Salmonella enterica
Salmonella enterica (formerly Salmonella choleraesuis) is a rod-shaped, flagellate, facultative anaerobic, Gram-negative bacterium and a species of the genus Salmonella.[1] A number of its serovars are serious human pathogens; many of them are (more specifically) serovars of Salmonella enterica subsp. enterica.
| Salmonella enterica | |
|---|---|
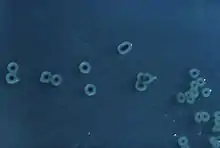 | |
| S. enterica Typhimurium colonies on a Hektoen enteric agar plate | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Gammaproteobacteria |
| Order: | Enterobacterales |
| Family: | Enterobacteriaceae |
| Genus: | Salmonella |
| Species: | S. enterica |
| Binomial name | |
| Salmonella enterica (ex Kauffmann & Edwards 1952) Le Minor & Popoff 1987 | |
| Subspecies | |
| |
Epidemiology
Most cases of salmonellosis are caused by food infected with S. enterica, which often infects cattle and poultry, though other animals such as domestic cats[2][3] and hamsters[4] have also been shown to be sources of infection in humans. Investigations of vacuum cleaner bags have shown that households can act as a reservoir of the bacterium; this is more likely if the household has contact with an infection source (i.e., members working with cattle or in a veterinary clinic).
Raw chicken eggs and goose eggs can harbor S. enterica, initially in the egg whites, although most eggs are not infected. As the egg ages at room temperature, the yolk membrane begins to break down and S. enterica can spread into the yolk. Refrigeration and freezing do not kill all the bacteria, but substantially slow or halt their growth. Pasteurizing and food irradiation are used to kill Salmonella for commercially produced foodstuffs containing raw eggs such as ice cream. Foods prepared in the home from raw eggs, such as mayonnaise, cakes, and cookies, can spread salmonellae if not properly cooked before consumption.
S. enterica genomes have been reconstructed from up 6,500 year old human remains across Western Eurasia, which provides evidence for geographic widespread infections with systemic S. enterica during prehistory, and a possible role of the Neolithization process in the evolution of host adaptation.[5] Additional reconstructed genomes from colonial Mexico suggest S. enterica as the cause of cocoliztli, an epidemic in 16th-century New Spain.[6]
Pathogenesis
Secreted proteins are of major importance for the pathogenesis of infectious diseases caused by S. enterica. A remarkably large number of fimbrial and nonfimbrial adhesins are present in Salmonella, and mediate biofilm formation and contact to host cells. Secreted proteins are also involved in host-cell invasion and intracellular proliferation, two hallmarks of Salmonella pathogenesis.[7]
DNA repair capability
Exposure of S. enterica to bile salts, such as sodium deoxycholate, induces the SOS DNA damage response indicating that in this organism bile salts cause DNA damage.[8] Bile salt exposure is found to increase GC to AT transition mutations and also to induce genes of the OxyR and SoxRS regulons suggesting further that bile salts specifically cause oxidative DNA damage.[8] Mutants of S. enterica that are defective in enzymes required for the process of base excision repair are sensitive to bile salts. This indicates that wild-type S. enterica uses base excision repair to remove DNA damages caused by the bile salts.[8] The RecBCD enzyme which functions in recombinational repair of DNA is also required for bile salt resistance.
Small noncoding RNA
Small nonprotein-coding RNAs (sRNA) are able to perform specific functions without being translated into proteins; 97 bacterial sRNAs from Salmonella Typhi were discovered.[9]
AsdA (antisense RNA of dnaA) is a cis-encoded antisense RNA of dnaA described in S. enterica serovar Typhi. It was discovered by deep sequencing and its transcription was confirmed by Northern blot and RACE analysis. AsdA is estimated to be about 540 nucleotides long, and represents the complementary strand to that encoding DnaA, a protein that plays a central role in the initiation of DNA replication and hence cellular division. In rich media, it is highly expressed only after reaching the stationary growth phase, but under limiting iron or osmotic stress, it is already expressed during exponential growth. Overexpression of AsdA stabilizes dnaA mRNA, increasing its levels and thereby enhancing its rate of translation. This suggests that AsdA is a regulator of DNA replication.[10]
Nomenclature
S. enterica has six subspecies, and each subspecies has associated serovars that differ by antigenic specificity.[11] S. enterica has over 2500 serovars.[12] Salmonella bongori was previously considered a subspecies of S. enterica, but it is now the other species in the genus Salmonella. Most of the human pathogenic Salmonella serovars belong to the enterica subspecies. These serogroups include S. Typhi, S. Enteritidis, S. Paratyphi, S. Typhimurium, and S. Choleraesuis. The serovars can be designated as written in the previous sentence (capitalized and nonitalicized following the genus), or as follows: "S. enterica subsp. enterica, serovar Typhi".
S. e. subsp. arizonae, named after the state of Arizona, is most commonly found in cold-blooded animals (especially snakes), but can also infect turkey, sheep, and humans. It is endemic in southwestern United States.[13] The similar S. e. subsp. diarizonae also infects snakes and occasionally humans.[14]
See also
References
- Giannella RA (1996). Baron S; et al. (eds.). Salmonella. In: Baron's Medical Microbiology (4th ed.). Univ of Texas Medical Branch. ISBN 978-0-9631172-1-2. (via NCBI Bookshelf).
- "Salmonellosis in Animals - Digestive System". Veterinary Manual. Retrieved 2021-01-01.
- Giacometti, Federica; Magarotto, Jacopo; Serraino, Andrea; Piva, Silvia (2017-07-24). "Highly suspected cases of salmonellosis in two cats fed with a commercial raw meat-based diet: health risks to animals and zoonotic implications". BMC Veterinary Research. 13 (1): 224. doi:10.1186/s12917-017-1143-z. ISSN 1746-6148. PMC 5525297. PMID 28738871.
- Swanson SJ, Snider C, Braden CR, et al. (2007). "Multidrug-resistant Salmonella enterica serotype Typhimurium associated with pet rodents". New England Journal of Medicine. 356 (1): 21–28. doi:10.1056/NEJMoa060465. PMID 17202452.
- Key, Felix M.; Posth, Cosimo; Esquivel-Gomez, Luis R.; Hübler, Ron; Spyrou, Maria A.; Neumann, Gunnar U.; Furtwängler, Anja; Sabin, Susanna; Burri, Marta; Wissgott, Antje; Lankapalli, Aditya Kumar; Vågene, Åshild J.; Meyer, Matthias; Nagel, Sarah; Tukhbatova, Rezeda; Khokhlov, Aleksandr; Chizhevsky, Andrey; Hansen, Svend; Belinsky, Andrey B.; Kalmykov, Alexey; Kantorovich, Anatoly R.; Maslov, Vladimir E.; Stockhammer, Philipp W.; Vai, Stefania; Zavattaro, Monica; Riga, Alessandro; Caramelli, David; Skeates, Robin; Beckett, Jessica; Gradoli, Maria Giuseppina; Steuri, Noah; Hafner, Albert; Ramstein, Marianne; Siebke, Inga; Lösch, Sandra; Erdal, Yilmaz Selim; Alikhan, Nabil-Fareed; Zhou, Zhemin; Achtman, Mark; Bos, Kirsten; Reinhold, Sabine; Haak, Wolfgang; Kühnert, Denise; Herbig, Alexander; Krause, Johannes (March 2020). "Emergence of human-adapted Salmonella enterica is linked to the Neolithization process". Nature Ecology & Evolution. 4 (3): 324–333. doi:10.1038/s41559-020-1106-9. ISSN 2397-334X. PMC 7186082. PMID 32094538.
- Vågene, Åshild J.; Herbig, Alexander; Campana, Michael G.; Robles García, Nelly M.; Warinner, Christina; Sabin, Susanna; Spyrou, Maria A.; Andrades Valtueña, Aida; Huson, Daniel; Tuross, Noreen; Bos, Kirsten I.; Krause, Johannes (2018). "Salmonella enterica genomes from victims of a major sixteenth-century epidemic in Mexico". Nature, Ecology & Evolution. 2 (3): 520–528. doi:10.1038/s41559-017-0446-6. PMID 29335577. S2CID 3358440.
- Hensel M (2009). "Secreted Proteins and Virulence in Salmonella enterica". Bacterial Secreted Proteins: Secretory Mechanisms and Role in Pathogenesis. Caister Academic Press. ISBN 978-1-904455-42-4.
- Prieto AI, Ramos-Morales F, Casadesús J. Repair of DNA damage induced by bile salts in Salmonella enterica. Genetics. 2006 Oct;174(2):575-84. doi: 10.1534/genetics.106.060889. Epub 2006 Aug 3. PMID 16888329; PMCID: PMC1602091
- Chinni, Suresh V.; Raabe, Carsten A.; Zakaria, Robaiza; Randau, Gerrit; Hoe, Chee Hock; Zemann, Anja; Brosius, Juergen; Tang, Thean-Hock; Rozhdestvensky, Timofey S. (2010-09-01). "Experimental identification and characterization of 97 novel npcRNA candidates in Salmonella enterica serovar Typhi". Nucleic Acids Research. 38 (17): 5893–5908. doi:10.1093/nar/gkq281. ISSN 1362-4962. PMC 2943607. PMID 20460466.
- Dadzie, Isaac; Xu, Shungao; Ni, Bin; Zhang, Xiaolei; Zhang, Haifang; Sheng, Xiumei; Xu, Huaxi; Huang, Xinxiang (2013-01-01). "Identification and characterization of a cis-encoded antisense RNA associated with the replication process of Salmonella enterica serovar Typhi". PLOS ONE. 8 (4): e61308. Bibcode:2013PLoSO...861308D. doi:10.1371/journal.pone.0061308. ISSN 1932-6203. PMC 3634043. PMID 23637809.
- Todar, Kenneth. "Salmonella and Salmonellosis". Todar's Online Textbook of Bacteriology.
- Murray PR, Rosenthal KS, Pfaller MA (2009). Medical Microbiology (6th ed.). Philadelphia, PA: Mosby Elsevier. p. 307.
- Lee, YC; Hung, MC; Hung, SC; Wang, HP; Cho, HL; Lai, MC; Wang, JT (9 December 2016). "Salmonella enterica subspecies arizonae infection of adult patients in Southern Taiwan: a case series in a non-endemic area and literature review". BMC Infectious Diseases. 16 (1): 746. doi:10.1186/s12879-016-2083-0. PMC 5148916. PMID 27938338.
- Schröter, M; Roggentin, P; Hofmann, J; Speicher, A; Laufs, R; Mack, D (January 2004). "Pet snakes as a reservoir for Salmonella enterica subsp. diarizonae (Serogroup IIIb): a prospective study". Applied and Environmental Microbiology. 70 (1): 613–5. Bibcode:2004ApEnM..70..613S. doi:10.1128/AEM.70.1.613-615.2004. PMC 321278. PMID 14711697.
External links
- Notes on Salmonella nomenclature
- Salmonella+enterica at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Current research on Salmonella typhimurium at the Norwich Research Park
- "Salmonella enterica". NCBI Taxonomy Browser. 28901.
- Type strain of Salmonella enterica at BacDive - the Bacterial Diversity Metadatabase