Sin Nombre orthohantavirus
Sin Nombre orthohantavirus (SNV) (from Spanish, meaning "without a name") is the prototypical etiologic agent of hantavirus cardiopulmonary syndrome (HCPS).[1]
| Sin Nombre orthohantavirus | |
|---|---|
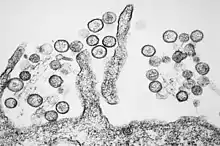 | |
| Transmission electron micrograph of Sin Nombre orthohantavirus | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Negarnaviricota |
| Class: | Ellioviricetes |
| Order: | Bunyavirales |
| Family: | Hantaviridae |
| Genus: | Orthohantavirus |
| Species: | Sin Nombre orthohantavirus |
| Member viruses | |
| Synonyms | |
| |
| Sin Nombre virus | |
|---|---|
| Specialty | Virology |
Discovered in 1993 near the Cañon de la Muerte on the Navajo Reservation, it was originally named the Muerto Canyon hantavirus, in keeping with the convention for naming new pathogens.[2] However, the Navajo Nation objected to the name in 1994.[3] It was also near the Four Corners point in the United States, so the virologists then tried naming it the "Four Corners virus". The name was changed after local residents raised objections.[4] In frustration, the virologists changed it to Sin Nombre, meaning "without a name" in Spanish.
History
It was first isolated in 1993 from rodents collected near the home of one of the initial patients with hantavirus pulmonary syndrome (HPS) in the Four Corners region of the western United States. Isolation was achieved through blind passage in Peromyscus maniculatus (deer mouse) and subsequent adaptation to growth in Vero E6 cells. Additional viral strains have also been isolated from P. maniculatus associated with a fatal case in California and P. leucopus from the vicinity of probable infection of a New York case. Black Creek Canal virus was isolated from S. hispidus collected near the residence of a human case in Dade County, Florida. Another etiologic agent of HCPS, Bayou virus, was first isolated from the vicinity of Monroe, Louisiana.[5]
Epidemiology
SNV occurs wherever its reservoir rodent carrier, the deer mouse Peromyscus maniculatus,[6] is found, which includes essentially the entire populated area of North America, except for the far southeastern region from eastern Texas through Florida, Alaska, and the far northern reaches of Canada. SNV and HCPS are especially common in western states; peak incidences for HCPS have been reported in regions in which there is a lot of contact between humans and mice (New Mexico, Arizona) and in states with exceptionally large rural populations such as California. All of the western provinces of Canada have also reported cases. SNV can be contracted through the inhalation of virus-contaminated deer mouse excreta.
The case fatality ratio of SNV-induced HCPS in the USA was reported to be about 66.7% (CDC, 1993). However, since that time the case fatality ratio has steadily declined as more mild cases came to be recognized. By 2007 the CFR had declined to about 35%.
Virus sequencing
The entire genomic sequence of SNV has subsequently been determined by using RNA extracted from autopsy material as well as RNA extracted from cell culture-adapted virus. The L RNA is 6562 nucleotides (nt) in length; the M RNA is 3696 nt long; and the S RNA is 2059 to 2060 nt long. When the prototype sequence (NMH15) of SNV detected in tissues from an HPS case was compared with the sequence of the SNV isolate (NMR11; isolated in Vero E6 cells from Peromyscus maniculatus trapped in the residence of the same case), only 16 nucleotide changes were found, and none of these changes resulted in alterations in amino acid sequences of viral proteins. It had been assumed that in the process of adaptation to cell culture, selection of SNV variants which grow optimally in cell culture would occur, and selected variants would differ genetically from the parental virus. Though NMH10 and NMR11 are identical in protein sequence, nucleotide substitutions in nontranslated regions of the genome could be responsible for altered viral phenotypes, as could changes in protein glycosylation or virus membrane components.
The nested RT-PCR assay developed during the initial HCPS outbreak provided a rapid method for the genetic characterization of novel hantaviruses that did not require a virus isolate. Numerous new hantaviruses have been detected by RT-PCR in rodent tissues but have yet to be associated with human disease. These include El Moro Canyon virus associated with the western harvest mouse, Reithrodontomys megalotis, Tula virus with Microtus arvalis and M. rossiaemeridionalis, Rio Segundo virus with the Mexican harvest mouse, R. mexicanus, Isla Vista virus with the California vole, M. californicus, and Prospect Hill-like viruses in Microtus species.
References
- Ye C, Prescott J, Nofchissey R, Goade D, Hjelle B (March 2004). "Neutralizing antibodies and Sin Nombre virus RNA after recovery from hantavirus cardiopulmonary syndrome". Emerging Infect. Dis. 10 (3): 478–82. doi:10.3201/eid1003.020821 (inactive 31 July 2022). PMC 3322788. PMID 15109416.
{{cite journal}}: CS1 maint: DOI inactive as of July 2022 (link) - Van Hook, Charles J. (November 2018). "Hantavirus Pulmonary Syndrome—The 25th Anniversary of the Four Corners Outbreak". Emerging Infectious Diseases. 24 (11): 2056–2060. doi:10.3201/eid2411.180381. PMC 6199996.
- "Navajos Decry Muerto Canyon Hantavirus Site". Los Angeles Times. April 24, 1994. Retrieved 3 July 2019.
- Strauss, Ellen G.; Strauss, James H. (2002). Viruses and human disease. Boston: Academic Press. p. 161. ISBN 978-0-12-673050-0.
- "Hantaviruses, with emphasis on Four Corners Hantavirus". Bvs.insp.mx. Archived from the original on 2013-04-20. Retrieved 2016-11-17.
- Lehmer EM, Clay CA, Pearce-Duvet J, St Jeor S, Dearing MD (March 2008). "Differential regulation of pathogens: the role of habitat disturbance in predicting prevalence of Sin Nombre virus". Oecologia. 155 (3): 429–39. Bibcode:2008Oecol.155..429L. doi:10.1007/s00442-007-0922-9. PMID 18064494. S2CID 19495085.