Nucleolus organizer region
Nucleolus organizer regions (NORs) are chromosomal regions crucial for the formation of the nucleolus. In humans, the NORs are located on the short arms of the acrocentric chromosomes 13, 14, 15, 21 and 22, the genes RNR1, RNR2, RNR3, RNR4, and RNR5 respectively.[1] These regions code for 5.8S, 18S, and 28S ribosomal RNA.[1] The NORs are "sandwiched" between the repetitive, heterochromatic DNA sequences of the centromeres and telomeres.[1] The exact sequence of these regions is not included in the human reference genome as of 2016[1] or the GRCh38.p10 released January 6, 2017.[2] On 28 February 2019, GRCh38.p13 was released, which added the NOR sequences for the short arms of chromosomes 13, 14, 15, 21, and 22.[3] However, it is known that NORs contain tandem copies of ribosomal DNA (rDNA) genes.[1] Some sequences of flanking sequences proximal and distal to NORs have been reported.[4] The NORs of a loris have been reported to be highly variable.[5] There are also DNA sequences related to rDNA that are on other chromosomes and may be involved in nucleoli formation.[6]
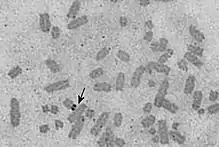
Visualization
Barbara McClintock first described the "nucleolar-organizing body" in Zea mays in 1934.[7] In karyotype analysis, a silver stain can be used to identify the NOR.[8][9] NORs can also be seen in nucleoli using silver stain, and that is being used to investigate cancerous changes.[10][11][12] NORs can also be seen using antibodies directed against the protein UBF, which binds to NOR DNA.[1]
Molecular biology
In addition to UBF, NORs also bind to ATRX protein, treacle, sirtuin-7 and other proteins.[1] UBF has been identified as a mitotic "bookmark" of expressed rDNA, which allows it to resume transcription quickly after mitosis.[1] The distal flanking junction (DJ) of the NORs has been shown to associate with the periphery of nucleoli.[4] rDNA operons in Escherichia coli have been found to cluster near each other, similar to a eukaryotic nucleolus.[13]
See also
References
- McStay B (2016). "Nucleolar organizer regions: genomic 'dark matter' requiring illumination". Genes & Development. 30 (14): 1598–610. doi:10.1101/gad.283838.116. PMC 4973289. PMID 27474438.
- anon. "Human Genome Overview". ncbi.nlm.nih.gov/. Genome Reference Consortium. Retrieved 17 June 2017.
- The Genome Reference Consortium. "GRCh38.p13 has been released". GenomeRef. Retrieved 16 August 2019.
- Floutsakou I, Agrawal S, Nguyen TT, Seoighe C, Ganley AR, McStay B (2013). "The shared genomic architecture of human nucleolar organizer regions". Genome Research. 23 (12): 2003–12. doi:10.1101/gr.157941.113. PMC 3847771. PMID 23990606.
- Baicharoen S, Hirai Y, Srikulnath K, Kongprom U, Hirai H (2016). "Hypervariability of Nucleolus Organizer Regions in Bengal Slow Lorises, Nycticebus bengalensis (Primates, Lorisidae)". Cytogenetic and Genome Research. 149 (4): 267–273. doi:10.1159/000449145. PMID 27648559.
- Kupriyanova NS, Netchvolodov KK, Sadova AA, Cherepanova MD, Ryskov AP (2015). "Non-canonical ribosomal DNA segments in the human genome, and nucleoli functioning". Gene. 572 (2): 237–42. doi:10.1016/j.gene.2015.07.019. PMID 26164756.
- McClintock B (1934). "The relation of a particular chromosomal element to the development of the nucleoli in Zea mays". Zeitschrift für Zellforschung und Mikroskopische Anatomie. 21 (2): 294–328. doi:10.1007/BF00374060.
- Bloom SE, Goodpasture C (October 1976). "An improved technique for selective silver staining of nucleolar organizer regions in human chromosomes". Human Genetics. 34 (2): 199–206. doi:10.1007/bf00278889. PMID 63440.
- Lau YF, Pfeiffer RA, Arrighi FE, Hsu TC (January 1978). "Combination of silver and fluorescent staining for metaphase chromosomes". American Journal of Human Genetics. 30 (1): 76–9. PMC 1685453. PMID 74950.
- Stepan A, Simionescu C, Pirici D, Ciurea R, Margaritescu C (2015). "Fractal analysis and the diagnostic usefulness of silver staining nucleolar organizer regions in prostate adenocarcinoma". Analytical Cellular Pathology. 2015: 250265. doi:10.1155/2015/250265. PMC 4558419. PMID 26366372.
- Gundog M, Yildiz OG, Imamoglu N, Aslan D, Aytekin A, Soyuer I, Soyuer S (2015). "Prognostic Significance of Two Dimensional AgNOR Evaluation in Local Advanced Rectal Cancer Treated with Chemoradiotherapy". Asian Pacific Journal of Cancer Prevention. 16 (18): 8155–61. doi:10.7314/apjcp.2015.16.18.8155. PMID 26745054.
- Sowmya GV, Nahar P, Astekar M, Agarwal H, Singh MP (2017). "Analysis of silver binding nucleolar organizer regions in exfoliative cytology smears of potentially malignant and malignant oral lesions". Biotechnic & Histochemistry. 92 (2): 115–121. doi:10.1080/10520295.2017.1283055. PMID 28296547.
- Gaal T, Bratton BP, Sanchez-Vazquez P, Sliwicki A, Sliwicki K, Vegel A, Pannu R, Gourse RL (2016). "Colocalization of distant chromosomal loci in space in E. coli: a bacterial nucleolus". Genes & Development. 30 (20): 2272–2285. doi:10.1101/gad.290312.116. PMC 5110994. PMID 27898392.