Bacterial transcription
Bacterial transcription is the process in which a segment of bacterial DNA is copied into a newly synthesized strand of messenger RNA (mRNA) with use of the enzyme RNA polymerase.
.jpg.webp)
The process occurs in three main steps: initiation, elongation, and termination; and the end result is a strand of mRNA that is complementary to a single strand of DNA. Generally, the transcribed region accounts for more than one gene.[1] In fact, many prokaryotic genes occur in operons, which are a series of genes that work together to code for the same protein or gene product and are controlled by a single promoter.[2] Bacterial RNA polymerase is made up of four subunits and when a fifth subunit attaches, called the σ-factor, the polymerase can recognize specific binding sequences in the DNA, called promoters.[3] The binding of the σ-factor to the promoter is the first step in initiation. Once the σ-factor releases from the polymerase, elongation proceeds.[4] The polymerase continues down the double stranded DNA, unwinding it and synthesizing the new mRNA strand until it reaches a termination site. There are two termination mechanisms that are discussed in further detail below. Termination is required at specific sites for proper gene expression to occur.[5] Gene expression determines how much gene product, such as protein, is made by the gene.[2] Transcription is carried out by RNA polymerase but its specificity is controlled by sequence-specific DNA binding proteins called transcription factors. Transcription factors work to recognize specific DNA sequences and based on the cells needs, promote or inhibit additional transcription.[6] Similar to other taxa, bacteria experience bursts of transcription.[7]: 125 [8][9][10][11][12][13] The work of the Jones team in Jones et al 2014 explains some of the underlying causes of bursts and other variability, including stability of the resulting mRNA,[7]: 125 the strength of promotion encoded in the relevant promoter[9] and the duration of transcription due to strength of the TF binding site.[9][10][11][12][13] They also found that bacterial TFs linger too briefly for TFs' binding characteristics to explain the sustained transcription of bursts.[8]
Bacterial transcription differs from eukaryotic transcription in several ways. In bacteria, transcription and translation can occur simultaneously in the cytoplasm of the cell, whereas in eukaryotes transcription occurs in the nucleus and translation occurs in the cytoplasm.[14] There is only one type of bacterial RNA polymerase whereas eukaryotes have 3 types.[2] Bacteria have a σ-factor that detects and binds to promoter sites but eukaryotes do not need a σ-factor. Instead, eukaryotes have transcription factors that allow the recognition and binding of promoter sites.[2]
Overall, transcription within bacteria is a highly regulated process that is controlled by the integration of many signals at a given time. Bacteria heavily rely on transcription and translation to generate proteins that help them respond specifically to their environment.[4]
RNA polymerase
RNA polymerase is composed of a core and a holoenzyme structure. The core enzymes contains the catalytic properties of RNA polymerase and is made up of ββ′α2ω subunits. This sequence is conserved across all bacterial species. The holoenzyme is composed of a specific component known as the sigma factor. The sigma factor functions in aiding in promoter recognition, correct placement of RNA polymerase, and beginning unwinding at the start site. After the sigma factor performs its required function, it dissociates, while the catalytic portion remains on the DNA and continues transcription.[4] Additionally, RNA polymerase contains a core Mg+ ion that assists the enzyme with its catalytic properties. RNA polymerase works by catalyzing the nucleophilic attack of 3’ OH of RNA to the alpha phosphate of a complementary NTP molecule to create a growing strand of RNA from the template strand of DNA. Furthermore, RNA polymerase also displays exonuclease activities, meaning that if improper base pairing is detected, it can cut out the incorrect bases and replace them with the proper, correct one.[15]
Initiation
Initiation of transcription requires promoter regions, which are specific nucleotide consensus sequences that tell the σ-factor on RNA polymerase where to bind to the DNA.[1] The promoters are usually located 15 to 19 bases apart and are most commonly found upstream of the genes they control.[2][1] RNA polymerase is made up of 4 subunits, which include two alphas, a beta, and a beta prime (α, α, β, and β'). A fifth subunit, sigma (called the σ-factor), is only present during initiation and detaches prior to elongation. Each subunit plays a role in the initiation of transcription, and the σ-factor must be present for initiation to occur. When all σ-factor is present, RNA polymerase is in its active form and is referred to as the holoenzyme. When the σ-factor detaches, it is in core polymerase form.[4][1] The σ-factor recognizes promoter sequences at -35 and -10 regions and transcription begins at the start site (+1). The sequence of the -10 region is TATAAT and the sequence of the -35 region is TTGACA.[1]
- The σ-factor binds to the -35 promoter region. At this point, the holoenzyme is referred to as the closed complex because the DNA is still double stranded (connected by hydrogen bonds).[4]
- Once the σ-factor binds, the remaining subunits of the polymerase attach to the site. The high concentration of adenine-thymine bonds at the -10 region facilitates the unwinding of the DNA. At this point, the holoenzyme is called the open complex.[16] This open complex is also called the transcription bubble.[14] Only one strand of DNA, called the template strand (also called the noncoding strand or nonsense/antisense strand), gets transcribed.[2]
- Transcription begins and short "abortive" nucleotide sequences approximately 10 base pairs long are produced. These short sequences are nonfunctional pieces of RNA that are produced and then released.[1] Generally, this nucleotide sequence consists of about twelve base pairs and aids in contributing to the stability of RNA polymerase so it is able to continue along the strand of DNA.[15]
- The σ-factor is needed to initiate transcription but is not needed to continue transcribing the DNA. The σ-factor dissociates from the core enzyme and elongation proceeds. This signals the end of the initiation phase and the holoenzyme is now in core polymerase form.[4]
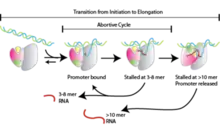
The promoter region is a prime regulator of transcription. Promoter regions regulate transcription of all genes within bacteria. As a result of their involvement, the sequence of base pairs within the promoter region is significant; the more similar the promoter region is to the consensus sequence, the tighter RNA polymerase will be able to bind. This binding contributes to the stability of elongation stage of transcription and overall results in more efficient functioning. Additionally, RNA polymerase and σ-factors are in limited supply within any given bacterial cell. Consequently, σ-factor binding to the promoter is affected by these limitations. All promoter regions contain sequences that are considered non-consensus and this helps to distribute σ-factors across the entirety of the genome.[17]
Elongation
During elongation, RNA polymerase slides down the double stranded DNA, unwinding it and transcribing (copying) its nucleotide sequence into newly synthesized RNA. The movement of the RNA-DNA complex is essential for the catalytic mechanism of RNA polymerase. Additionally, RNA polymerase increases the overall stability of this process by acting as a link between the RNA and DNA strands. [18] New nucleotides that are complementary to the DNA template strand are added to the 3' end of the RNA strand.[4] The newly formed RNA strand is practically identical to the DNA coding strand (sense strand or non-template strand), except it has uracil substituting thymine, and a ribose sugar backbone instead of a deoxyribose sugar backbone. Because nucleoside triphosphates (NTPs) need to attach to the OH- molecule on the 3' end of the RNA, transcription always occurs in the 5' to 3' direction. The four NTPs are adenosine-5'-triphosphate (ATP), guanoside-5'-triphosphate (GTP), uridine-5'-triphosphate (UTP), and cytidine-5'-triphosphate (CTP).[16] The attachment of NTPs onto the 3' end of the RNA transcript provides the energy required for this synthesis.[2] NTPs are also energy producing molecules that provide the fuel that drives chemical reactions in the cell.[4]
Multiple RNA polymerases can be active at once, meaning many strands of mRNA can be produced very quickly.[2] RNA polymerase moves down the DNA rapidly at approximately 40 bases per second. Due to the quick nature of this process, DNA is continually unwound ahead of RNA polymerase and then rewound once RNA polymerase moves along further. [18][1] The polymerase has a proofreading mechanism that limits mistakes to about 1 in 10,000 nucleotides transcribed.[19] RNA polymerase has lower fidelity (accuracy) and speed than DNA polymerase.[2] DNA polymerase has a very different proofreading mechanism that includes exonuclease activity, which contributes to the higher fidelity. The consequence of an error during RNA synthesis is usually harmless, where as an error in DNA synthesis could be detrimental.[2]
The promoter sequence determines the frequency of transcription of its corresponding gene.[1]
Termination
In order for proper gene expression to occur, transcription must stop at specific sites. Two termination mechanisms are well known:
- Intrinsic termination (also called Rho-independent termination): Specific DNA nucleotide sequences signal the RNA polymerase to stop. The sequence is commonly a palindromic sequence that causes the strand to loop which stalls the RNA polymerase.[16] Generally, this type of termination follows the same standard procedure. A pause will occur due to a polyuridine sequence that allows the formation of a hairpin loop. This hairpin loop will aid in forming a trapped complex, which will ultimately cause the dissociation of RNA polymerase from the template DNA strand and halt transcription.[15]
- Rho-dependent termination: ρ factor (rho factor) is a terminator protein that attaches to the RNA strand and follows behind the polymerase during elongation.[5] Once the polymerase nears the end of the gene it is transcribing, it encounters a series of G nucleotides which causes it to stall.[1] This stalling allows the rho factor to catch up to the RNA polymerase. The rho protein then pulls the RNA transcript from the DNA template and the newly synthesized mRNA is released, ending transcription.[5][1] Rho factor is a protein complex that also displays helicase activities (is able to unwind the nucleic acid strands). It will bind to the DNA in cytosine rich regions and when RNA polymerase encounters it, a trapped complex will form causing the dissociation of all molecules involved and end transcription.[15]
The termination of DNA transcription in bacteria may be stopped by certain mechanisms wherein the RNA polymerase will ignore the terminator sequence until the next one is reached. This phenomenon is known as antitermination and is utilized by certain bacteriophages.[20]
References
- "Prokaryotic Transcription and Translation | Biology for Majors I". courses.lumenlearning.com. Retrieved 2019-10-06.
- Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2008). Molecular Biology of the Cell (Sixth ed.). New York: Garland Science. ISBN 978-0-8153-4524-4.
- Bartee L (2017). Prokaryotic Transcription. Principles of Biology: Biology 211, 212, and 213. Open Oregon Educational Resources. Retrieved 2019-10-08.
- Lodish H, Berk A, Zipursky SL, Matsudaira P, Baltimore D, Darnel l J (2000). "Bacterial Transcription Initiation". Molecular Cell Biology (4th ed.).
- "Stages of transcription". Khan Academy. Retrieved 2019-10-07.
- Browning DF, Butala M, Busby SJ (September 2019). "Bacterial Transcription Factors: Regulation by Pick "N" Mix". Journal of Molecular Biology. 431 (20): 4067–4077. doi:10.1016/j.jmb.2019.04.011. PMID 30998934.
- El-Mansi, E. M. T.; Nielsen, Jens; Mousdale, David; Allman, Tony; Carlson, Ross (2019). El-Mansi, Mansi; Nielsen, Jens; Mousdale, David M.; Allman, Tony; Carlson, Ross (eds.). Fermentation Microbiology and Biotechnology (4 ed.). Boca Raton: CRC Press. pp. xix+419. doi:10.1201/9780429506987. ISBN 978-1-138-58102-9. OCLC 1080190329. S2CID 220766937. ISBN 978-0-429-50698-7.
- Symmons, Orsolya; Raj, Arjun (2016). "What's Luck Got to Do with It: Single Cells, Multiple Fates, and Biological Nondeterminism". Molecular Cell. Cell Press. 62 (5): 788–802. doi:10.1016/j.molcel.2016.05.023. ISSN 1097-2765. PMC 4900469. PMID 27259209.
- Payne, Joshua L.; Wagner, Andreas (2018-11-01). "The causes of evolvability and their evolution" (PDF). Nature Reviews Genetics. Nature Portfolio. 20 (1): 24–38. doi:10.1038/s41576-018-0069-z. ISSN 1471-0056. PMID 30385867. S2CID 53204518.
- Typas, Athanasios; Sourjik, Victor (2015-08-10). "Bacterial protein networks: properties and functions". Nature Reviews Microbiology. Nature Portfolio. 13 (9): 559–572. doi:10.1038/nrmicro3508. ISSN 1740-1526. PMID 26256789. S2CID 12498094.
- Bashor, Caleb J.; Collins, James J. (2018-05-20). "Understanding Biological Regulation Through Synthetic Biology". Annual Review of Biophysics. Annual Reviews. 47 (1): 399–423. doi:10.1146/annurev-biophys-070816-033903. hdl:1721.1/119222. ISSN 1936-122X. PMID 29547341.
- Xu, Heng; Skinner, Samuel O.; Sokac, Anna Marie; Golding, Ido (2016-09-13). "Stochastic Kinetics of Nascent RNA". Physical Review Letters. American Physical Society. 117 (12): 128101. Bibcode:2016PhRvL.117l8101X. doi:10.1103/physrevlett.117.128101. ISSN 0031-9007. PMC 5033037. PMID 27667861. NIHMS 816487.
- Eling, Nils; Morgan, Michael D.; Marioni, John C. (2019-05-21). "Challenges in measuring and understanding biological noise". Nature Reviews Genetics. Nature Portfolio. 20 (9): 536–548. doi:10.1038/s41576-019-0130-6. ISSN 1471-0056. PMC 7611518. PMID 31114032. EMSID: 85286
- "15.2: Prokaryotic Transcription". General Biology (OpenStax). LibreTexts. 2015-11-02. Retrieved 2019-10-08.
- Bębenek A, Ziuzia-Graczyk I (October 2018). "Fidelity of DNA replication-a matter of proofreading". Current Genetics. 64 (5): 985–996. doi:10.1007/s00294-018-0820-1. PMC 6153641. PMID 29500597.
- "7.6C: Prokaryotic Transcription and Translation Are Coupled". General Biology (OpenStax). LibreTexts. 2017-05-17. Retrieved 2019-10-07.
- Browning DF, Busby SJ (January 2004). "The regulation of bacterial transcription initiation". Nature Reviews. Microbiology. 2 (1): 57–65. doi:10.1038/nrmicro787. PMID 15035009. S2CID 680370.
- Clark, Mary Ann (5 March 2018). "Prokaryotic Transcription". Biology 2e. BC Open Textbooks. Retrieved 2019-11-29.
- Milo R, Phillips R. "What is the error rate in transcription and translation?". Cell Biology by the Numbers. Retrieved 2019-11-15.
- Lewin B, Krebs JE, Goldstein ES, Kilpatrick ST (2011). Lewin's genes X (10th ed.). Sudbury, Massachusetts: Jones and Bartlett. ISBN 978-0-7637-6632-0. OCLC 456641931.