Penicillin-binding proteins
Penicillin-binding proteins (PBPs) are a group of proteins that are characterized by their affinity for and binding of penicillin. They are a normal constituent of many bacteria; the name just reflects the way by which the protein was discovered. All β-lactam antibiotics (except for tabtoxinine-β-lactam, which inhibits glutamine synthetase) bind to PBPs, which are essential for bacterial cell wall synthesis. PBPs are members of a subgroup of enzymes called transpeptidases. Specifically, PBPs are DD-transpeptidases.
| Penicillin-binding protein, transpeptidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | PCN-bd_Tpept | ||||||||
| Pfam | PF00905 | ||||||||
| InterPro | IPR001460 | ||||||||
| OPM superfamily | 195 | ||||||||
| OPM protein | 5hlb | ||||||||
| Membranome | 541 | ||||||||
| |||||||||
| Penicillin-binding protein, dimerisation domain | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||
| Symbol | PBP_dimer | ||||||||||
| Pfam | PF03717 | ||||||||||
| InterPro | IPR005311 | ||||||||||
| |||||||||||
Diversity
There are a large number of PBPs, usually several in each organism, and they are found as both membrane-bound and cytoplasmic proteins. For example, Spratt (1977) reports that six different PBPs are routinely detected in all strains of E. coli ranging in molecular weight from 40,000 to 91,000.[3] The different PBPs occur in different numbers per cell and have varied affinities for penicillin. The PBPs are usually broadly classified into high-molecular-weight (HMW) and low-molecular-weight (LMW) categories.[4] Proteins that have evolved from PBPs occur in many higher organisms and include the mammalian LACTB protein.[5]
Function
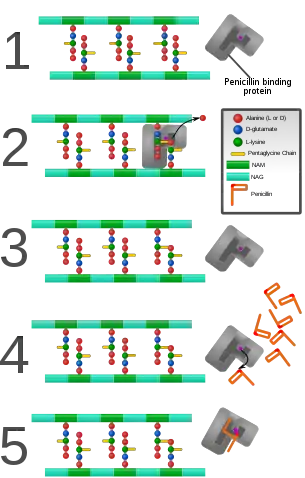
PBPs are all involved in the final stages of the synthesis of peptidoglycan, which is the major component of bacterial cell walls. Bacterial cell wall synthesis is essential to growth, cell division (thus reproduction) and maintaining the cellular structure in bacteria.[2] Inhibition of PBPs leads to defects in cell wall structure and irregularities in cell shape, for example filamentation, pseudomulticellular forms, lesions leading to spheroplast formation, and eventual cell death and lysis.[6]
PBPs have been shown to catalyze a number of reactions involved in the process of synthesizing cross-linked peptidoglycan from lipid intermediates and mediating the removal of D-alanine from the precursor of peptidoglycan. Purified enzymes have been shown to catalyze the following reactions: D-alanine carboxypeptidase, peptidoglycan transpeptidase, and peptidoglycan endopeptidase. In all bacteria that have been studied, enzymes have been shown to catalyze more than one of the above reactions.[3] The enzyme has a penicillin-insensitive transglycosylase N-terminal domain (involved in formation of linear glycan strands) and a penicillin-sensitive transpeptidase C-terminal domain (involved in cross-linking of the peptide subunits) and the serine at the active site is conserved in all members of the PBP family.[4]
Some low-molecular-weight PBPs associate with the MreB cytoskeleton and follow its rotation around the cell, inserting petipdoglycan in an oriented manner during cell growth.[7] In contrast, high-molecular-weight PBPs are independent from MreB and maintain cell wall integrity by detecting and repairing defects in the peptidoglycan.[8]
Antibiotics
PBPs bind to β-lactam antibiotics because they are similar in chemical structure to the modular pieces that form the peptidoglycan.[9] When they bind to penicillin, the β-lactam amide bond is ruptured to form a covalent bond with the catalytic serine residue at the PBPs active site. This is an irreversible reaction and inactivates the enzyme.
There has been a great deal of research into PBPs because of their role in antibiotics and resistance. Bacterial cell wall synthesis and the role of PBPs in its synthesis is a very good target for drugs of selective toxicity because the metabolic pathways and enzymes are unique to bacteria.[10] Resistance to antibiotics has come about through overproduction of PBPs and formation of PBPs that have low affinity for penicillins (among other mechanisms such as lactamase production). These experiments change the structure of PBP by adding different amino acids into the protein, allowing for new discovery of how the drug interacts with the protein. Research on PBPs has led to the discovery of new semi-synthetic β-lactams, wherein altering the side-chains on the original penicillin molecule has increased the affinity of PBPs for penicillin, and, thus, increased effectiveness in bacteria with developing resistance.
Presence of the protein penicillin binding protein 2A (PBP2A) is responsible for the antibiotic resistance seen in methicillin-resistant Staphylococcus aureus (MRSA).[11]
The β-lactam ring is a structure common to all β-lactam antibiotics.[12]
Other images
See also
PASTA domain
References
- Sainsbury S, Bird L, Rao V, Shepherd SM, Stuart DI, Hunter WN, Owens RJ, Ren J (January 2011). "Crystal structures of penicillin-binding protein 3 from Pseudomonas aeruginosa: comparison of native and antibiotic-bound forms". Journal of Molecular Biology. 405 (1): 173–84. doi:10.1016/j.jmb.2010.10.024. PMC 3025346. PMID 20974151.
- Miyachiro MM, Contreras-Martel C, Dessen A (January 2020). "Penicillin-binding proteins (PBPs) and bacterial cell wall elongation complexes". Subcellular Biochemistry. 93: 273–289. doi:10.1007/978-3-030-28151-9_8. ISBN 978-3-030-28150-2. PMID 31939154.
- Spratt BG (January 1977). "Properties of the penicillin-binding proteins of Escherichia coli K12,". European Journal of Biochemistry. 72 (2): 341–52. doi:10.1111/j.1432-1033.1977.tb11258.x. PMID 319999.
- Basu J, Chattopadhyay R, Kundu M, Chakrabarti P (July 1992). "Purification and partial characterization of a penicillin-binding protein from Mycobacterium smegmatis". Journal of Bacteriology. 174 (14): 4829–32. doi:10.1128/jb.174.14.4829-4832.1992. PMC 206282. PMID 1624470.
- Peitsaro N, Polianskyte Z, Tuimala J, Pörn-Ares I, Liobikas J, Speer O, Lindholm D, Thompson J, Eriksson O (January 2008). "Evolution of a family of metazoan active-site-serine enzymes from penicillin-binding proteins: a novel facet of the bacterial legacy". BMC Evolutionary Biology. 8: 16. doi:10.1186/1471-2148-8-26. PMC 2266909. PMID 18226203.
- Cushnie TP, O'Driscoll NH, Lamb AJ (December 2016). "Morphological and ultrastructural changes in bacterial cells as an indicator of antibacterial mechanism of action". Cellular and Molecular Life Sciences. 73 (23): 4471–4492. doi:10.1007/s00018-016-2302-2. hdl:10059/2129. PMID 27392605. S2CID 2065821.
- Dion, Michael F.; Kapoor, Mrinal; Sun, Yingjie; Wilson, Sean; Ryan, Joel; Vigouroux, Antoine; Teeffelen, Sven van; Oldenbourg, Rudolf; Garner, Ethan C. (2019-05-13). "Bacillus subtilis cell diameter is determined by the opposing actions of two distinct cell wall synthetic systems". Nature Microbiology. 4 (8): 1294–1305. doi:10.1038/s41564-019-0439-0. ISSN 2058-5276. PMC 6656618. PMID 31086310.
- Vigouroux, Antoine; Cordier, Baptiste; Aristov, Andrey; Alvarez, Laura; Özbaykal, Gizem; Chaze, Thibault; Oldewurtel, Enno Rainer; Matondo, Mariette; Cava, Felipe; Bikard, David; van Teeffelen, Sven (2020-01-06). Jie Xiao (ed.). "Class-A penicillin binding proteins do not contribute to cell shape but repair cell-wall defects". eLife. 9: –51998. doi:10.7554/eLife.51998. ISSN 2050-084X. PMC 7002073. PMID 31904338.
- Nguyen-Distèche M, Leyh-Bouille M, Ghuysen JM (October 1982). "Isolation of the membrane-bound 26 000-Mr penicillin-binding protein of Streptomyces strain K15 in the form of a penicillin-sensitive D-alanyl-D-alanine-cleaving transpeptidase". Biochemical Journal. 207 (1): 109–15. doi:10.1042/bj2070109. PMC 1153830. PMID 7181854.
- Chambers HF (March 1999). "Penicillin-binding protein-mediated resistance in pneumococci and staphylococci". Journal of Infectious Diseases. 179 Suppl 2: S353-9. doi:10.1086/513854. PMID 10081507.
- Ubukata K, Nonoguchi R, Matsuhashi M, Konno M (May 1989). "Expression and inducibility in Staphylococcus aureus of the mecA gene, which encodes a methicillin-resistant S. aureus-specific penicillin-binding protein". Journal of Bacteriology. 171 (5): 2882–5. doi:10.1128/jb.171.5.2882-2885.1989. PMC 209980. PMID 2708325.
- Pandey N, Cascella M (March 2020). "Beta Lactam Antibiotics". StatPearls. PMID 31424895.
- Bardal SK, Waechter JE, Martin DS (January 2011). "Chapter 18 - Infectious Diseases". Applied Pharmacology: 233–291. ISBN 9781437703108.

