Evolution of biological complexity
The evolution of biological complexity is one important outcome of the process of evolution.[1] Evolution has produced some remarkably complex organisms – although the actual level of complexity is very hard to define or measure accurately in biology, with properties such as gene content, the number of cell types or morphology all proposed as possible metrics.[2][3][4]
Many biologists used to believe that evolution was progressive (orthogenesis) and had a direction that led towards so-called "higher organisms", despite a lack of evidence for this viewpoint.[5] This idea of "progression" introduced the terms "high animals" and "low animals" in evolution. Many now regard this as misleading, with natural selection having no intrinsic direction and that organisms selected for either increased or decreased complexity in response to local environmental conditions.[6] Although there has been an increase in the maximum level of complexity over the history of life, there has always been a large majority of small and simple organisms and the most common level of complexity appears to have remained relatively constant.
Selection for simplicity and complexity
Usually organisms that have a higher rate of reproduction than their competitors have an evolutionary advantage. Consequently, organisms can evolve to become simpler and thus multiply faster and produce more offspring, as they require fewer resources to reproduce. A good example are parasites such as Plasmodium – the parasite responsible for malaria – and mycoplasma; these organisms often dispense with traits that are made unnecessary through parasitism on a host.[7]
A lineage can also dispense with complexity when a particular complex trait merely provides no selective advantage in a particular environment. Loss of this trait need not necessarily confer a selective advantage, but may be lost due to the accumulation of mutations if its loss does not confer an immediate selective disadvantage.[8] For example, a parasitic organism may dispense with the synthetic pathway of a metabolite where it can readily scavenge that metabolite from its host. Discarding this synthesis may not necessarily allow the parasite to conserve significant energy or resources and grow faster, but the loss may be fixed in the population through mutation accumulation if no disadvantage is incurred by loss of that pathway. Mutations causing loss of a complex trait occur more often than mutations causing gain of a complex trait.
With selection, evolution can also produce more complex organisms. Complexity often arises in the co-evolution of hosts and pathogens,[9] with each side developing ever more sophisticated adaptations, such as the immune system and the many techniques pathogens have developed to evade it. For example, the parasite Trypanosoma brucei, which causes sleeping sickness, has evolved so many copies of its major surface antigen that about 10% of its genome is devoted to different versions of this one gene. This tremendous complexity allows the parasite to constantly change its surface and thus evade the immune system through antigenic variation.[10]
More generally, the growth of complexity may be driven by the co-evolution between an organism and the ecosystem of predators, prey and parasites to which it tries to stay adapted: as any of these become more complex in order to cope better with the diversity of threats offered by the ecosystem formed by the others, the others too will have to adapt by becoming more complex, thus triggering an ongoing evolutionary arms race[9] towards more complexity.[11] This trend may be reinforced by the fact that ecosystems themselves tend to become more complex over time, as species diversity increases, together with the linkages or dependencies between species.
Types of trends in complexity
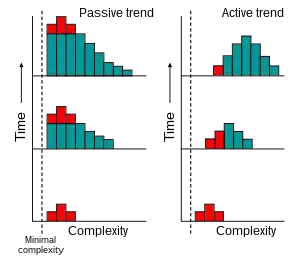
If evolution possessed an active trend toward complexity (orthogenesis), as was widely believed in the 19th century,[12] then we would expect to see an active trend of increase over time in the most common value (the mode) of complexity among organisms.[13]
However, an increase in complexity can also be explained through a passive process.[13] Assuming unbiased random changes of complexity and the existence of a minimum complexity leads to an increase over time of the average complexity of the biosphere. This involves an increase in variance, but the mode does not change. The trend towards the creation of some organisms with higher complexity over time exists, but it involves increasingly small percentages of living things.[4]
In this hypothesis, any appearance of evolution acting with an intrinsic direction towards increasingly complex organisms is a result of people concentrating on the small number of large, complex organisms that inhabit the right-hand tail of the complexity distribution and ignoring simpler and much more common organisms. This passive model predicts that the majority of species are microscopic prokaryotes, which is supported by estimates of 106 to 109 extant prokaryotes[14] compared to diversity estimates of 106 to 3·106 for eukaryotes.[15][16] Consequently, in this view, microscopic life dominates Earth, and large organisms only appear more diverse due to sampling bias.
Genome complexity has generally increased since the beginning of the life on Earth.[17][18] Some computer models have suggested that the generation of complex organisms is an inescapable feature of evolution.[19][20] Proteins tend to become more hydrophobic over time,[21] and to have their hydrophobic amino acids more interspersed along the primary sequence.[22] Increases in body size over time are sometimes seen in what is known as Cope's rule.[23]
Constructive neutral evolution
Recently work in evolution theory has proposed that by relaxing selection pressure, which typically acts to streamline genomes, the complexity of an organism increases by a process called constructive neutral evolution.[24] Since the effective population size in eukaryotes (especially multi-cellular organisms) is much smaller than in prokaryotes,[25] they experience lower selection constraints.
According to this model, new genes are created by non-adaptive processes, such as by random gene duplication. These novel entities, although not required for viability, do give the organism excess capacity that can facilitate the mutational decay of functional subunits. If this decay results in a situation where all of the genes are now required, the organism has been trapped in a new state where the number of genes has increased. This process has been sometimes described as a complexifying ratchet.[26] These supplemental genes can then be co-opted by natural selection by a process called neofunctionalization. In other instances constructive neutral evolution does not promote the creation of new parts, but rather promotes novel interactions between existing players, which then take on new moonlighting roles.[26]
Constructive neutral evolution has also been used to explain how ancient complexes, such as the spliceosome and the ribosome, have gained new subunits over time, how new alternative spliced isoforms of genes arise, how gene scrambling in ciliates evolved, how pervasive pan-RNA editing may have arisen in Trypanosoma brucei, how functional lncRNAs have likely arisen from transcriptional noise, and how even useless protein complexes can persist for millions of years.[24][27][26][28][29][30][31]
Mutational hazard hypothesis
The mutational hazard hypothesis is a non-adaptive theory for increased complexity in genomes.[32] The basis of mutational hazard hypothesis is that each mutation for non-coding DNA imposes a fitness cost.[33] Variation in complexity can be described by 2Neu, where Ne is effective population size and u is mutation rate.[34]
In this hypothesis, selection against non-coding DNA can be reduced in three ways: random genetic drift, recombination rate, and mutation rate.[35] As complexity increases from prokaryotes to multicellular eukaryotes, effective population size decreases, subsequently increasing the strength of random genetic drift.[32] This, along with low recombination rate[35] and high mutation rate,[35] allows non-coding DNA to proliferate without being removed by purifying selection.[32]
Accumulation of non-coding DNA in larger genomes can be seen when comparing genome size and genome content across eukaryotic taxa. There is a positive correlation between genome size and noncoding DNA genome content with each group staying within some variation.[32][33] When comparing variation in complexity in organelles, effective population size is replaced with genetic effective population size (Ng).[34] If looking at silent-site nucleotide diversity, then larger genomes are expected to have less diversity than more compact ones. In plant and animal mitochondria, differences in mutation rate account for the opposite directions in complexity, with plant mitochondria being more complex and animal mitochondria more streamlined.[36]
The mutational hazard hypothesis has been used to at least partially explain expanded genomes in some species. For example, when comparing Volvox cateri to a close relative with a compact genome, Chlamydomonas reinhardtii, the former had less silent-site diversity than the latter in nuclear, mitochondrial, and plastid genomes.[37] However when comparing the plastid genome of Volvox cateri to Volvox africanus, a species in the same genus but with half the plastid genome size, there was high mutation rates in intergenic regions.[38] In Arabiopsis thaliana, the hypothesis was used as a possible explanation for intron loss and compact genome size. When compared to Arabidopsis lyrata, researchers found a higher mutation rate overall and in lost introns (an intron that is no longer transcribed or spliced) compared to conserved introns.[39]
There are expanded genomes in other species that could not be explained by the mutational hazard hypothesis. For example, the expanded mitochondrial genomes of Silene noctiflora and Silene conica have high mutation rates, lower intron lengths, and more non-coding DNA elements compared to others in the same genus, but there was no evidence for long-term low effective population size.[40] The mitochondrial genomes of Citrullus lanatus and Cucurbita pepo differ in several ways. Citrullus lanatus is smaller, has more introns and duplications, while Cucurbita pepo is larger with more chloroplast and short repeated sequences.[41] If RNA editing sites and mutation rate lined up, then Cucurbita pepo would have a lower mutation rate and more RNA editing sites. However the mutation rate is four times higher than Citrullus lanatus and they have a similar number of RNA editing sites.[41] There was also an attempt to use the hypothesis to explain large nuclear genomes of salamanders, but researchers found opposite results than expected, including lower long-term strength of genetic drift.[42]
History
In the 19th century, some scientists such as Jean-Baptiste Lamarck (1744–1829) and Ray Lankester (1847–1929) believed that nature had an innate striving to become more complex with evolution. This belief may reflect then-current ideas of Hegel (1770–1831) and of Herbert Spencer (1820–1903) which envisaged the universe gradually evolving to a higher, more perfect state.
This view regarded the evolution of parasites from independent organisms to a parasitic species as "devolution" or "degeneration", and contrary to nature. Social theorists have sometimes interpreted this approach metaphorically to decry certain categories of people as "degenerate parasites". Later scientists regarded biological devolution as nonsense; rather, lineages become simpler or more complicated according to whatever forms had a selective advantage.[43]
In a 1964 book, The Emergence of Biological Organization, Quastler pioneered a theory of emergence, developing a model of a series of emergences from protobiological systems to prokaryotes without the need to invoke implausible very low probability events.[44]
The evolution of order, manifested as biological complexity, in living systems and the generation of order in certain non-living systems was proposed in 1983 to obey a common fundamental principal called “the Darwinian dynamic”.[45] The Darwinian dynamic was formulated by first considering how microscopic order is generated in simple non-biological systems that are far from thermodynamic equilibrium. Consideration was then extended to short, replicating RNA molecules assumed to be similar to the earliest forms of life in the RNA world. It was shown that the underlying order-generating processes in the non-biological systems and in replicating RNA are basically similar. This approach helped clarify the relationship of thermodynamics to evolution as well as the empirical content of Darwin’s theory.
In 1985 Morowitz[46] noted that the modern era of irreversible thermodynamics ushered in by Lars Onsager in the 1930s showed that systems invariably become ordered under a flow of energy, thus indicating that the existence of life involves no contradiction to the laws of physics.
See also
References
- Werner, Andreas; Piatek, Monica J.; Mattick, John S. (April 2015). "Transpositional shuffling and quality control in male germ cells to enhance evolution of complex organisms". Annals of the New York Academy of Sciences. 1341 (1): 156–163. Bibcode:2015NYASA1341..156W. doi:10.1111/nyas.12608. PMC 4390386. PMID 25557795.
- Adami, C. (2002). "What is complexity?". BioEssays. 24 (12): 1085–94. doi:10.1002/bies.10192. PMID 12447974.
- Waldrop, M.; et al. (2008). "Language: Disputed definitions". Nature. 455 (7216): 1023–1028. doi:10.1038/4551023a. PMID 18948925.
- Longo, Giuseppe; Montévil, Maël (2012-01-01). Dinneen, Michael J.; Khoussainov, Bakhadyr; Nies, André (eds.). Computation, Physics and Beyond. Lecture Notes in Computer Science. Springer Berlin Heidelberg. pp. 289–308. CiteSeerX 10.1.1.640.1835. doi:10.1007/978-3-642-27654-5_22. ISBN 9783642276538. S2CID 16929949.
- McShea, D. (1991). "Complexity and evolution: What everybody knows". Biology and Philosophy. 6 (3): 303–324. doi:10.1007/BF00132234. S2CID 53459994.
- Ayala, F. J. (2007). "Darwin's greatest discovery: design without designer". PNAS. 104 (Suppl 1): 8567–73. Bibcode:2007PNAS..104.8567A. doi:10.1073/pnas.0701072104. PMC 1876431. PMID 17494753.
- Sirand-Pugnet, P.; Lartigue, C.; Marenda, M.; et al. (2007). "Being Pathogenic, Plastic, and Sexual while Living with a Nearly Minimal Bacterial Genome". PLOS Genet. 3 (5): e75. doi:10.1371/journal.pgen.0030075. PMC 1868952. PMID 17511520.
- Maughan, H.; Masel, J.; Birky, W. C.; Nicholson, W. L. (2007). "The roles of mutation accumulation and selection in loss of sporulation in experimental populations of Bacillus subtilis". Genetics. 177 (2): 937–948. doi:10.1534/genetics.107.075663. PMC 2034656. PMID 17720926.
- Dawkins, Richard; Krebs, J. R. (1979). "Arms Races between and within Species". Proceedings of the Royal Society B. 205 (1161): 489–511. Bibcode:1979RSPSB.205..489D. doi:10.1098/rspb.1979.0081. PMID 42057. S2CID 9695900.
- Pays, E. (2005). "Regulation of antigen gene expression in Trypanosoma brucei". Trends Parasitol. 21 (11): 517–20. doi:10.1016/j.pt.2005.08.016. PMID 16126458.
- Heylighen, F. (1999a) "The Growth of Structural and Functional Complexity during Evolution", in F. Heylighen, J. Bollen & A. Riegler (eds.) The Evolution of Complexity Kluwer Academic, Dordrecht, 17–44.
- Ruse, Michael (1996). Monad to man: the Concept of Progress in Evolutionary Biology. Harvard University Press. pp. 526–529 and passim. ISBN 978-0-674-03248-4.
- Carroll SB (2001). "Chance and necessity: the evolution of morphological complexity and diversity". Nature. 409 (6823): 1102–9. Bibcode:2001Natur.409.1102C. doi:10.1038/35059227. PMID 11234024. S2CID 4319886.
- Oren, A. (2004). "Prokaryote diversity and taxonomy: current status and future challenges". Philos. Trans. R. Soc. Lond. B Biol. Sci. 359 (1444): 623–38. doi:10.1098/rstb.2003.1458. PMC 1693353. PMID 15253349.
- May, R. M.; Beverton, R. J. H. (1990). "How Many Species?". Philosophical Transactions of the Royal Society of London. Series B: Biological Sciences. 330 (1257): 293–304. doi:10.1098/rstb.1990.0200.
- Schloss, P.; Handelsman, J. (2004). "Status of the microbial census". Microbiol Mol Biol Rev. 68 (4): 686–91. doi:10.1128/MMBR.68.4.686-691.2004. PMC 539005. PMID 15590780.
- Markov, A. V.; Anisimov, V. A.; Korotayev, A. V. (2010). "Relationship between genome size and organismal complexity in the lineage leading from prokaryotes to mammals". Paleontological Journal. 44 (4): 363–373. doi:10.1134/s0031030110040015. S2CID 10830340.
- Sharov, Alexei A (2006). "Genome increase as a clock for the origin and evolution of life". Biology Direct. 1 (1): 17. doi:10.1186/1745-6150-1-17. PMC 1526419. PMID 16768805.
- Furusawa, C.; Kaneko, K. (2000). "Origin of complexity in multicellular organisms". Phys. Rev. Lett. 84 (26 Pt 1): 6130–3. arXiv:nlin/0009008. Bibcode:2000PhRvL..84.6130F. doi:10.1103/PhysRevLett.84.6130. PMID 10991141. S2CID 13985096.
- Adami, C.; Ofria, C.; Collier, T. C. (2000). "Evolution of biological complexity". PNAS. 97 (9): 4463–8. arXiv:physics/0005074. Bibcode:2000PNAS...97.4463A. doi:10.1073/pnas.97.9.4463. PMC 18257. PMID 10781045.
- Wilson, Benjamin A.; Foy, Scott G.; Neme, Rafik; Masel, Joanna (24 April 2017). "Young genes are highly disordered as predicted by the preadaptation hypothesis of de novo gene birth". Nature Ecology & Evolution. 1 (6): 0146–146. doi:10.1038/s41559-017-0146. PMC 5476217. PMID 28642936.
- Foy, Scott G.; Wilson, Benjamin A.; Bertram, Jason; Cordes, Matthew H. J.; Masel, Joanna (April 2019). "A Shift in Aggregation Avoidance Strategy Marks a Long-Term Direction to Protein Evolution". Genetics. 211 (4): 1345–1355. doi:10.1534/genetics.118.301719. PMC 6456324. PMID 30692195.
- Heim, N. A.; Knope, M. L.; Schaal, E. K.; Wang, S. C.; Payne, J. L. (2015-02-20). "Cope's rule in the evolution of marine animals". Science. 347 (6224): 867–870. Bibcode:2015Sci...347..867H. doi:10.1126/science.1260065. PMID 25700517.
- Stoltzfus, Arlin (1999). "On the Possibility of Constructive Neutral Evolution". Journal of Molecular Evolution. 49 (2): 169–181. Bibcode:1999JMolE..49..169S. CiteSeerX 10.1.1.466.5042. doi:10.1007/PL00006540. ISSN 0022-2844. PMID 10441669. S2CID 1743092.
- Sung, W.; Ackerman, M. S.; Miller, S. F.; Doak, T. G.; Lynch, M. (2012). "Drift-barrier hypothesis and mutation-rate evolution". Proceedings of the National Academy of Sciences. 109 (45): 18488–18492. Bibcode:2012PNAS..10918488S. doi:10.1073/pnas.1216223109. PMC 3494944. PMID 23077252.
- Lukeš, Julius; Archibald, John M.; Keeling, Patrick J.; Doolittle, W. Ford; Gray, Michael W. (2011). "How a neutral evolutionary ratchet can build cellular complexity". IUBMB Life. 63 (7): 528–537. doi:10.1002/iub.489. PMID 21698757. S2CID 7306575.
- Gray, M. W.; Lukes, J.; Archibald, J. M.; Keeling, P. J.; Doolittle, W. F. (2010). "Irremediable Complexity?". Science. 330 (6006): 920–921. Bibcode:2010Sci...330..920G. doi:10.1126/science.1198594. ISSN 0036-8075. PMID 21071654. S2CID 206530279.
- Daniel, Chammiran; Behm, Mikaela; Öhman, Marie (2015). "The role of Alu elements in the cis-regulation of RNA processing". Cellular and Molecular Life Sciences. 72 (21): 4063–4076. doi:10.1007/s00018-015-1990-3. ISSN 1420-682X. PMID 26223268. S2CID 17960570.
- Covello, PatrickS.; Gray, MichaelW. (1993). "On the evolution of RNA editing". Trends in Genetics. 9 (8): 265–268. doi:10.1016/0168-9525(93)90011-6. PMID 8379005.
- Palazzo, Alexander F.; Koonin, Eugene V. (2020). "Functional Long Non-coding RNAs Evolve from Junk Transcripts". Cell. 183 (5): 1151–1161. doi:10.1016/j.cell.2020.09.047. ISSN 0092-8674. PMID 33068526. S2CID 222815635.
- Hochberg, GKA; Liu, Y; Marklund, EG; Metzger, BPH; Laganowsky, A; Thornton, JW (December 2020). "A hydrophobic ratchet entrenches molecular complexes". Nature. 588 (7838): 503–508. Bibcode:2020Natur.588..503H. doi:10.1038/s41586-020-3021-2. PMC 8168016. PMID 33299178.
- Lynch, Michael; Conery, John S. (2003-11-21). "The Origins of Genome Complexity". Science. 302 (5649): 1401–1404. Bibcode:2003Sci...302.1401L. doi:10.1126/science.1089370. ISSN 0036-8075. PMID 14631042. S2CID 11246091.
- Lynch, Michael; Bobay, Louis-Marie; Catania, Francesco; Gout, Jean-François; Rho, Mina (2011-09-22). "The Repatterning of Eukaryotic Genomes by Random Genetic Drift". Annual Review of Genomics and Human Genetics. 12 (1): 347–366. doi:10.1146/annurev-genom-082410-101412. ISSN 1527-8204. PMC 4519033. PMID 21756106.
- Lynch, M. (2006-03-24). "Mutation Pressure and the Evolution of Organelle Genomic Architecture". Science. 311 (5768): 1727–1730. Bibcode:2006Sci...311.1727L. doi:10.1126/science.1118884. ISSN 0036-8075. PMID 16556832. S2CID 2678365.
- Lynch, Michael (2006-02-01). "The Origins of Eukaryotic Gene Structure". Molecular Biology and Evolution. 23 (2): 450–468. doi:10.1093/molbev/msj050. ISSN 1537-1719. PMID 16280547.
- Lynch, Michael (2006-10-13). "Streamlining and Simplification of Microbial Genome Architecture". Annual Review of Microbiology. 60 (1): 327–349. doi:10.1146/annurev.micro.60.080805.142300. ISSN 0066-4227. PMID 16824010.
- Smith, D. R.; Lee, R. W. (2010-10-01). "Low Nucleotide Diversity for the Expanded Organelle and Nuclear Genomes of Volvox carteri Supports the Mutational-Hazard Hypothesis". Molecular Biology and Evolution. 27 (10): 2244–2256. doi:10.1093/molbev/msq110. ISSN 0737-4038. PMID 20430860.
- Gaouda, Hager; Hamaji, Takashi; Yamamoto, Kayoko; Kawai-Toyooka, Hiroko; Suzuki, Masahiro; Noguchi, Hideki; Minakuchi, Yohei; Toyoda, Atsushi; Fujiyama, Asao; Nozaki, Hisayoshi; Smith, David Roy (2018-09-01). Chaw, Shu-Miaw (ed.). "Exploring the Limits and Causes of Plastid Genome Expansion in Volvocine Green Algae". Genome Biology and Evolution. 10 (9): 2248–2254. doi:10.1093/gbe/evy175. ISSN 1759-6653. PMC 6128376. PMID 30102347.
- Yang, Yu-Fei; Zhu, Tao; Niu, Deng-Ke (April 2013). "Association of Intron Loss with High Mutation Rate in Arabidopsis: Implications for Genome Size Evolution". Genome Biology and Evolution. 5 (4): 723–733. doi:10.1093/gbe/evt043. ISSN 1759-6653. PMC 4104619. PMID 23516254.
- Sloan, Daniel B.; Alverson, Andrew J.; Chuckalovcak, John P.; Wu, Martin; McCauley, David E.; Palmer, Jeffrey D.; Taylor, Douglas R. (2012-01-17). Gray, Michael William (ed.). "Rapid Evolution of Enormous, Multichromosomal Genomes in Flowering Plant Mitochondria with Exceptionally High Mutation Rates". PLOS Biology. 10 (1): e1001241. doi:10.1371/journal.pbio.1001241. ISSN 1545-7885. PMC 3260318. PMID 22272183.
- Alverson, Andrew J; Wei, XioXin; Rice, Danny W; Stern, David B; Barry, Kerrie; Palmer, Jeffrey D (2010-01-29). "Insights into the Evolution of Mitochondrial Genome Size from Complete Sequences of Citrus lanatus and Cucurbita pepo (Cucurbitaceae)". Molecular Biology and Evolution. 27 (6): 1436–1448. doi:10.1093/molbev/msq029. PMC 2877997. PMID 20118192.
- Mohlhenrich, Erik Roger; Lockridge Mueller, Rachel (2016-09-27). "Genetic drift and mutational hazard in the evolution of salamander genomic gigantism". Evolution. 70 (12): 2865–2878. doi:10.1111/evo.13084. hdl:10217/173461. PMID 27714793. S2CID 205125025 – via JSTOR.
- Dougherty, Michael J. (July 1998). "Is the human race evolving or devolving?". Scientific American.
From a biological perspective, there is no such thing as devolution. All changes in the gene frequencies of populations—and quite often in the traits those genes influence—are by definition evolutionary changes. [...] When species do evolve, it is not out of need but rather because their populations contain organisms with variants of traits that offer a reproductive advantage in a changing environment.
- Quastler, H. (1964) The Emergence of Biological Organization. Yale University Press
- Bernstein H, Byerly HC, Hopf FA, Michod RA, Vemulapalli GK. (1983) The Darwinian Dynamic. Quarterly Review of Biology 58, 185-207. JSTOR 2828805
- Morowitz HJ. (1985) Mayonnaise and the origin of life. (Berkley Books, NY)