Nidovirales
| Nidovirales | |
|---|---|
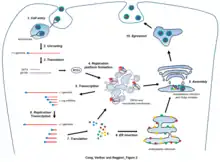 | |
| Life cycle of nidoviruses | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Pisuviricota |
| Class: | Pisoniviricetes |
| Order: | Nidovirales |
Nidovirales is an order of enveloped, positive-strand RNA viruses which infect vertebrates and invertebrates. Host organisms include mammals, birds, reptiles, amphibians, fish, arthropods, molluscs, and helminths.[1] The order includes the families Coronaviridae, Arteriviridae, Roniviridae, and Mesoniviridae.[2]
Member viruses have a viral envelope and a positive-sense, single-stranded RNA genome which is capped and polyadenylated.[3] Nidoviruses are named for the Latin nidus, meaning nest, as all viruses in this order produce a 3' co-terminal nested set of subgenomic mRNAs during infection.[4]
Virology
Structure
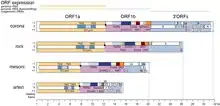
Nidoviruses have a viral envelope and a positive-sense, single-stranded RNA genome which is capped and polyadenylated.[3] The group expresses structural proteins separately from the nonstructural ones. The structural proteins are encoded at the 3’ region of the genome and are expressed from a set of subgenomic mRNAs.
Member viruses encode one main proteinase and between one and three accessory proteinases which are mainly involved in expressing the replicase gene. These proteinases are also responsible for activating or inactivating specific proteins at the correct time in the virus life cycle, ensuring replication occurs at the right time.
Genome
Nidoviruses can be distinguished from other RNA viruses by a constellation of seven conserved domains—5'-TM2-3CLpro-TM3-RdRp-Zm-HEL1-NendoU-3'—with the first three being encoded in ORF1a and the remaining four in ORF1b. TM2 and TM3 and transmembrane domains; RdRp is the RNA-dependent RNA polymerase; Zm is a Zn-cluster binding domain fused with a helicase (HEL1); 3CLpro is a 3C-like protease; and NendoU is an uridylate-specific endonuclease. The 3CLpro has a catalytic His-Cys dyad, and is related to the SARS coronavirus main proteinase (Mpro).
Most, but not all, nidovirus subgenomic RNAs contain a 5′ leader sequence derived from the 5′ end of the genomic RNA. The frameshift that generates ORF1b frameshift occurs at a UUUAAAC heptanucleotide 'slippery' sequence located upstream of the ORF1a stop codon and a putative RNA pseudoknot structure.
Many proteins have been identified on the genomes of Nidovirales, but their function has not yet been determined. Other enzymes that may be present in the genome include papain-like proteases, ADP-ribose/poly(ADP-ribose)-binding or ADP-ribose 1''-phosphate phosphatase activities and cyclic nucleotide phosphodiesterase.
Phylogenetics
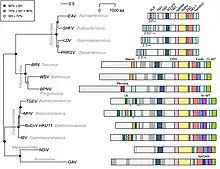
The order Nidovirales can be divided into two clades depending on the size of the genome: those with large genomes (26.3–31.7 kilobases) which included the Coronaviridae and Roniviridae (the large nidoviruses) and those with small genomes (the small nidoviruses)—a clade that includes the distantly related Arteriviridae (12.7–15.7 kb).
The large nidoviruses encode both a 2'-O-methyltransferase and a 3'–5' exoribonuclease (ExoN)—the latter being very unusual for an RNA virus. They also encode a superfamily 1 helicase, uridylate-specific endonuclease (an enzyme unique to nidoviruses) and several proteases.
Nidoviruses as a group have the largest RNA genomes of viruses. Group member planarian secretory cell nidovirus (PSCNV) has the largest known nonsegmented RNA genome of 41.1kb.[6] Its host is the planarian flatworm.[7]
Taxonomy
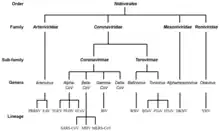
The following suborders and families are recognized (-virineae denotes suborders and -viridae denotes families):[8]
- Abnidovirineae
- Arnidovirineae
- Arteriviridae
- Cremegaviridae
- Gresnaviridae
- Olifoviridae
- Cornidovirineae
- Coronaviridae
- Mesnidovirineae
- Medioniviridae
- Mesoniviridae
- Monidovirineae
- Nanidovirinae
- Nanghoshaviridae
- Nanhypoviridae
- Ronidovirineae
- Euroniviridae
- Roniviridae
- Tornidovirineae
See also
- Animal viruses
- Coronaviruses
References
- ↑ Ogando, Natacha S.; Ferron, Francois; Decroly, Etienne; Canard, Bruno; Posthuma, Clara C.; Snijder, Eric J. (2019). "The Curious Case of the Nidovirus Exoribonuclease: Its Role in RNA Synthesis and Replication Fidelity". Frontiers in Microbiology. 10: 1813. doi:10.3389/fmicb.2019.01813. ISSN 1664-302X. PMC 6693484. PMID 31440227.
- ↑ "International Committee on Taxonomy of Viruses (ICTV)". talk.ictvonline.org. Retrieved 2020-06-08.
- 1 2 King, Andrew M. Q.; Adams, Michael J.; Carstens, Eric B.; Lefkowitz, Elliot J., eds. (2012-01-01), "Order - Nidovirales", Virus Taxonomy, Elsevier: 784–794, doi:10.1016/B978-0-12-384684-6.00066-5, ISBN 978-0-12-384684-6, retrieved 2020-06-08
- ↑ Antoine A.F. de Vries, Marian C. Horzinek, Peter J. M. Rottier, Raoul J. de Groot (1997). "The Genome Organization of the Nidovirales: Similarities and Differences between Arteri-, Toro-, and Coronaviruses". Seminars in Virology. 8 (1): 33–47. CiteSeerX 10.1.1.462.1825. doi:10.1006/smvy.1997.0104. PMC 7128191. PMID 32288441. S2CID 85383257.
{{cite journal}}: CS1 maint: uses authors parameter (link) - ↑ Gulyaeva, Anastasia A.; Gorbalenya, Alexander E. (January 2021). "A nidovirus perspective on SARS-CoV-2". Biochemical and Biophysical Research Communications. 538: 24–34. doi:10.1016/j.bbrc.2020.11.015. PMC 7664520.
- ↑ "Taxonomy browser (Planidovirus 1)". www.ncbi.nlm.nih.gov. Retrieved 2020-06-08.
- ↑ Saberi, Amir; Gulyaeva, Anastasia A.; Brubacher, John L.; Newmark, Phillip A.; Gorbalenya, Alexander E. (2018-11-01). "A planarian nidovirus expands the limits of RNA genome size". PLOS Pathogens. 14 (11): e1007314. doi:10.1371/journal.ppat.1007314. ISSN 1553-7374. PMC 6211748. PMID 30383829.
- ↑ "Virus Taxonomy: 2019 Release". talk.ictvonline.org. International Committee on Taxonomy of Viruses. Retrieved 30 April 2020.
- Ziebuhr, J; Snijder, EJ; Gorbalenya, AE (April 2000). "Virus-encoded proteinases and proteolytic processing in the Nidovirales". The Journal of General Virology. 81 (Pt 4): 853–79. doi:10.1099/0022-1317-81-4-853. PMID 10725411.
External links
- Nidovirales—ICTVdB—The Universal Virus Database, version 4.
- Nidovirales at the US National Library of Medicine Medical Subject Headings (MeSH)
- NIH/MeSH
- "The Nidoviruses" ISBN 0306466341