Transmissible gastroenteritis virus
Transmissible gastroenteritis virus or Transmissible gastroenteritis coronavirus (TGEV) is a coronavirus which infects pigs. It is an enveloped, positive-sense, single-stranded RNA virus which enters its host cell by binding to the APN receptor.[2] The virus is a member of the genus Alphacoronavirus, subgenus Tegacovirus, species Alphacoronavirus 1.[3][4]
| Transmissible gastroenteritis virus | |
|---|---|
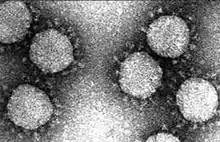 | |
| Electron micrograph of transmissible gastroenteritis coronavirus (TGEV) | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Pisuviricota |
| Class: | Pisoniviricetes |
| Order: | Nidovirales |
| Family: | Coronaviridae |
| Genus: | Alphacoronavirus |
| Subgenus: | Tegacovirus |
| Species: | Alphacoronavirus 1 |
| Virus: | Transmissible gastroenteritis virus |
| Isolates | |
| |
Proteins that contribute to the overall structure of TGEV include the spike (S), envelope (E), membrane (M) and nucleocapsid (N). The genomic size of coronaviruses ranges from approximately 28.6 kilobases.[5] Other coronaviruses that belong to the species Alphacoronavirus 1 are Feline coronavirus, Canine coronavirus and Feline infectious peritonitis virus.
Biology
TGEV belongs to the family Coronaviridae, genus Alphacoronavirus, species Alphacoronavirus 1. It is an enveloped virus with a positive single stranded RNA genome. TGEV has three major structural proteins, which are phosphoprotein (N), integral membrane protein (E1), and large glycoprotein (E2). The N protein encapsulates the genomic RNA, and the S protein forms viral projections.
The 3' segment of about 8000 nucleotides encodes subgenomic RNAs. The remaining part of the genome encodes viral replicase. The three largest gene sequence from 5' to 3' is in the order of E2 to E1 to N. There are about seven other open reading frames that are not structurally related. There are very little overlaps among the genes, and is densely packed. A negative strand is synthesized to serve as a template for transcribing RNAs of one genome size and several subgenome sized RNAs.
The E2 protein forms a petal-shaped 20 nm long projection from the virus's surface. The E2 protein is thought to be involved in pathogenesis by helping the virus enter the host cytoplasm. The E2 protein initially has 1447residues, and then a short hydrophobic sequence is cleaved. After glycosylation of the protein in the golgi, the protein is then incorporated into the new virus. There are several functional domains within the E2 protein. A 20 residue hydrophobic segment at the C-terminus anchors the protein in the lipid membrane. The rest of the protein is divided into two parts, a hydrophilic stretch that is inside the virus and a cysteine rich stretch that are possibly fatty acylation sites. The E1 protein is mostly embedded in the lipid envelop and hence plays an essential role in virus architecture. The E1 protein is postulated to interact with the lymphocyte membrane, which leads to the induction of IFN-coding genes.
Coronaviruses enter the host by first attaching to the host cell using the spike glycoprotein. The S protein interacts with the porcine aminopeptidase N (pAPN), a cellular receptor, to aide in its entry. The same cell receptor is also a point of contact for Human Coronaviruses. A domain in the S spike protein is recognized by pAPN, and transfection of pAPN occurs to nonpermissive cells and infects them with TGEV.
Morphology
The morphology of TGEV was mostly determined by electron microscopy techniques. The morphology is similar to myxovirus and oncogenic virus in that they have surface projections and an envelop. The viruses are mainly circular in shape with a diameter ranging from 100 to 150 nm including the surface projections. The projections were mainly petal-shaped attached by a very narrow stalk. The projections seemed to be very easily detached from the virus and were only found on select areas.
Pathology
TGEV infects pigs. In piglets less than 1 week old, the mortality rate is close to 100%. The pathology of TGEV is similar to that of other coronaviruses. Once the virus infects the host, it multiplies in the cell lining of the small intestine resulting in the loss of absorptive cells that in turn leads to shortening of villi. The infected swine then have reduced capability for digesting food and die from dehydration.[6]
Occurrence
TGE was prevalent in the US when it was originally discovered in the early 20th century. It became more scarce in the late 80's with the rise of porcine respiratory coronavirus (PRCV). It is thought that PRCV provides some immunity to TGE.[7]
Engineering TGEV coronavirus
The Transmissible Gastroenteritis Virus has been engineered as an expression vector. The vector was constructed by replacing the nonessential 3a and 3b ORF, which is driven by the transcription-regulating sequences (TRS) with green fluorescent protein. The resulting construct was still enteropathogenic, but with reduced growth. The infection of cells with this altered virus elicits a specific lactogenic immune response against the heterologous protein. The application of this vector is in the development of a vaccine or even gene therapy. The motivation for engineering the TGEV genome is that coronaviruses have large genomes, so they have room for insertion of foreign genes. Coronaviruses also infect the respiratory tract, and they can be used to target antigens to that area and generate some immune response.
References
- Fehr AR, Perlman S (2015). "Coronaviruses: an overview of their replication and pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.). Coronaviruses. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1–23. doi:10.1007/978-1-4939-2438-7_1. ISBN 978-1-4939-2438-7. PMC 4369385. PMID 25720466.
See Table 1.
- Woo, Patrick C. Y.; Huang, Yi; Lau, Susanna K. P.; Yuen, Kwok-Yung (24 August 2010). "Coronavirus Genomics and Bioinformatics Analysis". Viruses. 2 (8): 1804–1820. doi:10.3390/v2081803. ISSN 1999-4915. PMC 3185738. PMID 21994708.
Figure 2. Phylogenetic analysis of RNA-dependent RNA polymerases (Pol) of coronaviruses with complete genome sequences available. The tree was constructed by the neighbor-joining method and rooted using Breda virus polyprotein.
- "Taxonomy browser (Alphacoronavirus 1)". www.ncbi.nlm.nih.gov. Retrieved 29 February 2020.
- Thiel V, ed. (2007). Coronaviruses: Molecular and Cellular Biology (1st ed.). Caister Academic Press. ISBN 978-1-904455-16-5.
- Harris, D. L. Hank. "Transmissible Gastroenteritis in Pigs". Merck Veterinary Manual. Merck. Retrieved 7 July 2019.
- Transboundary and Emerging Diseases of Animals. Iowa State University. 2016. ISBN 978-0-9846270-5-9.
Internal links
External links
- Laude H, Rasschaert D, Delmas B, Godet M, Gelfi J, Charley B (June 1990). "Molecular biology of transmissible gastroenteritis virus". Veterinary Microbiology. 23 (1–4): 147–54. doi:10.1016/0378-1135(90)90144-K. PMC 7117338. PMID 2169670.
- Sola I, Alonso S, Zúñiga S, Balasch M, Plana-Durán J, Enjuanes L (April 2003). "Engineering the Transmissible Gastroenteritis Virus Genome as an Expression Vector Inducing Lactogenic Immunity". Journal of Virology. 77 (7): 4357–69. doi:10.1128/JVI.77.7.4357-4369.2003. PMC 150661. PMID 12634392.
- Tajima M (March 1970). "Morphology of transmissible gastroenteritis virus of pigs". Archives of Virology. 29 (1): 105–8. doi:10.1007/BF01253886. PMC 7086923. PMID 4195092.