Hsp90
Hsp90 (heat shock protein 90) is a chaperone protein that assists other proteins to fold properly, stabilizes proteins against heat stress, and aids in protein degradation. It also stabilizes a number of proteins required for tumor growth, which is why Hsp90 inhibitors are investigated as anti-cancer drugs.
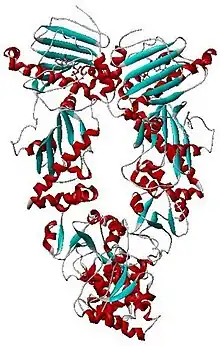
| Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| Symbol | HATPase_c | ||||||||
| Pfam | PF02518 | ||||||||
| Pfam clan | CL0025 | ||||||||
| InterPro | IPR003594 | ||||||||
| SMART | SM00387 | ||||||||
| SCOP2 | 1ei1 / SCOPe / SUPFAM | ||||||||
| |||||||||
| Hsp90 protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
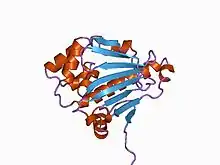 Structure of the N-terminal domain of the yeast Hsp90 chaperone.[2] | |||||||||
| Identifiers | |||||||||
| Symbol | Hsp90 | ||||||||
| Pfam | PF00183 | ||||||||
| InterPro | IPR020576 | ||||||||
| PROSITE | PDOC00270 | ||||||||
| SCOP2 | 1ah6 / SCOPe / SUPFAM | ||||||||
| |||||||||
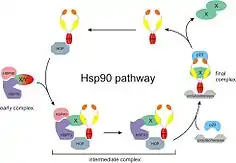
Heat shock proteins, as a class, are among the most highly expressed cellular proteins across all species.[3] As their name implies, heat shock proteins protect cells when stressed by elevated temperatures. They account for 1–2% of total protein in unstressed cells. However, when cells are heated, the fraction of heat shock proteins increases to 4–6% of cellular proteins.[4]
Heat shock protein 90 (Hsp90) is one of the most common of the heat-related proteins. The "90" comes from the fact that it has a mass of roughly 90 kilodaltons. A 90 kDa protein is considered fairly large for a non-fibrous protein. Hsp90 is found in bacteria and all branches of eukarya, but it is apparently absent in archaea.[5] Whereas cytoplasmic Hsp90 is essential for viability under all conditions in eukaryotes, the bacterial homologue HtpG is dispensable under non-heat stress conditions.[6]
This protein was first isolated by extracting proteins from cells stressed by heating, dehydrating or by other means, all of which caused the cell's proteins to begin to denature.[7] However it was later discovered that Hsp90 also has essential functions in unstressed cells.
Isoforms
Hsp90 is highly conserved and expressed in a variety of different organisms from bacteria to mammals – including the prokaryotic analogue HtpG (high-temperature protein G) with 40% sequence identity and 55% similarity to the human protein.[5] Yeast Hsp90 is 60% identical to human Hsp90α.
In mammalian cells, there are two or more genes encoding cytosolic Hsp90 homologues,[5] with the human Hsp90α showing 85% sequence identity to Hsp90β.[8] The α- and the β-forms are thought to be the result of a gene duplication event that occurred millions of years ago.[5]
The five functional human genes encoding Hsp90 protein isoforms are listed below:[8]
| family | subcellular location |
subfamily | gene | protein |
|---|---|---|---|---|
| HSP90A | cytosolic | HSP90AA (inducible) |
HSP90AA1 | Hsp90-α1 |
| HSP90AA2 | Hsp90-α2 | |||
| HSP90AB (constitutively expressed) |
HSP90AB1 | Hsp90-β | ||
| HSP90B | endoplasmic reticulum |
HSP90B1 | Endoplasmin/ GRP-94 | |
| TRAP | mitochondrial | TRAP1 | TNF Receptor- Associated Protein 1 |
There are 12 human pseudogenes (non-functional genes) that encode additional Hsp90 isoforms that are not expressed as proteins.
A membrane-associated variant of cytosolic Hsp90, lacking an ATP-binding site, has recently been identified and was named Hsp90N.[9] This HSP90α-Δ-N transcript is a chimera, with the first 105 bp of the coding sequence derived from the CD47 gene on chromosome 3q13.2, and the remaining coding sequence derived from HSP90AA1.[8] However, gene-encoding Hsp90N was later proven to be non-existent in human genome. It is possibly a cloning artifact or a product of chromosomal rearrangement occurring in a single cell line.[10]
Structure
Common features
The overall structure of Hsp90 is similar to that of other proteins in that it contains all of the common secondary structural elements (i.e., alpha helixes, beta pleated sheets, and random coils). Being a cytoplasmic protein requires that the protein be globular in structure, that is largely non-polar on the inside and polar on the outside, so as to be dissolved by water. Hsp90 contains nine helices and eight anti-parallel beta pleated sheets, which combine to form several alpha/beta sandwiches. The 310 helices make up approximately 11% of the protein's amino acid residues, which is much higher than the average 4% in other proteins.[11]
Domain structure
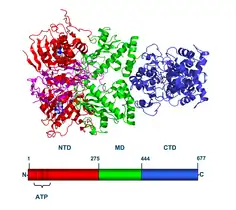
Hsp90 consists of four structural domains:[12][13][14]
- a highly conserved N-terminal domain (NTD) of ~25 kDa
- a "charged linker" region, that connects the N-terminus with the middle domain
- a middle domain (MD) of ~40 kDa
- a C-terminal domain (CTD) of ~12 kDa.
Crystal structures are available for the N-terminal domain of yeast and human Hsp90,[15][16][17] for complexes of the N-terminus with inhibitors and nucleotides,[15][16] and for the middle domain of yeast Hsp90.[18] Recently structures for full length Hsp90 from E. coli (2IOP, 2IOQ),[19] yeast (2CG9, 2CGE),[20] and the dog endoplasmic reticulum (2O1U, 2O1V)[21] were elucidated.[22]
Hsp90 forms homodimers where the contact sites are localized within the C-terminus in the open conformation of the dimer. The N-termini also come in contact in the closed conformation of the dimer.[18]
N-terminal domain
The N-terminal domain shows homology not only among members of the Hsp90 chaperone family but also to members of the ATPase/kinase GHKL (Gyrase, Hsp90, Histidine Kinase, MutL) superfamily.[13]
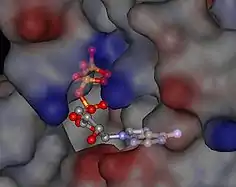
A common binding pocket for ATP and the inhibitor geldanamycin is situated in the N-terminal domain.[15][16] Amino acids that are directly involved in the interaction with ATP are Leu34, Asn37, Asp79, Asn92, Lys98, Gly121, and Phe124. In addition, Mg2+ and several water molecules form bridging electrostatic and hydrogen bonding interactions, respectively, between Hsp90 and ATP. In addition, Glu33 is required for ATP hydrolysis.
Middle domain
The middle domain is divided into three regions:
- a 3-layer α-β-α sandwich
- a 3-turn α-helix and irregular loops
- a 6-turn α-helix.[13]
The MD is also involved in client protein binding. For example, proteins known to interact this the Hsp90 MD include PKB/Akt1, eNOS,[23][24] Aha1, Hch1. Furthermore, substrate binding (e.g., by Aha1 and Hch1) to the MD is also known to increase the ATPase activity of Hsp90.[18][25]
C-terminal domain
The C-terminal domain possesses an alternative ATP-binding site, which becomes accessible when the N-terminal Bergerat pocket is occupied.[26][27]
At the very C-terminal end of the protein is the tetratricopeptide repeat (TPR) motif recognition site, the conserved MEEVD pentapeptide, that is responsible for the interaction with co-factors such as the immunophilins FKBP51 and FKBP52, the stress induced phosphoprotein 1 (Sti1/Hop), cyclophilin-40, PP5, Tom70, and many more.[28]
Mechanism
The Hsp90 protein contains three functional domains, the ATP-binding, protein-binding, and dimerizing domain, each of which playing a crucial role in the function of the protein.
ATP binding
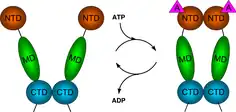
The region of the protein near the N-terminus has a high-affinity ATP-binding site. The ATP binds to a sizable cleft in the side of protein, which is 15 Å (1.5 nanometres) deep. This cleft has a high affinity for ATP, and when given a suitable protein substrate, Hsp90 cleaves the ATP into ADP and Pi. Direct inhibitors of ATP binding or allosteric inhibitors of either ATP binding or ATPase activity can block Hsp90 function.[11] Another interesting feature of the ATP-binding region of Hsp90 is that it has a “lid” that is open during the ADP-bound state and closed in the ATP-bound state.[29] In the open conformation, the lid has no intraprotein interaction, and when closed comes into contact with several residues.[30] The contribution of this lid to the activity of Hsp90 has been probed with site-directed mutagenesis. The Ala107Asp mutant stabilizing the closed conformation of the protein through the formation of additional hydrogen bonds substantially increases ATPase activity while leaving the AMP+PnP conformation unchanged.[30]
The ATPase-binding region of Hsp90 is currently under intense study, because it is the principal binding site of drugs targeting this protein.[31] Antitumor drugs targeting this section of Hsp90 include the antibiotics geldanamycin,[11][32] herbimycin, radicicol, deguelin,[33] derrubone,[34] macbecin,[35] and beta-lactams.[36]
Protein binding
The protein-binding region of Hsp90 is located toward the C-terminus of the amino sequence. The Hsp90 protein can adopt two major conformational states. The first is an open ATP-bound state and the second is a closed ADP-bound state. Thus, ATP hydrolysis drives what is commonly referred to as a “pincer-type” conformational change in the protein binding site.[37]
Hsp90, while in the open conformation, leaves some hydrophobic residues exposed, to which unfolded and misfolded proteins that have unusual hydrophobic regions exposed are recruited with high affinity.[38] When a bound substrate is in place, the energy-releasing ATP hydrolysis by the ATPase function near the N-terminal domain forces conformational changes that clamp the Hsp90 down onto the substrate.[30] In a reaction similar to that of other molecular clamp proteins like GyrB and MutL, this site drives virtually all of the protein folding functions that Hsp90 plays a role in. In contrast, MutL and GyrB function as topoisomerases and use a charge clamp with a high amount of positively charged sidechains that is electrostatically attracted to the negative backbone of DNA.[39]
The ability of Hsp90 to clamp onto proteins allows it to perform several functions including assisting folding, preventing aggregation, and facilitating transport.
Function
Normal cells
In unstressed cells, Hsp90 plays a number of important roles, which include assisting folding, intracellular transport, maintenance, and degradation of proteins as well as facilitating cell signaling.
Protein folding and role as chaperone
Hsp90 is known to associate with the non-native structures of many proteins, which has led to the proposal that Hsp90 is involved in protein folding in general.[40] Furthermore, Hsp90 has been shown to suppress the aggregation of a wide range of "client" or "substrate" proteins and hence acts as a general protective chaperone.[41][42][43] However Hsp90 is somewhat more selective than other chaperones.[44]
Protein degradation
Eukaryotic proteins that are no longer needed or are misfolded or otherwise damaged are usually marked for destruction by the polyubiquitation pathway. These ubiquitinated proteins are recognized and degraded by the 26S proteasome.[45][46] Hence the 26S proteasome is an integral part of the cell's mechanism to degrade proteins. Furthermore, a constant supply of functional Hsp90 is needed to maintain the tertiary structure of the proteasome.[47] Finally experiments done with heat sensitive Hsp90 mutants and the 26S proteasome suggest that Hsp90 is responsible for most, if not all, of the ATPase activity of the proteasome.[45]
Interaction with steroid receptors
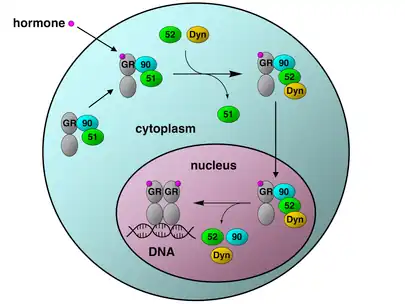
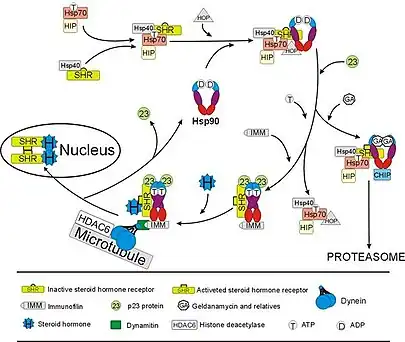
The glucocorticoid receptor (GR) is the most thoroughly studied example of a steroid receptor whose function is crucially dependent on interactions with Hsp90.[50][51] In the absence of the steroid hormone cortisol, GR resides in the cytosol complexed with several chaperone proteins including Hsp90 (see figure to the right). These chaperones maintain the GR in a state capable of binding hormone. A second role of Hsp90 is to bind immunophilins (e.g., FKBP52) that attach the GR complex to the dynein protein trafficking pathway, which translocates the activated receptor from the cytoplasm into the nucleus.[52] Once in the nucleus, the GR dimerizes and binds to specific sequences of DNA and thereby upregulates the expression of GR responsive genes. Hsp90 is also required for the proper functioning of several other steroid receptors, including those responsible for the binding of aldosterone,[53] androgen,[54] estrogen,[55] and progesterone.[56]
Cancerous cells
Cancerous cells overexpress a number of proteins, including growth factor receptors, such as EGFR,[57] or signal transduction proteins such as PI3K and AKT (Inhibition of these proteins may trigger apoptosis). Hsp90 stabilizes various growth factor receptors[58] and some signaling molecules including PI3K and AKT proteins. Hence inhibition of Hsp90 downregulates the PI3K/AKT pathway leading to downregulation of the anti-apoptotic protein Bcl-w resulting in apoptosis of cancerous and senescent cells.[59][15][60]
Interestingly, the disruption of HSP90 with nano-therapeutics has been implicated in targeting drug-induced resistance and relieves the suppression of Natural Killer (NK) immune cells in breast cancer.[61] Another important role of Hsp90 in cancer is the stabilization of mutant proteins such as v-Src, the fusion oncogene Bcr/Abl, and mutant forms of p53 that appear during cell transformation. It appears that Hsp90 can act as a "protector" of less stable proteins produced by DNA mutations.[62]
Hsp90 is also required for induction of vascular endothelial growth factor (VEGF) and nitric oxide synthase (NOS).[24] Both are important for de novo angiogenesis that is required for tumour growth beyond the limit of diffusion distance of oxygen in tissues.[62] It also promotes the invasion step of metastasis by assisting the matrix metalloproteinase MMP2.[63] Together with its co-chaperones, Hsp90 modulates tumour cell apoptosis "mediated through effects on AKT,[23] tumor necrosis factor receptors (TNFR) and nuclear factor-κB (NF-κB) function.".[64] Also, Hsp90 participates in many key processes in oncogenesis such as self-sufficiency in growth signals, stabilization of mutant proteins, angiogenesis, and metastasis.
Clinical significance
Hsp90 plays apparently conflicting roles in the cell, as it is essential for both the creation and the maintenance as well as the destruction of proteins. Its normal function is critical to maintaining the health of cells, whereas its dysregulation may contribute to carcinogenesis. The ability of this chaperone to both stabilize the 26S proteasome (which enables the cell to degrade unwanted and/or harmful proteins) and to stabilize kinases against the same proteasome demonstrates its functional diversity. The uses of Hsp90 inhibitors in cancer treatment highlight Hsp90's importance as a therapeutic target.[65]
Targeting Hsp90 with drugs has shown promising effects in clinical trials. For example, the Hsp90 inhibitor geldanamycin has been used as an anti-tumor agent.[11] The drug was originally thought to function as a kinase inhibitor but was subsequently shown to be an Hsp90 inhibitor where it uses a compact conformation to insert itself into the ATP binding site.[11]
HSP90 beta has been identified as one of the autoantigenic biomarkers and targets involved in human ovarian autoimmune disease leading to ovarian failure and thereby infertility.[66]
Prediction and validation of the immunodominant epitope/s of HSP90 beta protein has been demonstrated using sera from infertile women having anti-HSP90 autoantibodies. The decapeptide EP6 (380-389)is a major immunogenic epitope of HSP90 followed by EP1 (1-12) and EP8 (488-498). Knowledge of binding epitopes on the autoantigen is necessary to understand the subsequent pathologic events. Predicted 3D structures of these peptides demonstrated that they exist in the loop conformation, which is the most mobile part of the protein. Also, analysis of the sequences of HSP90 beta across several species reveals that EP6 peptide forms a part of a well-conserved motif. A polyclonal antibody generated to the immunodominant epitope- EP6 confirms similar biochemical and cellular immunoreactivity as seen with the patients' sera with anti-HSP90 autoantibodies. The study might generate new tools for the detection of disease-inducing epitopes and a possible therapeutic intervention.[67]
Evolution
Sequence alignments of Hsp90 have shown the protein to have about 40% sequence identity across all homologs, indicating that it is a highly conserved protein. There are two homologs, found in the cytosol and endoplasmic reticulum respectively. The presence of these two homologs was likely caused by a gene duplication event very early in the evolution of eukaryotes that may have accompanied the evolution of the endoplasmic reticulum or the nucleus. This inference is supported by the fact that the duplication is found in Giardia lamblia, one of the earliest branching eukaryotic species. At least 2 other subsequent gene duplications occurred, which explains the different forms of Hsp90 found in fungi and vertebrates. One divergence produced cognate and heat-induced forms of Hsp90 in Saccharomyces cerevisiae, while the second gene duplication event in the cytosolic branch produced the alpha and beta subfamilies of sequences that are found in all vertebrates. In a phylogenetic tree based on Hsp90 sequences, it was found that plants and animals are more closely related to each other than to fungi.[68] Similar to the Hsp90 protein, the gene for Hsp70 protein also underwent duplication at a very early stage in the formation of eukaryotic cells and the homologs in the cytosol and endoplasmic reticulum resulted from this gene duplication event.[69] These gene duplication events are important in terms of the origin of the eukaryotic cell and of the endoplasmic reticulum.[70][71]
References
- PDB: 2CG9; Ali MM, Roe SM, Vaughan CK, Meyer P, Panaretou B, Piper PW, Prodromou C, Pearl LH (April 2006). "Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex". Nature. 440 (7087): 1013–7. doi:10.1038/nature04716. PMC 5703407. PMID 16625188.
- Prodromou C, Roe SM, Piper PW, Pearl LH (June 1997). "A molecular clamp in the crystal structure of the N-terminal domain of the yeast Hsp90 chaperone". Nat. Struct. Biol. 4 (6): 477–82. doi:10.1038/nsb0697-477. PMID 9187656. S2CID 38764610.
- Csermely P, Schnaider T, Soti C, Prohászka Z, Nardai G (August 1998). "The 90-kDa molecular chaperone family: structure, function, and clinical applications. A comprehensive review". Pharmacol. Ther. 79 (2): 129–68. doi:10.1016/S0163-7258(98)00013-8. PMID 9749880.
- Crevel G, Bates H, Huikeshoven H, Cotterill S (1 June 2001). "The Drosophila Dpit47 protein is a nuclear Hsp90 co-chaperone that interacts with DNA polymerase alpha". J. Cell Sci. 114 (Pt 11): 2015–25. doi:10.1242/jcs.114.11.2015. PMID 11493638.
- Chen B, Zhong D, Monteiro A (2006). "Comparative genomics and evolution of the HSP90 family of genes across all kingdoms of organisms". BMC Genomics. 7: 156. doi:10.1186/1471-2164-7-156. PMC 1525184. PMID 16780600.
- Thomas JG, Baneyx F (October 1998). "Roles of the Escherichia coli Small Heat Shock Proteins IbpA and IbpB in Thermal Stress Management: Comparison with ClpA, ClpB, and HtpG In Vivo". J. Bacteriol. 180 (19): 5165–72. doi:10.1128/JB.180.19.5165-5172.1998. PMC 107554. PMID 9748451.
- Prodromou C, Panaretou B, Chohan S, Siligardi G, O'Brien R, Ladbury JE, Roe SM, Piper PW, Pearl LH (August 2000). "The ATPase cycle of Hsp90 drives a molecular 'clamp' via transient dimerization of the N-terminal domains". EMBO J. 19 (16): 4383–92. doi:10.1093/emboj/19.16.4383. PMC 302038. PMID 10944121.
- Chen B, Piel WH, Gui L, Bruford E, Monteiro A (December 2005). "The HSP90 family of genes in the human genome: insights into their divergence and evolution". Genomics. 86 (6): 627–37. doi:10.1016/j.ygeno.2005.08.012. PMID 16269234.
- Grammatikakis N, Vultur A, Ramana CV, Siganou A, Schweinfest CW, Watson DK, Raptis L (March 2002). "The role of Hsp90N, a new member of the Hsp90 family, in signal transduction and neoplastic transformation". J. Biol. Chem. 277 (10): 8312–20. doi:10.1074/jbc.M109200200. PMID 11751906.
- Zurawska A, Urbanski J, Bieganowski P (November 2008). "Hsp90n - An accidental product of a fortuitous chromosomal translocation rather than a regular Hsp90 family member of human proteome". Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics. 1784 (11): 1844–6. doi:10.1016/j.bbapap.2008.06.013. PMID 18638579.
- Goetz MP, Toft DO, Ames MM, Erlichman C (August 2003). "The Hsp90 chaperone complex as a novel target for cancer therapy". Ann. Oncol. 14 (8): 1169–76. doi:10.1093/annonc/mdg316. PMID 12881371.
- Pearl LH, Prodromou C (February 2000). "Structure and in vivo function of Hsp90". Curr. Opin. Struct. Biol. 10 (1): 46–51. doi:10.1016/S0959-440X(99)00047-0. PMID 10679459.
- Prodromou C, Pearl LH (October 2003). "Structure and functional relationships of Hsp90". Curr Cancer Drug Targets. 3 (5): 301–23. doi:10.2174/1568009033481877. PMID 14529383.
- Pearl LH, Prodromou C (2001). "Structure, function, and mechanism of the Hsp90 molecular chaperone". Protein Folding in the Cell. pp. 157–86. doi:10.1016/S0065-3233(01)59005-1. ISBN 978-0-12-034259-4. PMID 11868271.
{{cite book}}:|journal=ignored (help) - Stebbins CE, Russo AA, Schneider C, Rosen N, Hartl FU, Pavletich NP (April 1997). "Crystal structure of an Hsp90-geldanamycin complex: targeting of a protein chaperone by an antitumor agent". Cell. 89 (2): 239–50. doi:10.1016/S0092-8674(00)80203-2. PMID 9108479.
- Prodromou C, Roe SM, O'Brien R, Ladbury JE, Piper PW, Pearl LH (July 1997). "Identification and structural characterization of the ATP/ADP-binding site in the Hsp90 molecular chaperone". Cell. 90 (1): 65–75. doi:10.1016/S0092-8674(00)80314-1. PMID 9230303.
- Prodromou C, Roe SM, Piper PW, Pearl LH (June 1997). "A molecular clamp in the crystal structure of the N-terminal domain of the yeast Hsp90 chaperone". Nat. Struct. Biol. 4 (6): 477–82. doi:10.1038/nsb0697-477. PMID 9187656. S2CID 38764610.
- Meyer P, Prodromou C, Hu B, Vaughan C, Roe SM, Panaretou B, Piper PW, Pearl LH (March 2003). "Structural and functional analysis of the middle segment of hsp90: implications for ATP hydrolysis and client protein and cochaperone interactions" (PDF). Mol. Cell. 11 (3): 647–58. doi:10.1016/S1097-2765(03)00065-0. PMID 12667448.
- Shiau AK, Harris SF, Southworth DR, Agard DA (October 2006). "Structural Analysis of E. coli hsp90 reveals dramatic nucleotide-dependent conformational rearrangements". Cell. 127 (2): 329–40. doi:10.1016/j.cell.2006.09.027. PMID 17055434.
- Ali MM, Roe SM, Vaughan CK, Meyer P, Panaretou B, Piper PW, Prodromou C, Pearl LH (April 2006). "Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex". Nature. 440 (7087): 1013–7. doi:10.1038/nature04716. PMC 5703407. PMID 16625188.
- Dollins DE, Warren JJ, Immormino RM, Gewirth DT (October 2007). "Structures of GRP94-nucleotide complexes reveal mechanistic differences between the hsp90 chaperones". Mol. Cell. 28 (1): 41–56. doi:10.1016/j.molcel.2007.08.024. PMC 2094010. PMID 17936703.
- Wandinger SK, Richter K, Buchner J (July 2008). "The hsp90 chaperone machinery". J. Biol. Chem. 283 (27): 18473–7. doi:10.1074/jbc.R800007200. PMID 18442971.
- Sato S, Fujita N, Tsuruo T (September 2000). "Modulation of Akt kinase activity by binding to Hsp90". Proc. Natl. Acad. Sci. U.S.A. 97 (20): 10832–7. doi:10.1073/pnas.170276797. PMC 27109. PMID 10995457.
- Fontana J, Fulton D, Chen Y, Fairchild TA, McCabe TJ, Fujita N, Tsuruo T, Sessa WC (May 2002). "Domain mapping studies reveal that the M domain of hsp90 serves as a molecular scaffold to regulate Akt-dependent phosphorylation of endothelial nitric oxide synthase and NO release". Circ. Res. 90 (8): 866–73. doi:10.1161/01.RES.0000016837.26733.BE. PMID 11988487.
- Panaretou B, Siligardi G, Meyer P, Maloney A, Sullivan JK, Singh S, Millson SH, Clarke PA, Naaby-Hansen S, Stein R, Cramer R, Mollapour M, Workman P, Piper PW, Pearl LH, Prodromou C (December 2002). "Activation of the ATPase activity of hsp90 by the stress-regulated cochaperone aha1" (PDF). Mol. Cell. 10 (6): 1307–18. doi:10.1016/S1097-2765(02)00785-2. PMID 12504007.
- Marcu MG, Chadli A, Bouhouche I, Catelli M, Neckers LM (November 2000). "The heat shock protein 90 antagonist novobiocin interacts with a previously unrecognized ATP-binding domain in the carboxyl terminus of the chaperone". J. Biol. Chem. 275 (47): 37181–6. doi:10.1074/jbc.M003701200. PMID 10945979.
- Söti C, Rácz A, Csermely P (March 2002). "A Nucleotide-dependent molecular switch controls ATP binding at the C-terminal domain of Hsp90. N-terminal nucleotide binding unmasks a C-terminal binding pocket". J. Biol. Chem. 277 (9): 7066–75. doi:10.1074/jbc.M105568200. PMID 11751878.
- Young JC, Obermann WM, Hartl FU (July 1998). "Specific binding of tetratricopeptide repeat proteins to the C-terminal 12-kDa domain of hsp90". J. Biol. Chem. 273 (29): 18007–10. doi:10.1074/jbc.273.29.18007. PMID 9660753.
- Didenko T, Duarte AM, Karagöz GE, Rüdiger SG (March 2012). "Hsp90 structure and function studied by NMR spectroscopy". Biochim. Biophys. Acta. 1823 (3): 636–47. doi:10.1016/j.bbamcr.2011.11.009. PMID 22155720.
- Wegele H, Müller L, Buchner J (2004). Hsp70 and Hsp90--a relay team for protein folding. pp. 1–44. doi:10.1007/s10254-003-0021-1. ISBN 978-3-540-22096-1. PMID 14740253.
{{cite book}}:|journal=ignored (help) - Chiosis G, Caldas Lopes E, Solit D (June 2006). "Heat shock protein-90 inhibitors: a chronicle from geldanamycin to today's agents". Curr Opin Investig Drugs. 7 (6): 534–41. PMID 16784024.
- Pratt WB, Toft DO (1 February 2003). "Regulation of signaling protein function and trafficking by the hsp90/hsp70-based chaperone machinery". Exp. Biol. Med. (Maywood). 228 (2): 111–33. CiteSeerX 10.1.1.334.341. doi:10.1177/153537020322800201. PMID 12563018. S2CID 9162123.
- Oh SH, Woo JK, Yazici YD, Myers JN, Kim WY, Jin Q, Hong SS, Park HJ, Suh YG, Kim KW, Hong WK, Lee HY (June 2007). "Structural basis for depletion of heat shock protein 90 client proteins by deguelin". J. Natl. Cancer Inst. 99 (12): 949–61. doi:10.1093/jnci/djm007. PMID 17565155.
- Hadden MK, Galam L, Gestwicki JE, Matts RL, Blagg BS (December 2007). "Derrubone, an inhibitor of the Hsp90 protein folding machinery". J. Nat. Prod. 70 (12): 2014–8. doi:10.1021/np070190s. PMID 18020309.
- Martin CJ, Gaisser S, Challis IR, Carletti I, Wilkinson B, Gregory M, Prodromou C, Roe SM, Pearl LH, Boyd SM, Zhang MQ (May 2008). "Molecular characterization of macbecin as an Hsp90 inhibitor". J. Med. Chem. 51 (9): 2853–7. doi:10.1021/jm701558c. PMID 18357975.
- O'Boyle NM, Knox AJ, Price TT, Williams DC, Zisterer DM, Lloyd DG, Meegan MJ (October 2011). "Lead identification of β-lactam and related imine inhibitors of the molecular chaperone heat shock protein 90". Bioorg. Med. Chem. 19 (20): 6055–68. doi:10.1016/j.bmc.2011.08.048. PMID 21920765.
- Grenert JP, Sullivan WP, Fadden P, Haystead TA, Clark J, Mimnaugh E, Krutzsch H, Ochel HJ, Schulte TW, Sausville E, Neckers LM, Toft DO (September 1997). "The amino-terminal domain of heat shock protein 90 (hsp90) that binds geldanamycin is an ATP/ADP switch domain that regulates hsp90 conformation". J. Biol. Chem. 272 (38): 23843–50. doi:10.1074/jbc.272.38.23843. PMID 9295332.
- Xu Z, Horwich AL, Sigler PB (August 1997). "The crystal structure of the asymmetric GroEL-GroES-(ADP)7 chaperonin complex". Nature. 388 (6644): 741–50. doi:10.1038/41944. PMID 9285585. S2CID 19423648.
- Kampranis SC, Bates AD, Maxwell A (July 1999). "A model for the mechanism of strand passage by DNA gyrase". Proc. Natl. Acad. Sci. U.S.A. 96 (15): 8414–9. doi:10.1073/pnas.96.15.8414. PMC 17530. PMID 10411889.
- Buchner J (April 1999). "Hsp90 & Co. - a holding for folding". Trends Biochem. Sci. 24 (4): 136–41. doi:10.1016/S0968-0004(99)01373-0. PMID 10322418.
- Miyata Y, Yahara I (April 1992). "The 90-kDa heat shock protein, Hsp90, binds and protects casein kinase II from self-aggregation and enhances its kinase activity". J. Biol. Chem. 267 (10): 7042–7. doi:10.1016/S0021-9258(19)50533-6. PMID 1551911.
- Wiech H, Buchner J, Zimmermann R, Jakob U (July 1992). "Hsp90 chaperones protein folding in vitro". Nature. 358 (6382): 169–70. doi:10.1038/358169a0. PMID 1614549. S2CID 4316363.
- Jakob U, Lilie H, Meyer I, Buchner J (March 1995). "Transient interaction of Hsp90 with early unfolding intermediates of citrate synthase. Implications for heat shock in vivo". J. Biol. Chem. 270 (13): 7288–94. doi:10.1074/jbc.270.13.7288. PMID 7706269.
- Picard D (October 2002). "Heat-shock protein 90, a chaperone for folding and regulation". Cell. Mol. Life Sci. 59 (10): 1640–8. doi:10.1007/PL00012491. PMID 12475174. S2CID 34094587.
- Imai J, Maruya M, Yashiroda H, Yahara I, Tanaka K (July 2003). "The molecular chaperone Hsp90 plays a role in the assembly and maintenance of the 26S proteasome". EMBO J. 22 (14): 3557–67. doi:10.1093/emboj/cdg349. PMC 165619. PMID 12853471.
- Correia MA, Sadeghi S, Mundo-Paredes E (2005). "Cytochrome P450 ubiquitination: branding for the proteolytic slaughter?". Annu. Rev. Pharmacol. Toxicol. 45: 439–64. doi:10.1146/annurev.pharmtox.45.120403.100127. PMID 15822184.
- Kimura Y, Matsumoto S, Yahara I (March 1994). "Temperature-sensitive mutants of hsp82 of the budding yeast Saccharomyces cerevisiae". Mol. Gen. Genet. 242 (5): 517–27. doi:10.1007/BF00285275. PMID 8121410. S2CID 36722145.
- Davies TH, Ning YM, Sánchez ER (February 2002). "A new first step in activation of steroid receptors: hormone-induced switching of FKBP51 and FKBP52 immunophilins". J. Biol. Chem. 277 (7): 4597–600. doi:10.1074/jbc.C100531200. PMID 11751894.
- Pałyga J, Kozłowski Ł (2007). "Structure and function of molecular chaperone HSP90". Sowriemiennyj Naucznyj Wiestnik Ser. Biologija Chimija. 15 (23): 46–65.
- Pratt WB, Morishima Y, Murphy M, Harrell M (2006). "Chaperoning of glucocorticoid receptors". Molecular Chaperones in Health and Disease. Handbook of Experimental Pharmacology. Vol. 172. pp. 111–38. doi:10.1007/3-540-29717-0_5. ISBN 978-3-540-25875-9. PMID 16610357.
- Grad I, Picard D (September 2007). "The glucocorticoid responses are shaped by molecular chaperones". Mol. Cell. Endocrinol. 275 (1–2): 2–12. doi:10.1016/j.mce.2007.05.018. PMID 17628337. S2CID 22117642.
- Pratt WB, Galigniana MD, Morishima Y, Murphy PJ (2004). "Role of molecular chaperones in steroid receptor action". Essays Biochem. 40: 41–58. doi:10.1042/bse0400041. PMID 15242338.
- Rafestin-Oblin ME, Couette B, Radanyi C, Lombes M, Baulieu EE (June 1989). "Mineralocorticosteroid receptor of the chick intestine. Oligomeric structure and transformation". J. Biol. Chem. 264 (16): 9304–9. doi:10.1016/S0021-9258(18)60531-9. PMID 2542305.
- Joab I, Radanyi C, Renoir M, Buchou T, Catelli MG, Binart N, Mester J, Baulieu EE (1984). "Common non-hormone binding component in non-transformed chick oviduct receptors of four steroid hormones". Nature. 308 (5962): 850–3. doi:10.1038/308850a0. PMID 6201744. S2CID 4303649.
- Redeuilh G, Moncharmont B, Secco C, Baulieu EE (May 1987). "Subunit composition of the molybdate-stabilized "8-9 S" nontransformed estradiol receptor purified from calf uterus". J. Biol. Chem. 262 (15): 6969–75. doi:10.1016/S0021-9258(18)48188-4. PMID 3584104.
- Catelli MG, Binart N, Jung-Testas I, Renoir JM, Baulieu EE, Feramisco JR, Welch WJ (December 1985). "The common 90-kd protein component of non-transformed '8S' steroid receptors is a heat-shock protein". EMBO J. 4 (12): 3131–5. doi:10.1002/j.1460-2075.1985.tb04055.x. PMC 554632. PMID 2419124.
- Lurje G, Lenz HJ (2009). "EGFR Signaling and Drug Discovery". Oncology. 77 (6): 400–410. doi:10.1159/000279388. PMID 20130423. S2CID 7638236.
- Sawai A, Chandarlapaty S, Greulich H, Gonen M, Ye Q, Arteaga CL, Sellers W, Rosen N, Solit DB (January 2008). "Inhibition of Hsp90 down-regulates mutant epidermal growth factor receptor (EGFR) expression and sensitizes EGFR mutant tumors to paclitaxel". Cancer Res. 68 (2): 589–96. doi:10.1158/0008-5472.CAN-07-1570. PMC 4011195. PMID 18199556.
- Paez-Ribes M, González-Gualda E, Doherty GJ, Muñoz-Espín D (2019). "Targeting senescent cells in translational medicine". EMBO Molecular Medicine. 11 (12): e10234. doi:10.15252/emmm.201810234. PMC 6895604. PMID 31746100.
- Mohsin SK, Weiss HL, Gutierrez MC, Chamness GC, Schiff R, Digiovanna MP, Wang CX, Hilsenbeck SG, Osborne CK, Allred DC, Elledge R, Chang JC (April 2005). "Neoadjuvant trastuzumab induces apoptosis in primary breast cancers". J. Clin. Oncol. 23 (11): 2460–8. doi:10.1200/JCO.2005.00.661. PMID 15710948.
- Smalley M, Natarajan SK, Mondal J, Best D, Goldman D, Shanthappa B, et al. (December 2020). "Nanoengineered Disruption of Heat Shock Protein 90 Targets Drug-Induced Resistance and Relieves Natural Killer Cell Suppression in Breast Cancer". Cancer Research. 80 (23): 5355–5366. doi:10.1158/0008-5472.CAN-19-4036. PMC 7718318. PMID 33077554.
- Calderwood SK, Khaleque MA, Sawyer DB, Ciocca DR (March 2006). "Heat shock proteins in cancer: chaperones of tumorigenesis". Trends in Biochemical Sciences. 31 (3): 164–72. doi:10.1016/j.tibs.2006.01.006. PMID 16483782.
- Eustace BK, Sakurai T, Stewart JK, Yimlamai D, Unger C, Zehetmeier C, Lain B, Torella C, Henning SW, Beste G, Scroggins BT, Neckers L, Ilag LL, Jay DG (June 2004). "Functional proteomic screens reveal an essential extracellular role for hsp90 alpha in cancer cell invasiveness". Nat. Cell Biol. 6 (6): 507–14. doi:10.1038/ncb1131. PMID 15146192. S2CID 40025264.
- Whitesell L, Lindquist SL (October 2005). "Hsp90 and the chaperoning of cancer". Nat. Rev. Cancer. 5 (10): 761–72. doi:10.1038/nrc1716. PMID 16175177. S2CID 22098282.
- Kim YS, Alarcon SV, Lee S, Lee MJ, Giaccone G, Neckers L, Trepel JB (2009). "Update on Hsp90 inhibitors in clinical trial". Curr Top Med Chem. 9 (15): 1479–92. doi:10.2174/156802609789895728. PMC 7241864. PMID 19860730.
- Pires ES, Khole VV (2009). "A block in the road to fertility: autoantibodies to heat-shock protein 90-beta in human ovarian autoimmunity". Fertil Steril. 92 (4): 1395–1409. doi:10.1016/j.fertnstert.2008.08.068. PMID 19022436.
- Pires ES, Choudhury AK, Idicula-Thomas S, Khole VV (2011). "Anti-HSP90 autoantibodies in sera of infertile women identify a dominant, conserved epitope EP6 (380-389) of HSP90 beta protein". Reprod Biol Endocrinol. 9 (16): 13. doi:10.1186/1477-7827-9-16. PMC 3039567. PMID 21272367.
- Gupta RS (November 1995). "Phylogenetic analysis of the 90 kD heat shock family of protein sequences and an examination of the relationship among animals, plants, and fungi species". Mol. Biol. Evol. 12 (6): 1063–73. doi:10.1093/oxfordjournals.molbev.a040281. PMID 8524040.
- Gupta RS, Aitken K, Falah M, Singh B (April 1994). "Cloning of Giardia lamblia heat shock protein HSP70 homologs: implications regarding origin of eukaryotic cells and of endoplasmic reticulum". Proc. Natl. Acad. Sci. U.S.A. 91 (8): 2895–9. doi:10.1073/pnas.91.8.2895. PMC 43480. PMID 8159675.
- Gupta RS, Golding GB (May 1996). "The origin of the eukaryotic cell". Trends Biochem. Sci. 21 (5): 166–71. doi:10.1016/S0968-0004(96)20013-1. PMID 8871398.
- Gupta RS (December 1998). "Protein phylogenies and signature sequences: A reappraisal of evolutionary relationships among archaebacteria, eubacteria, and eukaryotes". Microbiol. Mol. Biol. Rev. 62 (4): 1435–91. doi:10.1128/MMBR.62.4.1435-1491.1998. PMC 98952. PMID 9841678.
External links
- Hsp90+Heat-Shock+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Didier Picard. "Hsp90 web site". VisibilityWeb. Archived from the original on 2004-11-23. Retrieved 2008-07-07.
A web site for the community of scientists interested in the Hsp90 molecular chaperone machine.