12171937.
- Rutter J, Probst BL, McKnight SL (October 2002). "Coordinate regulation of sugar flux and translation by PAS kinase". Cell. 111 (1): 17–28. doi:10.1016/s0092-8674(02)00974-1. PMID 12372297. S2CID 6883785.
- Kleczkowski LA, Geisler M, Ciereszko I, Johansson H (March 2004). "UDP-glucose pyrophosphorylase. An old protein with new tricks". Plant Physiology. 134 (3): 912–8. doi:10.1104/pp.103.036053. PMC 523891. PMID 15020755.
- Komatsu S, Yamada E, Furukawa K (January 2009). "Cold stress changes the concanavalin A-positive glycosylation pattern of proteins expressed in the basal parts of rice leaf sheaths". Amino Acids. 36 (1): 115–23. doi:10.1007/s00726-008-0039-4. PMID 18278531. S2CID 1797342.
- Meng M, Geisler M, Johansson H, Harholt J, Scheller HV, Mellerowicz EJ, Kleczkowski LA (May 2009). "UDP-glucose pyrophosphorylase is not rate limiting, but is essential in Arabidopsis". Plant & Cell Physiology. 50 (5): 998–1011. doi:10.1093/pcp/pcp052. PMID 19366709.
- Wells, Lance; Hart, Gerald W. (2003-07-03). "O-GlcNAc turns twenty: functional implications for post-translational modification of nuclear and cytosolic proteins with a sugar". FEBS Letters. 546 (1): 154–158. doi:10.1016/S0014-5793(03)00641-0. ISSN 1873-3468. PMID 12829252. S2CID 24587552.
- Duggleby RG, Chao YC, Huang JG, Peng HL, Chang HY (January 1996). "Sequence differences between human muscle and liver cDNAs for UDPglucose pyrophosphorylase and kinetic properties of the recombinant enzymes expressed in Escherichia coli". European Journal of Biochemistry. 235 (1–2): 173–9. doi:10.1111/j.1432-1033.1996.00173.x. PMID 8631325.
- Lai K, Elsas LJ (May 2000). "Overexpression of human UDP-glucose pyrophosphorylase rescues galactose-1-phosphate uridyltransferase-deficient yeast". Biochemical and Biophysical Research Communications. 271 (2): 392–400. doi:10.1006/bbrc.2000.2629. PMID 10799308.
- Nigam VN, Macdonald HL, Cantero A (February 1962). "Limiting factors for glycogen storage in tumors. I. Limiting enzymes". Cancer Research. 22 (2): 131–8. PMID 14479721.
- Jiang SS, Lin TY, Wang WB, Liu MC, Hsueh PR, Liaw SJ (May 2010). "Characterization of UDP-glucose dehydrogenase and UDP-glucose pyrophosphorylase mutants of Proteus mirabilis: defectiveness in polymyxin B resistance, swarming, and virulence". Antimicrobial Agents and Chemotherapy. 54 (5): 2000–9. doi:10.1128/AAC.01384-09. PMC 2863647. PMID 20160049.
- Klein KA, Fukuto HS, Pelletier M, Romanov G, Grabenstein JP, Palmer LE, Ernst R, Bliska JB (February 2012). "A transposon site hybridization screen identifies galU and wecBC as important for survival of Yersinia pestis in murine macrophages". Journal of Bacteriology. 194 (3): 653–62. doi:10.1128/JB.06237-11. PMC 3264090. PMID 22139502.
- Bonofiglio L, García E, Mollerach M (October 2005). "Biochemical characterization of the pneumococcal glucose 1-phosphate uridylyltransferase (GalU) essential for capsule biosynthesis". Current Microbiology. 51 (4): 217–21. doi:10.1007/s00284-005-4466-0. PMID 16132460. S2CID 13591083.
- Perenthaler E, Nikoncuk A, Yousefi S, Berdowski WM, Alsagob M, Capo I, et al. (March 2020). "Loss of UGP2 in brain leads to a severe epileptic encephalopathy, emphasizing that bi-allelic isoform-specific start-loss mutations of essential genes can cause genetic diseases". Acta Neuropathologica. 139 (3): 415–442. doi:10.1007/s00401-019-02109-6. PMC 7035241. PMID 31820119.
- "#618744: Epileptic Encephalopathy, Early Infantile 83; EIEE83". Online Mendelian Inheritance in Man (OMIM).
External links
- UDP+Glucose+Pyrophosphorylase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
