Rickettsiales
The Rickettsiales, informally called rickettsias, are an order of small Alphaproteobacteria. They are obligate intracellular parasites, and some are notable pathogens, including Rickettsia, which causes a variety of diseases in humans, and Ehrlichia, which causes diseases in livestock. Another genus of well-known Rickettsiales is the Wolbachia, which infect about two-thirds of all arthropods and nearly all filarial nematodes.[2] Genetic studies support the endosymbiotic theory according to which mitochondria and related organelles developed from members of this group.[3]
| Rickettsiales | |
|---|---|
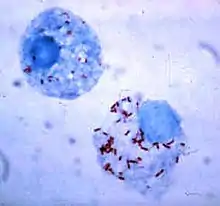 | |
| Rickettsia rickettsii (red dots) in the cell of a deer tick | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Alphaproteobacteria |
| Order: | Rickettsiales Gieszczykiewicz 1939 (Approved Lists 1980) |
| Families | |
| |
The Rickettsiales are difficult to culture, as they rely on living eukaryotic host cells for their survival.
Rickettsiales phylogeny
The Rickettsiales further consist of three known families, the Rickettsiaceae, the Midichloriaceae, and the Ehrlichiaceae. Most studies also support the inclusion of the Holosporaceae, but one study has challenged this view.[4] In that alternative, the Holosporaceae are the sole representatives of their own order, the Holosporales, and as such not part of the Rickettsiales (see the schematic tree below). Other lineages, not clearly part of any family, have been described, as well. Examples include Candidatus Arcanobacter lacustris [5] and Rickettsiales bacterium Ac37b.
| Schematic ribosomal RNA phylogeny of Alphaproteobacteria | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The cladogram of Rickettsidae has been inferred by Ferla et al. [6] from the comparison of 16S + 23S ribosomal RNA sequences. |
Phylogenetic relationship between Rickettsiales and Pelagibacterales (SAR11)
The phylogenetic relationship between these two groups has yet to reach consensus in the scientific literature.
Early reports suggested that they represented sister clades to each other.[7][8] However, later studies suggested that this relationship is false and was due to a phylogenetic artefact, which artificially groups independent AT-rich and fast-evolving lineages (Rickettsiales and Pelagibacterales have both properties) together.[9][10] Upon correcting for this artefact, the Pelagibacterales form a sister clade to the Hyphomicrobiales, Rhodobacterales and Caulobacterales instead.
Another study [4] adheres to the sister relationship between the two clades (see schematic tree). In their classification, the relation between the two orders is retained in the subclass, the Rickettsidae, which include the Rickettsiales, the Pelagibacteriales, and the extinct protomitochondrion (mitochondria themselves are not bacteria, but organelles).[4]
Reductive evolution
Rickettsiales genomes are undergoing reductive evolution and are typically small (generally < 1,5 Mbp), AT-rich (generally < 40% GC) with a low coding density (generally < 85%) and a relatively high number of pseudogenes.[11] Reduction in genome size, % GC and coding density and genes are generally attributed to genetic drift and Muller's ratchet. Genetic drift is enhanced in Rickettsiales genomes due to low population sizes (given their endosymbiotic nature) and frequent population bottlenecks. Similarly, Muller's ratchet is activated through the lack of recombination and horizontal gene transfer (the eukaryotic host cell is a natural barrier).
References
- Li D, Fang J, Wen B, Wua X. (2021). "Molecular identification of a novel intracellular proteobacteria from scallop Chlamys farreri". Aquaculture. 539: 736565. doi:10.1016/j.aquaculture.2021.736565. S2CID 233921822.
{{cite journal}}: CS1 maint: uses authors parameter (link) - Werren JH, Baldo L, Clark ME (2008). "Wolbachia: master manipulators of invertebrate biology". Nat. Rev. Microbiol. 6 (10): 741–51. doi:10.1038/nrmicro1969. PMID 18794912. S2CID 12816914.
- Thomas S. (2016). Rickettsiales:Biology, Molecular Biology, Epidemiology, and Vaccine Development. pp.529. Springer•ISBN 978-3-319-46857-0
- Ferla, M. P.; Thrash, J. C.; Giovannoni, S. J.; Patrick, W. M. (2013). "New rRNA gene-based phylogenies of the Alphaproteobacteria provide perspective on major groups, mitochondrial ancestry and phylogenetic instability". PLOS ONE. 8 (12): e83383. Bibcode:2013PLoSO...883383F. doi:10.1371/journal.pone.0083383. PMC 3859672. PMID 24349502.
- Martijn J, Schulz F, Zaremba-Niedzwiedzka K, et al. (2015). "Single-cell genomics of a rare environmental alphaproteobacterium provides unique insights into Rickettsiaceae evolution". ISME J. 9 (11): 2373–85. doi:10.1038/ismej.2015.46. PMC 4497978. PMID 25848874.
- Ferla MP, Thrash JC, Giovannoni SJ, Patrick WM (2013). "New rRNA gene-based phylogenies of the Alphaproteobacteria provide perspective on major groups, mitochondrial ancestry and phylogenetic instability". PLOS ONE. 8 (12): e83383. Bibcode:2013PLoSO...883383F. doi:10.1371/journal.pone.0083383. PMC 3859672. PMID 24349502.
- Williams KP, Sobral BW, Dickerman AW (2007). "A robust species tree for the alphaproteobacteria". J. Bacteriol. 189 (13): 4578–86. doi:10.1128/JB.00269-07. PMC 1913456. PMID 17483224.
- Thrash JC, Boyd A, Huggett MJ, et al. (2011). "Phylogenomic evidence for a common ancestor of mitochondria and the SAR11 clade". Sci Rep. 1: 13. Bibcode:2011NatSR...1E..13T. doi:10.1038/srep00013. PMC 3216501. PMID 22355532.
- Viklund J, Ettema TJ, Andersson SG (2012). "Independent genome reduction and phylogenetic reclassification of the oceanic SAR11 clade". Mol. Biol. Evol. 29 (2): 599–615. doi:10.1093/molbev/msr203. PMID 21900598.
- Rodríguez-Ezpeleta N, Embley TM (2012). "The SAR11 group of alpha-proteobacteria is not related to the origin of mitochondria". PLOS ONE. 7 (1): e30520. Bibcode:2012PLoSO...730520R. doi:10.1371/journal.pone.0030520. PMC 3264578. PMID 22291975.
- Darby AC, Cho NH, Fuxelius HH, Westberg J, Andersson SG (2007). "Intracellular pathogens go extreme: genome evolution in the Rickettsiales". Trends Genet. 23 (10): 511–20. doi:10.1016/j.tig.2007.08.002. PMID 17822801.